内容补充--细胞游离DNA片段组学在癌症中的应用
原创内容补充--细胞游离DNA片段组学在癌症中的应用
原创
追风少年i
修改于 2025-10-06 11:21:19
修改于 2025-10-06 11:21:19
作者,Evil Genius
肿瘤早筛:全称是肿瘤早期筛查,指的是在表面健康、尚未出现任何癌症症状的人群中,通过简便、有效的检查方法,发现那些已经患有早期癌症或癌前病变的个体。
基于血液的液体活检是当前肿瘤早筛领域最前沿的方向。
原理:通过抽血,检测血液中微量的、由肿瘤细胞释放的循环肿瘤DNA(ctDNA)。
肿瘤早筛目前NGS主要的分析手段
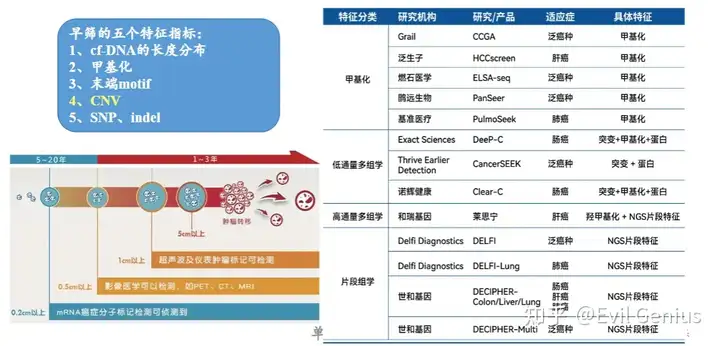
分析维度 | 核心思路 | 技术特点 |
|---|---|---|
DNA甲基化 | 检测DNA分子上特定的甲基化修饰(5mC)模式。癌细胞会表现出全基因组的低甲基化和特定区域(如抑癌基因启动子)的高甲基化。 | 直接反映基因的表达调控,是当前最主流的技术路径之一。通过检测血浆中循环肿瘤DNA(ctDNA)的特异性甲基化信号,可实现癌症的早期筛查和组织溯源定位。 |
片段组学 | 分析血液中游离DNA(cfDNA)的片段大小、末端序列、核小体保护模式等特征。癌来源的ctDNA与正常细胞释放的DNA在这些特征上存在差异。 | 是一种间接的分析方法,不直接检测序列变异。它能提供与甲基化互补的信息,可与甲基化等多组学数据整合,构建更精准的AI模型。 |
基因突变 | 检测与癌症驱动相关的基因序列改变,例如单核苷酸变异(SNV)、插入/缺失(Indel)等。 | 是肿瘤基因检测的经典方法。但在早筛中,因早期肿瘤ctDNA含量极低,突变信号较弱,其灵敏度通常低于甲基化,更多作为补充信息 |
基因突变是我们课程主要的内容,甲基化需要另外的课程部分,也是目前早筛最成熟的技术,今天我们来补充片段组学。
Cell-free DNA片段组学的发展
细胞游离DNA(cfDNA)是指存在于血液、尿液及其他体液中、未包裹在细胞内的片段化DNA分子。早期对cfDNA的兴趣源于三大发现:(1)通过基于PCR的肿瘤相关基因变异检测,在癌症患者血浆中发现循环肿瘤DNA(ctDNA);(2)通过PCR检测Y染色体DNA,在孕妇血浆和血清中发现胎儿cfDNA;(3)通过供体特异性基因变异检测,在器官移植受者血浆中发现供体来源cfDNA。这些发现推动了用于癌症检测的微创液体活检技术、靶向治疗设计、产前检测及移植物排斥反应监测的发展。
cfDNA片段组学致力于研究cfDNA的片段化模式,包括但不限于片段大小、末端基序、末端密度("偏好性末端")及核小体占据情况。早期对cfDNA尺寸谱的分析显示,人血浆DNA的模态长度为166 bp,相当于一个核小体DNA加上连接DNA的长度。胎儿与肿瘤DNA的模态尺寸较短(143 bp),且<143 bp的片段呈现10 bp周期性。近期,第三代单分子测序技术解析了Illumina平台此前无法检测的"长链cfDNA"(>500 bp),其癌症诊断潜力正被积极探索¹。另一方面,传统cfDNA提取及双链建库方法可能丢弃的"超短cfDNA"(∼50 bp),正逐渐显现出具有重要生物学意义的价值。
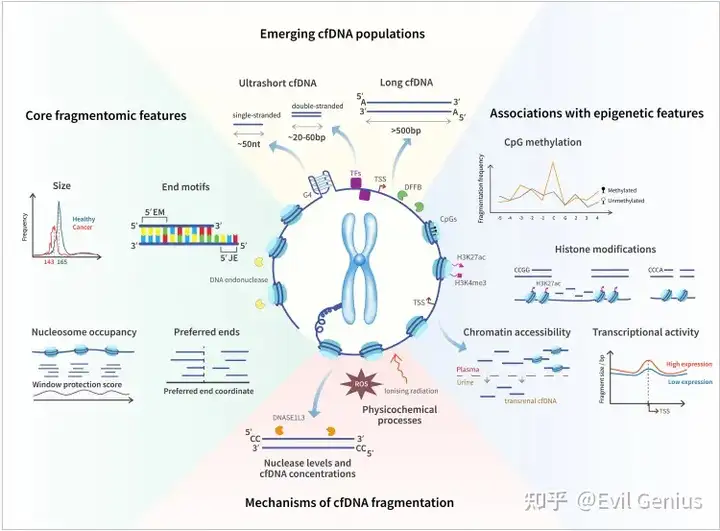
事实上,学界长期认为细胞死亡是cfDNA生成的重要来源。
研究发现核酸酶活性不仅与cfDNA尺寸谱相关,还与血浆DNA片段末端的序列模式(末端基序)有关。
核酸酶参与cfDNA片段化过程,为cfDNA片段组学模式与基因组调控机制(如染色质结构与表观遗传DNA修饰)及其转录组输出之间的新兴关联提供了生物学基础。早期证据包括"偏好性末端"的发现——即某些基因组坐标位点存在cfDNA末端的过度呈现,且其分布在肿瘤与非肿瘤基因组中存在差异。这种cfDNA覆盖模式的不均匀性还可用于核小体及转录因子足迹分析,可能携带组织或肿瘤特异性特征。在分子层面研究发现,CpG位点周围的片段化程度取决于其甲基化状态,尽管介导此类核酸酶甲基化敏感行为的机制尚未完全阐明。目前已知CpG甲基化影响cfDNA覆盖度推断的核小体定位;此外,近期研究表明调节染色质可及性、可作为典型癌症生物标志物的组蛋白修饰,与特征性片段化模式相关。这些关联提供了生物学依据,使得通过将核小体占据与转录行为相关联,可直接从cfDNA片段化谱推断RNA表达。
癌症是一种涉及多种表观遗传与转录组改变以及细胞更新异常的多因素疾病。有证据表明DNA片段化受底层染色质结构影响,而染色质结构与DNA甲基化、组蛋白修饰及转录活性密切相关。此外,cfDNA片段组学可能反映细胞死亡模式,其中核酸酶介导的DNA降解被认为是重要环节。本质上,cfDNA片段组学可能贯通了癌症生物学的这些多元维度。
新型片段组学特征为癌症检测开辟全新路径
cfDNA的片段大小分布、末端基序及足迹模式已被确立为核心片段组学特征
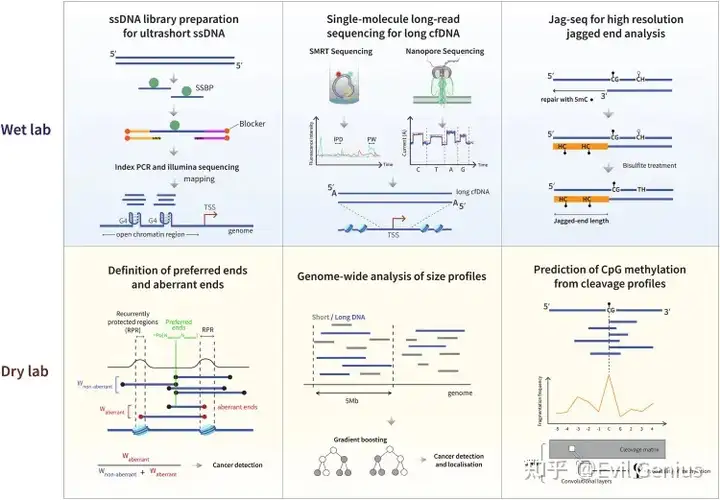
传统核小体相关片段(~166 bp)之外的两种特殊cfDNA:
超短cfDNA (~50 bp):主要采用单链建库技术捕获,富含启动子等调控区域信息,并能透过肾小球形成尿源cfDNA,在癌症检测中显示出独特价值。
基因组特征与癌症关联:
- 基因组来源:超短cfDNA在启动子区域有普遍富集。
- 癌症变化:在晚期癌症患者中,转录起始点上游的超短cfDNA覆盖度显著降低,这可能与疾病分期相关。
- 潜在来源:其形成可能与G-四链体结构或转录因子结合位点有关,并能揭示组织特异性信号。
长链cfDNA (>500 bp):需用第三代测序技术分析,富含在转录活跃的常染色质区域,能提供更完整的单分子表观遗传信息,但当前测序通量低是主要应用瓶颈。
在癌症中的表现与生成机制:
- 癌症关联:在肝细胞癌患者中,映射到高表达基因的长链cfDNA丰度显著高于健康对照或乙肝携带者,能有效区分HCC与非HCC。
- 生成机制:DFFB核酸酶在生成长链cfDNA中起重要作用,其切割活性受高阶染色质结构调控。长链cfDNA的末端密度在CTCF结合位点等区域显著激增,有望用于描绘癌症中特异的染色质组织异常模式。
从细胞游离DNA片段组学解码表观遗传与转录信号
通过分析cfDNA的片段化模式(片段组学),可以反向推断其来源细胞的表观遗传状态(如DNA甲基化、组蛋白修饰)和转录活性,而无需依赖传统的、具有破坏性的实验方法(如亚硫酸盐处理、ChIP-seq)。
1. 预测DNA甲基化状态
核心发现:cfDNA在CpG位点周围的切割模式与该位点的甲基化状态直接相关(甲基化胞嘧啶的切割率是未甲基化的2倍)。
技术方法:
- FRAGMA:利用这种定量关系,通过支持向量机等模型,无需亚硫酸盐测序即可推断甲基化状态,并用于肝癌检测(AUC高达0.98)。
- FRAGMAXR:将FRAGMA原理扩展至基因组区域,通过分析核小体足迹的周期性振幅变化来检测癌症。在区分肝癌时(AUC=0.93),其性能优于传统的片段大小等指标,与FRAGMA结合后AUC可达0.98。该技术也适用于肺、乳腺和卵巢癌的检测。
2. 预测特异性组蛋白修饰
核心发现:特定的组蛋白修饰(如与活跃增强子相关的H3K27ac)与特征性的片段化模式(如特定长度片段的相对频率、末端基序)存在强关联。
技术方法:
- FRAGHA:通过分析这些片段特征(大小、末端基序),可以用线性回归或SVM模型来推断血浆中的H3K27ac水平,并成功用于区分肝癌与非肝癌病例(AUC = 0.93–0.97)。
3. 推断高阶染色质结构与转录特征
核心原理:活性基因的启动子区域通常存在核小体耗竭,因此可以通过cfDNA推断的核小体占据数据来反推基因表达水平。
应用与进展:
- 转录活性推断:TSS周围的片段大小多样性可以参数化基因表达水平,用于肺癌的检测和分型。
- 组织溯源:通过分析组织特异性表达基因的启动子区域的核小体足迹,可以推导cfDNA的组织来源。一项研究甚至能将低至0.3x测序深度的数据反卷积为490多种细胞类型,从而检测多种癌症。
- 探索肿瘤发生机制:高分辨率片段组学能揭示与癌症遗传背景相关的细微染色质结构变化。例如,在携带胚系TP53突变的李-佛美尼综合征患者中,即使未患癌,也观察到了癌症样的片段组学改变(如核小体相位减弱、染色质可及性增加),这表明该技术能捕捉癌前状态的早期生物学变化。
4. 多组学整合的临床实践案例
MERGE模型:一项肺癌检测研究展示了整合表观遗传-片段组学的威力。
- 方法:首先通过多种表观遗传学数据定义出“多表观遗传调控基因”区域,然后专门提取这些区域内的片段组学特征(大小、末端基序等)进行建模。
- 性能:该模型在检测肺癌(包括I期)和区分良恶性肺结节方面表现出色(AUC=0.94),并复现了与肺癌发生相关的关键基因,为靶向治疗提供了生物学见解。
5. 挑战与注意事项
- 尽管前景广阔,但基于片段组学的染色质组织分析仍面临一些挑战:
- 预分析偏差:cfDNA样本完整性、GC含量等可能影响结果,需考虑标准化。
- 数据稀疏性:在低测序覆盖度区域,数据稀疏可能带来分析困难。
- 成本与深度权衡:在保证足够测序深度以获取可靠片段特征与控制样本成本之间需要权衡。
新的片段组学机制的探索
当前片段组学研究正从表征已知核酸酶作用机制向系统性探索全新生物学过程深化。一方面,通过末端基序解析发现未知片段模式(如与氧化应激相关的F-profile VI)在肝癌检测中展现卓越效能,提示存在非酶促生物学机制。另一方面,全基因组关联研究在确认DNASE1L3等核心核酸酶作用的同时,更发现PANX1膜通道蛋白等非核酸酶基因与片段特征显著相关,揭示了细胞膜通道介导的细胞死亡等全新调控维度。这些发现突破了传统核酸酶中心论,为开发基于新机制的多维癌症诊断标志物体系开辟了道路。
基于cfDNA片段组学的癌症检测新计算方法
Algorithm | Input features | Machine learning algorithm | Cancers analyzed | Sequencing depth required for robustness | Performance summary |
|---|---|---|---|---|---|
DNA evaluation of fragments for early interception (DELFI) | Fragment size ratios and distributions in 5 Mb genomic bins | Gradient boosting | Breast, colorectal, lung, ovarian, pancreatic, gastric, bile duct | WGS 0.1× coverage | 57%–99% sensitivity at 98% specificity; overall AUC = 0.94 Tissue-of-origin prediction: 75% accuracy |
Genome-wide analysis of fragment ends (GALYFRE) | Information-weighted fraction of aberrant fragments (iwFAF) which parametrizes overall aberrancy of fragment end positions relative to previously defined recurrently protected regions; nucleotide frequencies 10 bp either side of fragment ends | Random forest | Breast, cholangiocarcinoma, glioblastoma, melanoma | 1 million reads per sample (WGS ∼0.05× coverage) | 45.5%–94.3% sensitivity at 95% specificity; overall AUC = 0.91; stage I AUC = 0.87 |
Examination of cfDNA with end selection (EXCEL) | Discordance metric (N-index): parametrizes difference between cfDNA-inferred and expected hematopoietic nucleosome positions in 5 Mb genomic bins | Gradient boosting | Bile duct, breast, colorectal, lung, ovarian, pancreatic | Not shown | 78.4%–100% sensitivity at 95% specificity across all cancer types and stages; overall AUC = 0.95 |
Instruction-tuned large language model for the assessment of cancer (iLLMAC) | Sentence containing tokens of 5′ 4-mer end motifs ordered by frequency; end motif “founder profiles” calculated by non-negative matrix factorization; motif diversity score | Large language model (LLaMA, 7B parameters) | Liver (hepatocellular), cervical, colorectal, esophageal, ovarian, head and neck, lung | 0.1 million reads per sample | 100% sensitivity at 66.7% specificity; overall AUC = 0.912 |
End motif inspection via transformer (EMIT) | Sentence containing tokens of 5′ 4-mer end motifs ordered by frequency | Large language model (up to 32 Mb parameter size) | Lung | 10 million reads per sample (WGS ∼0.5× coverage) | AUC = 0.962; identified several end motifs with higher attention scores in cancer |
基于片段末端基因组坐标的分析
核心思想:分析cfDNA片段末端在基因组上的特定位置分布,癌细胞死亡产生的cfDNA其末端会出现在不同于健康细胞的基因组区域。
代表性方法:
- GALYFRE:通过计算一个名为 iwFAF 的综合指标,来量化一个样本中cfDNA片段末端位置的总体“异常程度”。该指标与肿瘤负荷高度相关,即使在极低测序深度下也能有效检测早期癌症,并对胶质母细胞瘤等难检癌症表现优异。
- N-index:通过参数化cfDNA推断的核小体位置与预期的造血细胞核小体位置之间的差异来检测多种癌症,性能卓越。
2. 基于末端序列基序的“语言”分析
核心思想:将不同的末端序列基序(如"ACGT")视为“单词”,并将它们按频率组成“句子”,利用强大的大语言模型进行分析。
代表性方法:
- iLLMAC 和 EMIT:是两个基于Transformer架构的模型。它们通过分析这些末端基序“句子”,在多种癌症的检测中取得了很高的准确率。
- 技术优势:Transformer模型不仅能实现高性能,其注意力机制还能帮助识别对癌症检测最重要的关键末端基序,增强了模型的可解释性。
3. 前景与挑战
巨大潜力:文中特别提到DeepSeek等采用“专家混合”架构的模型,有望以更经济的计算成本实现顶尖性能,并便于在本地服务器部署,对医疗应用极具吸引力。
当前局限:
iLLMAC 直接套用自然语言模型,可能不适合基因组数据的底层逻辑。
EMIT 虽专为cfDNA设计,但为了适配模型而简化了输入数据(例如仅使用频率排序,丢弃了具体的频率数值和片段的原始基因组位置信息),这导致丢失了与染色质结构等相关的关键生物学信号。
总结:该领域正迅速从传统的生物信息分析向更复杂的AI驱动发现演进。未来的方向是开发能更好保留并利用cfDNA高维空间结构信息的新一代模型,以更全面地解读癌症在片段组学中留下的“指纹”。
多模态片段组学分析
多种片段组学特征的同步发展,推动了对多模态模型架构的探索,旨在整合各类片段组学生物标志物及其他维度信息。这些模型常采用"堆叠"算法——通过元学习器(如广义线性模型)将不同分类器的预测结果进行融合。
尽管多模态分析在理论上具有优势,但相较于结构更简单的模型,多模态片段组学模型对于极早期癌症的检测效能提升仍然有限。液体活检领域长期面临的核心挑战在于:早期病变细胞释放的cfDNA比例极低,且其在循环中被快速清除,导致肿瘤源性cfDNA信号的信噪比严重不足。单纯增加模型中的片段组学特征数量未必能带来性能的实质提升,甚至可能因不必要的模型复杂性产生反效果。例如在尤文肉瘤检测研究中,采用堆叠算法的综合模型性能并未显著优于仅使用区域片段覆盖度的单一特征模型,且在区分不同肉瘤类型时表现更差。这提示:特征选择必须基于疾病的特定生物学背景,未来需要深入研究早期癌变行为如何通过细胞死亡相关的基因组降解模式在cfDNA中呈现,这将为基于cfDNA的早期癌症诊断提供关键补充。
片段组学临床验证
癌症检测与组织溯源
DELFI(肺癌筛查):
- 研究设计:美国多中心大型筛查研究,旨在评估DELFI作为低剂量CT一线筛查工具的效能。
- 性能:在验证集中达到84%的灵敏度与53%的特异性,且在不同人群中都表现稳定。
- 公共卫生价值:模型预测,若将其用于CT前筛查,可在五年内预防超过7,600例肺癌死亡,显示出巨大的潜在公共卫生效益。
SPOT-MAS(多癌种筛查):
- 研究设计:在越南进行的多中心试验,针对40岁以上无症状人群,整合了片段组学、甲基化与拷贝数变异进行多癌种检测与组织溯源。
- 初步结果:在初步报告中,60% 的检测阳性者经确诊后,其癌症类型与模型预测的前两位组织来源之一相符。但由于大多数阴性受试者尚未完成12个月随访,其阴性预测值尚不明确。
PreCar(肝癌筛查):
- 研究设计:中国多中心研究,评估整合了5hmC、核小体足迹等多维特征的PreCar模型在高风险肝硬化人群中筛查肝癌的价值。
性能与挑战:
- **在模型构建阶段表现出色(灵敏度93.8%,特异性95.4%)。
- **在后续的横断面验证中,对早期肝癌的灵敏度降至51.3%。
关键启示:这一性能下降凸显了诊断模型必须使用能代表筛查人群特征的数据(尤其是足够的早期病例)进行训练,否则可能导致在真实场景中表现不佳。
2. 治疗监测与风险预测
范式转变:传统的治疗后监测(如微小残留病检测)严重依赖事先从肿瘤组织中找到的基因突变。
片段组学新优势:新兴的片段组学方法在此领域展现出独特前景,其最大优势在于无需事先进行肿瘤组织测序,仅通过分析血液中的cfDNA片段特征即可实现无创监测,为临床提供了更大便利。
总结:当前的多项大型临床试验初步验证了片段组学液体活检在癌症筛查、溯源和监测中的巨大潜力,标志着该技术正从研究走向临床。然而,未来仍需解决早期检测灵敏度、模型泛化能力以及长期预后价值的精确评估等关键挑战。
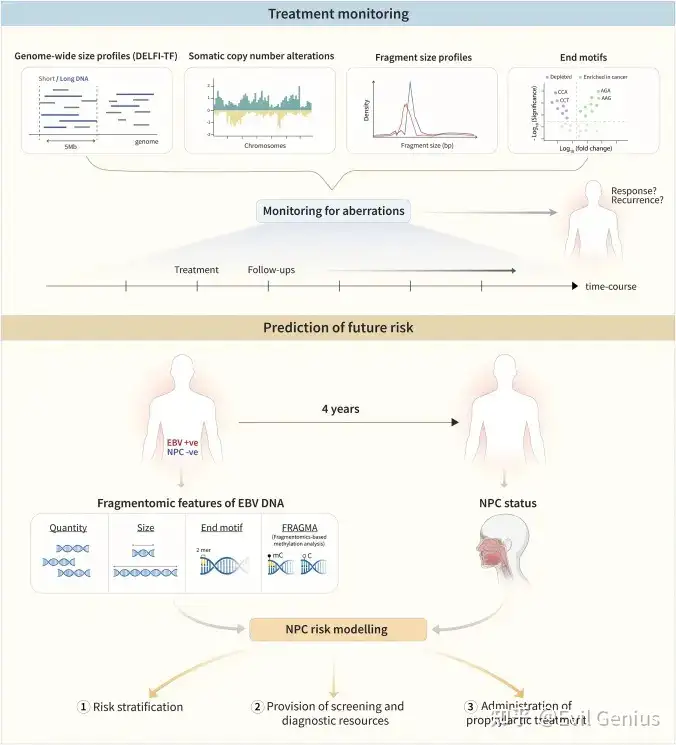
DELFI方法作为一种肿瘤类型无关的工具,在治疗监测和疗效预测领域展现出显著优势。研究表明,通过分析全基因组片段组学特征估算的DELFI肿瘤分数与传统基因突变检测结果高度一致,并能精准预测患者总生存期和无进展生存期。其动态变化率甚至在传统影像学无法区分部分缓解与疾病稳定时,就能有效鉴别治疗反应。与此同时,片段组学在鼻咽癌风险预测中也取得突破:通过整合血浆EBV DNA的片段大小、末端基序及FRAGMA甲基化推断等多维特征,可将未来4年内发病的相对风险提升至87.1倍。
尽管片段组学在癌症诊断中展现出巨大潜力,该领域仍面临诸多核心挑战:在机制层面,需明确不同细胞死亡模式对cfDNA产生的特异性贡献、解析组织释放cfDNA的选择性机制、量化核酸酶降解与细胞吞噬等清除途径的相对权重;在技术层面,需开发能准确区分肿瘤源性片段的高维特征、建立标准化流程以消除批次效应、并通过种群特异性GWAS研究识别真正的生物学调控因子;在临床转化层面,亟需开展大规模多中心试验系统比较不同算法的诊断效能,并验证特定片段特征组合在特定癌种与人群中的优势。解决这些关键问题将推动片段组学最终实现癌症液体活检的精准化革命。
生活很好,有你更好
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
评论
登录后参与评论
推荐阅读
目录

