内容补充--衰老细胞的单细胞空间检测
原创内容补充--衰老细胞的单细胞空间检测
原创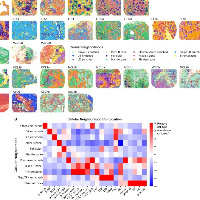
追风少年i
发布于 2026-03-14 19:53:59
发布于 2026-03-14 19:53:59
作者,Evil Genius
今天我们分享一个方法,对单细胞空间数据进行衰老细胞的检测。大家还知道常见的衰老细胞marker是什么么?
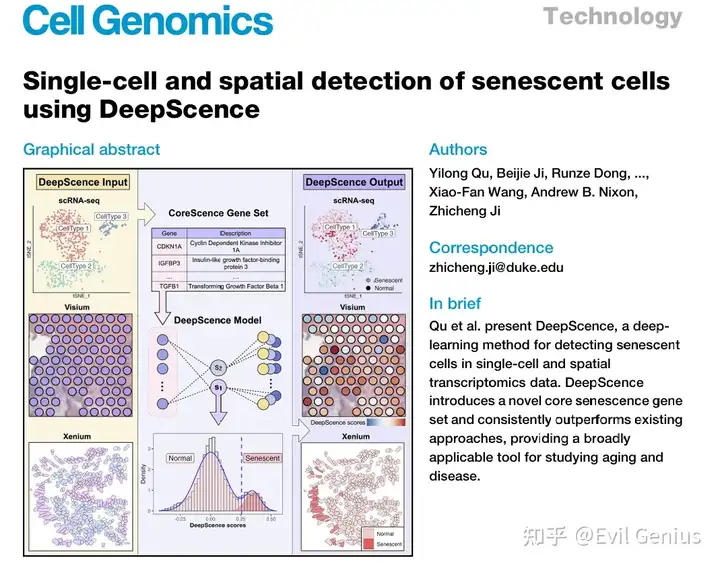

细胞衰老是一个基本的生物学过程,细胞在此过程中进入永久性的细胞周期停滞状态,丧失其增殖能力。这种状态主要作为一种防御机制,对抗在癌症中观察到的失控性细胞分裂。然而,随着时间的推移,衰老细胞(SnCs)的累积可能导致组织和器官的物理与功能完整性下降,在衰老过程中扮演重要角色。此外,SnCs还与多种年龄相关疾病的发生发展有关,例如骨关节炎、肺纤维化和阿尔茨海默病。 SnCs分泌促炎性蛋白质,这些蛋白质会改变细胞微环境,影响邻近细胞,并导致慢性炎症。
目前的衰老评判的基因集并不统一。
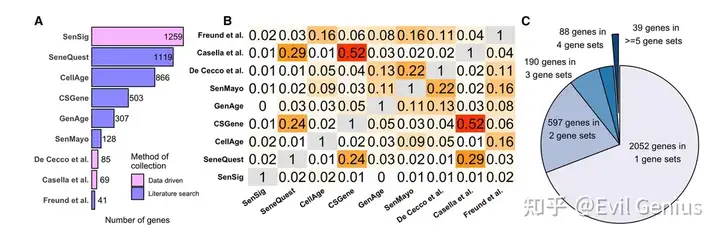

构建了一个名为“CoreScence”的核心衰老基因集。
构建方法:作者综合分析了之前提到的9个已发表的衰老相关基因集,选取了至少被其中5个基因集共同报道的基因,最终形成了一个由39个基因组成的核心基因集——CoreScence。其依据是,被多个独立研究共同认定的基因,更不容易受到单一研究纳入标准偏差的影响,因此与衰老过程的相关性更高。
基因构成:CoreScence包含了一些经典的细胞衰老标志基因,如 CDKN1A 和 CDKN2A。同时作者也指出,这些基因除了参与衰老,也可能涉及免疫或发育等其他功能,这一点将通过后续的DeepScence模型来解决。
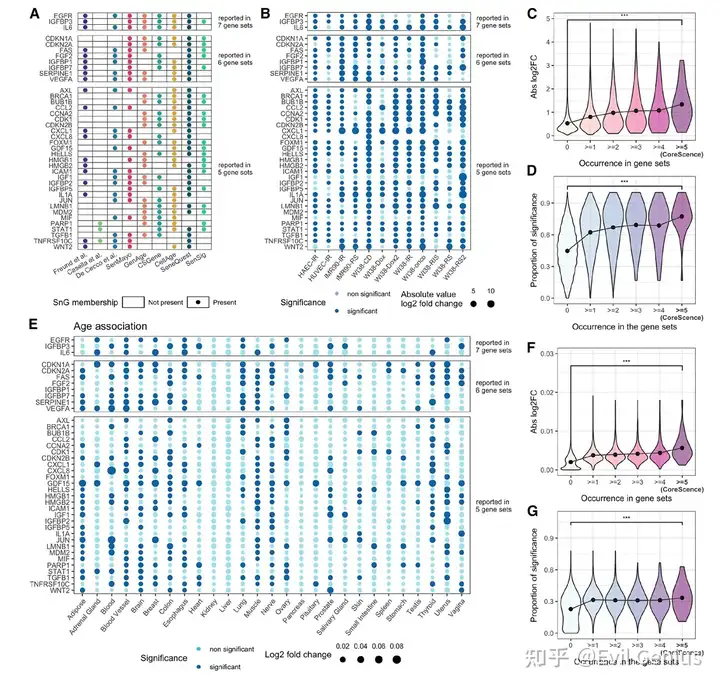
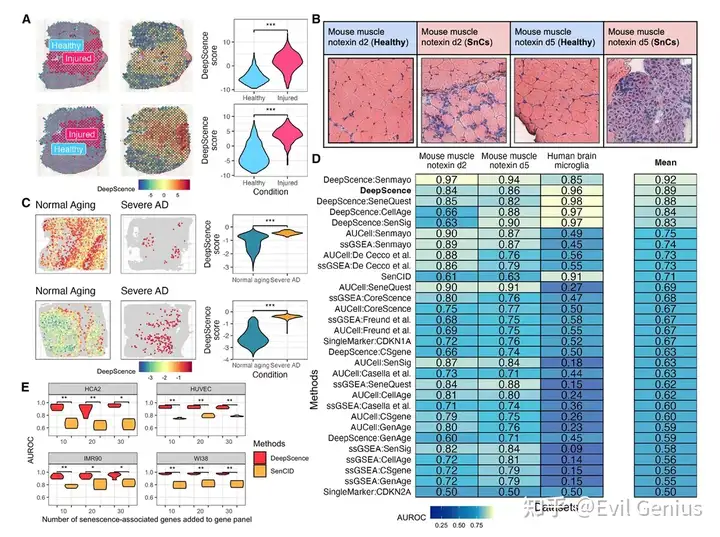
方法思路设计
主要输出:模型拟合完成后,捕捉衰老信息的神经元的值被定义为该细胞的衰老得分,这是一个连续的数值,代表细胞的衰老程度。
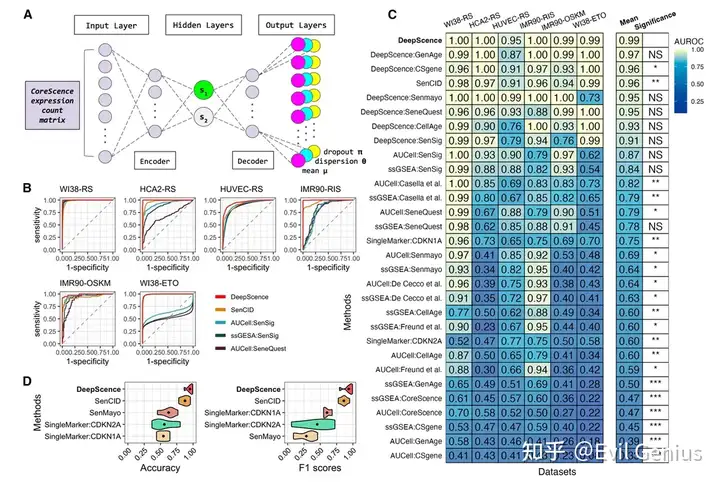
单细胞的衰老细胞检测
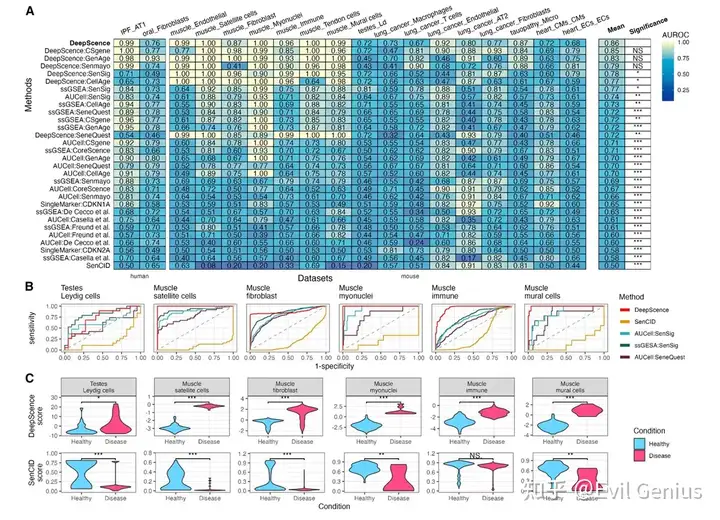
空间组学的衰老细胞检测
示例代码
import scanpy as sc
import numpy as np
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
from DeepScence.api import DeepScence
from dca.api import dca
adata = sc.read_h5ad("./mouse_muscle.h5ad")
sc.pp.filter_genes(adata, min_cells=1)
dca(adata)
#Run DeepScence and Visualize
# running mouse dataset
adata.obs["b"] = adata.obs["celltype"].astype(str)
adata = DeepScence(adata, binarize=False, species="mouse")
# Plot senescence score distributions comparing conditions
plt.figure(figsize=(12, 6))
sns.boxplot(
data=adata.obs,
x="celltype",
y="ds",
hue="stim",
palette=["#60efff", "#0061ff"]
)
plt.show()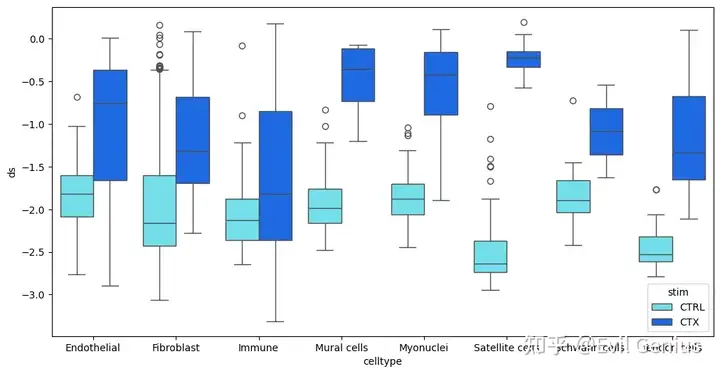
示例2
counts = pd.read_csv('./origin_matrix_GSE94980.txt', sep = '\t').T
adata = sc.AnnData(counts)
adata.obs['condition'] = [x.split('_')[-1]=='OSKM' for x in adata.obs_names]
adata.obs["SnC"] = (adata.obs["condition"]).astype(int)
adata.X = adata.X.astype(np.float32)
# denoise
sc.pp.filter_genes(adata, min_cells=1)
dca(adata)
adata = DeepScence(adata, binarize=True)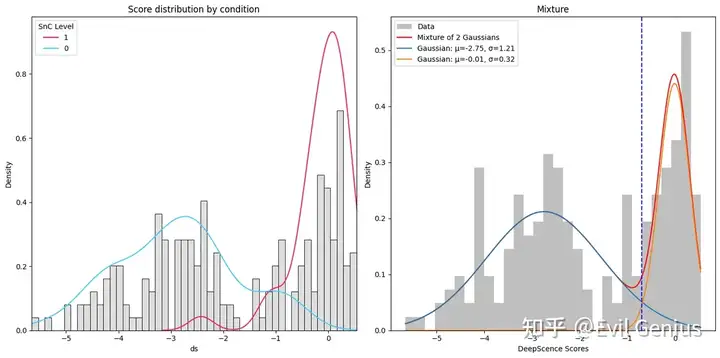
from sklearn.metrics import roc_auc_score, confusion_matrix, accuracy_score
import matplotlib.pyplot as plt
binary_numeric = adata.obs["binary"].map({"SnC": 1, "Normal": 0}).values
auroc = roc_auc_score(adata.obs["SnC"].values, adata.obs["ds"].values)
accuracy = accuracy_score(binary_numeric, adata.obs["SnC"])
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 4))
sns.boxplot(x=adata.obs["SnC"], y=adata.obs["ds"], ax=ax1)
ax1.set_title(f"Score by cell state (AUC={auroc:.3f})")
ax1.set_xlabel("True label (1: SnCs, 0: Normal)")
ax1.set_ylabel("DeepScence scores")
cm = confusion_matrix(binary_numeric, adata.obs["SnC"])
sns.heatmap(cm, annot=True, fmt='d', cmap='Blues', ax=ax2)
ax2.set_title(f"Binarization (Accuracy: {accuracy:.3f})")
ax2.set_xlabel("Predicted SnCs")
ax2.set_ylabel("True label")
plt.tight_layout()
plt.show()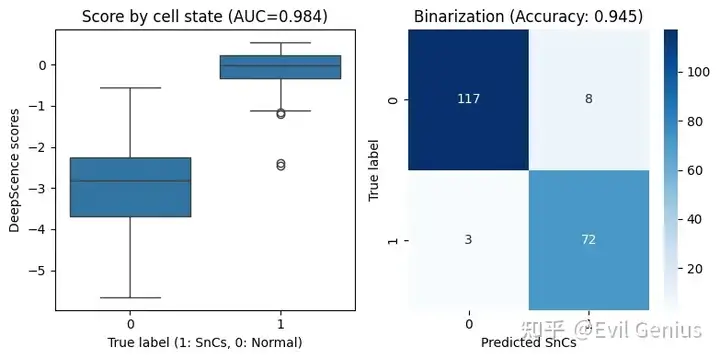
生活很好,有你更好。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
评论
登录后参与评论
推荐阅读
目录

