脚本更新--高精度空转分析差异富集的细胞生态位
原创脚本更新--高精度空转分析差异富集的细胞生态位
原创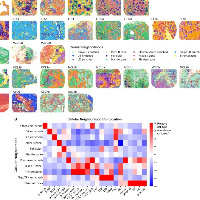
追风少年i
发布于 2026-03-06 10:06:42
发布于 2026-03-06 10:06:42
作者,Evil Genius
关于空间转录组的niche分析,相信大家都已经不陌生了,那么针对特定的组织结构,其区域显著富集的细胞niche,以及不同组(疾病间)显著差异的细胞niche,将会是分析的一个重点。
低精度如下面的分析
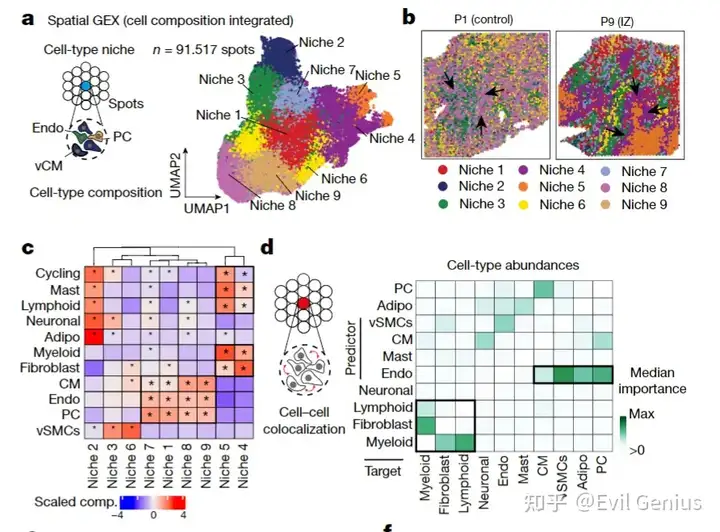
niche5明显聚集在P9样本。
高精度则需要更加的精细化
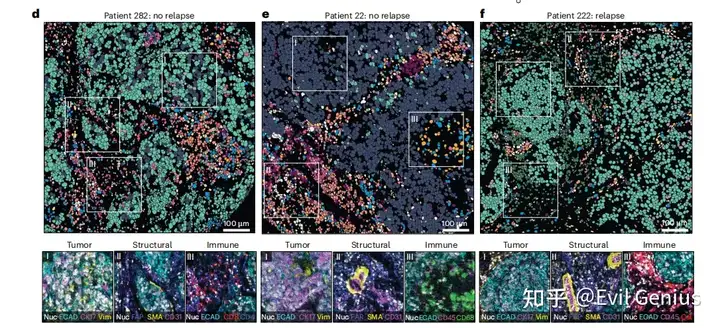
组织之间,不同区域之间,不同组之间的细胞结构需要更加小心的处理,发现其中niche的细微变化。
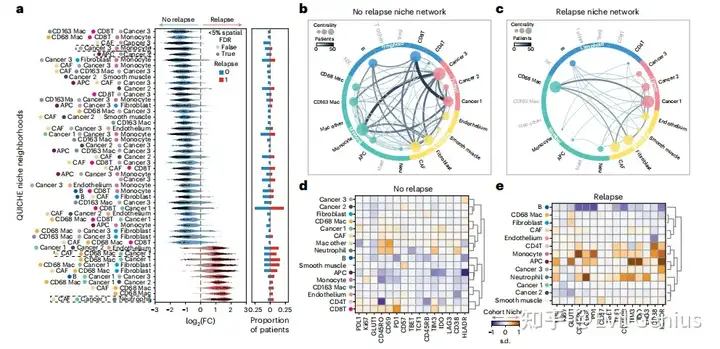
整体思路
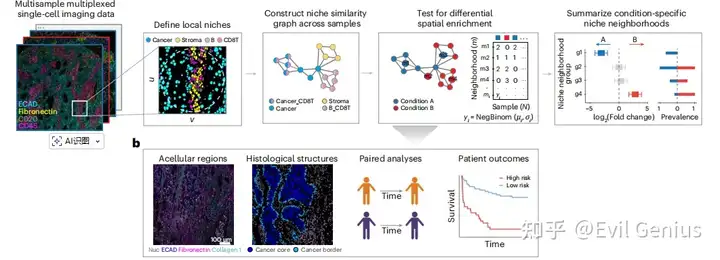
今天我们来分享一下这部分代码,前期的处理需要大家完成。
import os
import pandas as pd
import numpy as np
import quiche as qu
import scanpy as sc
import anndata
%reload_ext autoreload
%load_ext autoreload
%autoreload 2
%matplotlib inline
sc.set_figure_params(dpi = 400, dpi_save = 400, fontsize = 14)
####读取数据
adata = anndata.read_h5ad('多样本整合的h5ad')
adata.obs['Relapse'] = adata.obs['Relapse'].astype('int').astype('str')
adata.obs['Study'] = adata.obs['Study'].map(dict(zip(adata.obs['Study'].cat.categories,['A', 'B', 'C', 'D', 'E'])))
adata.raw = adata
adata.X = qu.pp.standardize(adata.X)
## Removes FOVs that are less than the downsampling (i.e. sketch) size
sketch_size = 1000
adata = qu.pp.filter_fovs(adata, 'Patient_ID', sketch_size)Perform differential spatial enrichment analysis with QUICHE
### initialize class
quiche_op = qu.tl.QUICHE(adata = adata, labels_key = 'cell_cluster', spatial_key = 'spatial', fov_key = 'fov', patient_key = 'Patient_ID', segmentation_label_key = 'label')
### step 1: compute spatial niches
## in this case we are computing a knn of 30 neighbors bounded by 200 pixel radius
quiche_op.compute_spatial_niches(radius = 200, n_neighbors = 30, min_cell_threshold = 3)
### step 2: perform distribution-focused downsampling by sampling niches from each patient according to the sketch size parameter
quiche_op.subsample(sketch_size = sketch_size, sketch_key = 'Patient_ID', n_jobs = 8)
### step 3: test for differential spatial enrichment across conditions.
## in this case, we are testing for differential enrichment across relapse groups while adjusting for medical center (i.e. study)
## for more details on model contrasts, see edgeR documentation
quiche_op.differential_enrichment(design = '~Study+Relapse', model_contrasts = 'Relapse1', k_sim = 100)
### step 4: annotate niche neighborhoods by the most abundant cell types
## in this case, annotate by the top 3
quiche_op.annotate_niches(nlargest = 3, annotation_scheme = 'neighborhood', annotation_key = 'quiche_niche_neighborhood')
###Access significant niche neighborhoods
## aggregates and returns 'significant' annotated niche neighborhoods
## in this case, we are filtering to ensure niches have median logFC > 1 or < -1 and have median Spatial FDR < 0.05
## feel free to intially relax your thresholds when examining results to ensure you're not missing information based on too strict of cutoffs
niche_scores = qu.tl.filter_niches(quiche_op,
thresholds = {'logFC': {'median' : [-1, 1]}, 'SpatialFDR': {'median' : 0.05}},
min_niche_count = 5,
annotation_key = 'quiche_niche_neighborhood')
niche_scores.sort_values(by = 'logFC').head()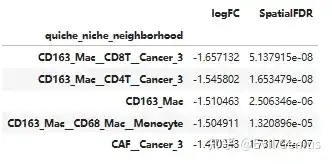
## computes summary statistics
niche_metadata = qu.tl.compute_niche_metadata(quiche_op,
niches = list(niche_scores.index),
annotation_key = 'quiche_niche_neighborhood',
patient_key = 'Patient_ID',
condition_key = 'Relapse',
niche_threshold = 0,
condition_type = 'binary',
metrics = ['logFC', 'SpatialFDR', 'PValue'])
niche_metadata.head()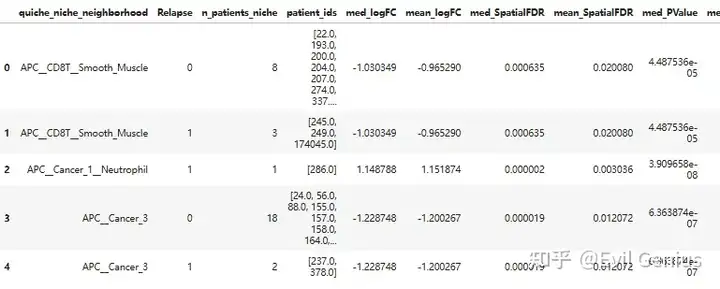
Plot significant niche neighborhoods of interest
qu.pl.beeswarm_proportion(quiche_op,
niche_metadata = niche_metadata,
niches = niche_scores.index,
xlim = [-3,3],
xlim_proportion = [-0.3, 0.3],
logfc_key = 'logFC',
pvalue_key = 'SpatialFDR',
annotation_key = 'quiche_niche_neighborhood',
condition_key = 'Relapse',
figsize = (6, 12),
fontsize = 10,
colors_dict = {'0': '#377eb8', '1': '#e41a1c'},
save_directory = None,
filename_save = None)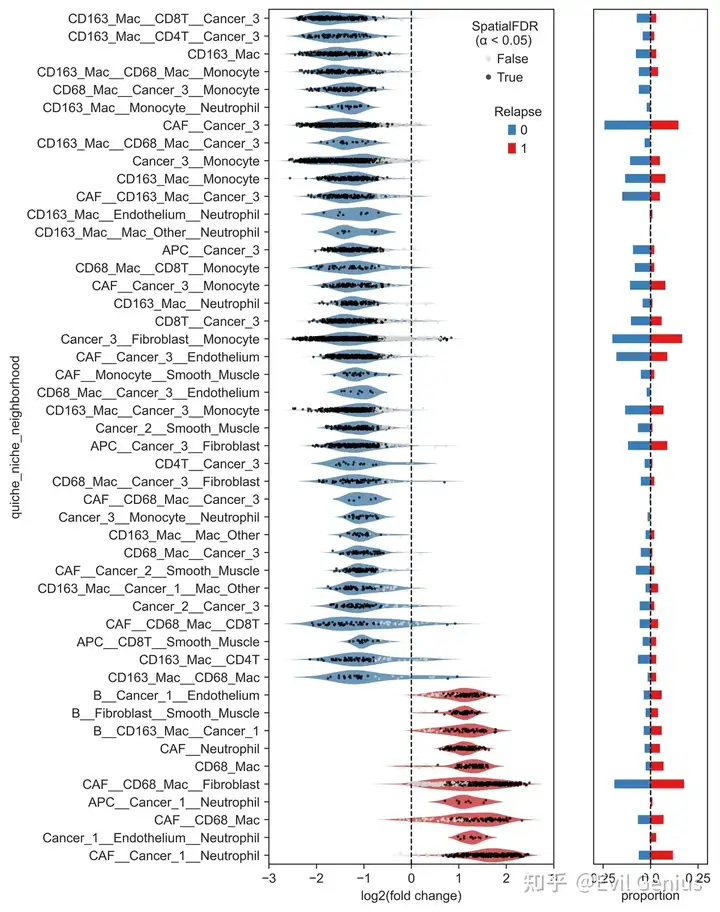
niche网络
lineage_dict = {'APC':'myeloid', 'B':'lymphoid', 'CAF': 'structural', 'CD4T': 'lymphoid', 'CD8T': 'lymphoid',
'CD68_Mac': 'myeloid', 'CD163_Mac': 'myeloid', 'Cancer_1': 'tumor', 'Cancer_2': 'tumor', 'Cancer_3': 'tumor',
'Endothelium':'structural', 'Fibroblast': 'structural', 'Mac_Other': 'myeloid', 'Mast':'myeloid', 'Monocyte':'myeloid',
'NK':'lymphoid', 'Neutrophil':'myeloid', 'Smooth_Muscle':'structural', 'T_Other':'lymphoid', 'Treg':'lymphoid'}
cell_ordering = ['Cancer_1', 'Cancer_2', 'Cancer_3', 'CD4T', 'CD8T', 'Treg', 'T_Other', 'B',
'NK', 'CD68_Mac', 'CD163_Mac', 'Mac_Other', 'Monocyte', 'APC','Mast', 'Neutrophil',
'CAF', 'Fibroblast', 'Smooth_Muscle', 'Endothelium']
##niche network for patients that did not relapse, ie logFC < 0
niche_metadata_neg = niche_metadata[(niche_metadata['mean_logFC'] < 0) & (niche_metadata['n_patients_niche'] > 1)]
G1 = qu.tl.compute_niche_network(niche_df = niche_metadata_neg,
colors_dict = colors_dict,
lineage_dict = lineage_dict,
annotation_key = 'quiche_niche_neighborhood')
qu.pl.plot_niche_network_donut(G = G1,
figsize=(6, 6),
node_order = cell_ordering,
centrality_measure = 'eigenvector',
colors_dict = colors_dict,
lineage_dict=lineage_dict,
donut_radius_inner = 1.15,
donut_radius_outer = 1.25,
edge_cmap = 'bone_r',
edge_label = 'Patients',
save_directory=None,
filename_save=None)
##niche network for patients that relapsed, ie logFC > 0
niche_metadata_pos = niche_metadata[(niche_metadata['mean_logFC'] > 0) & (niche_metadata['n_patients_niche'] > 1)]
G2 = qu.tl.compute_niche_network(niche_df = niche_metadata_pos,
colors_dict = colors_dict,
lineage_dict = lineage_dict,
annotation_key = 'quiche_niche_neighborhood')
qu.pl.plot_niche_network_donut(G = G2,
figsize=(6, 6),
node_order = cell_ordering,
centrality_measure = 'eigenvector',
colors_dict = colors_dict,
lineage_dict=lineage_dict,
donut_radius_inner = 1.15,
donut_radius_outer = 1.25,
edge_cmap = 'bone_r',
edge_label = 'Patients',
save_directory=None,
filename_save=None)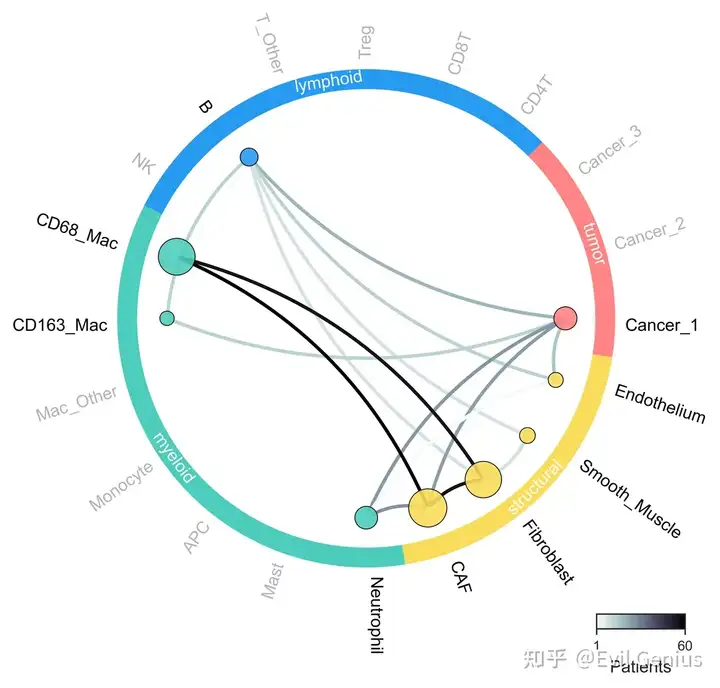

niche的空间展示
niche = 'CD8T__Cancer_1'
seg_dir = './'
qu.pl.plot_niches(quiche_op,
niche = niche,
fovs = ['TMA44_R14C7'],
segmentation_directory = seg_dir,
save_directory = os.path.join('figures', 'quiche_niche_neighborhood'),
fov_key = "fov",
segmentation_label_key = "label",
labels_key='cell_cluster',
annotation_key = "quiche_niche_neighborhood",
seg_suffix = "_whole_cell.tiff",
colors_dict = {'Cancer_1': '#66cdaa', 'CD8T': '#ff5666'},
figsize = (6, 6),
dpi = 300)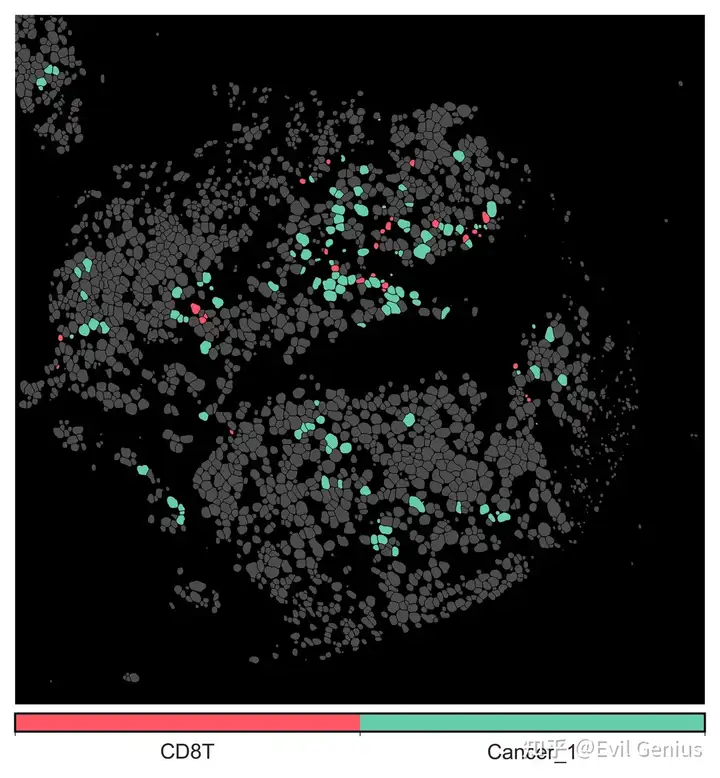
qu.pl.plot_niche_scores(quiche_op,
niche = niche,
fovs = ['TMA44_R14C7'],
segmentation_directory = seg_dir,
save_directory = os.path.join('figures', 'logFC'),
fov_key = "fov",
segmentation_label_key = "label",
labels_key='cell_cluster',
annotation_key = "quiche_niche_neighborhood",
metric = "logFC",
seg_suffix = "_whole_cell.tiff",
vmin = None,
vmax = None,
cmap = "vlag",
figsize = (6, 6),
dpi = 300)
qu.pl.plot_niche_scores(quiche_op,
niche = niche,
fovs = ['TMA44_R14C7'],
segmentation_directory = seg_dir,
save_directory = os.path.join('figures', '-log10(SpatialFDR)'),
fov_key = "fov",
segmentation_label_key = "label",
labels_key='cell_cluster',
annotation_key = "quiche_niche_neighborhood",
metric = "-log10(SpatialFDR)",
seg_suffix = "_whole_cell.tiff",
vmin = None,
vmax = None,
cmap = "Reds",
figsize = (6, 6),
dpi = 300)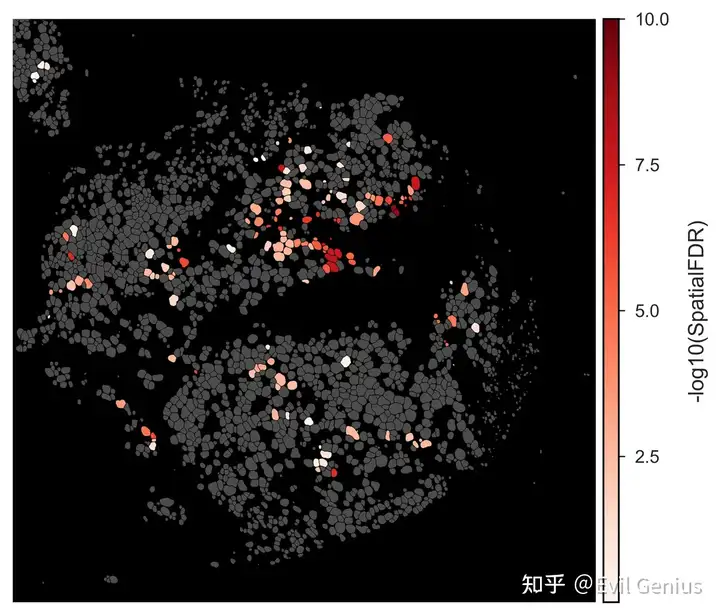

差异niche分析
functional_markers = ['PDL1', 'Ki67', 'GLUT1', 'CD45RO', 'CD69', 'PD1', 'CD57', 'TBET', 'TCF1', 'CD45RB', 'TIM3', 'IDO', 'LAG3', 'CD38', 'HLADR']
quiche_op.compute_functional_expression(niches=niche_scores.index,
annotation_key = 'quiche_niche_neighborhood',
min_cell_threshold = 3,
foldchange_key = 'logFC',
markers = functional_markers,
n_jobs = 8)
qu.pl.plot_differential_expression(quiche_op,
niches = np.unique(niche_metadata_neg.quiche_niche_neighborhood),
annotation_key = 'quiche_niche_neighborhood',
labels_key = 'cell_cluster',
markers = functional_markers,
figsize = (5.25, 4.5),
cmap = 'vlag',
save_directory = 'figures',
filename_save = 'matrixplot')
qu.pl.plot_differential_expression(quiche_op,
niches = np.unique(niche_metadata_pos.quiche_niche_neighborhood),
annotation_key = 'quiche_niche_neighborhood',
labels_key = 'cell_cluster',
markers = functional_markers,
figsize = (5.25, 4),
cmap = 'vlag',
vmin = -1,
vmax = 1,
vcenter = 0,
dendrogram = True,
save_directory = 'figures',
filename_save = 'matrixplot')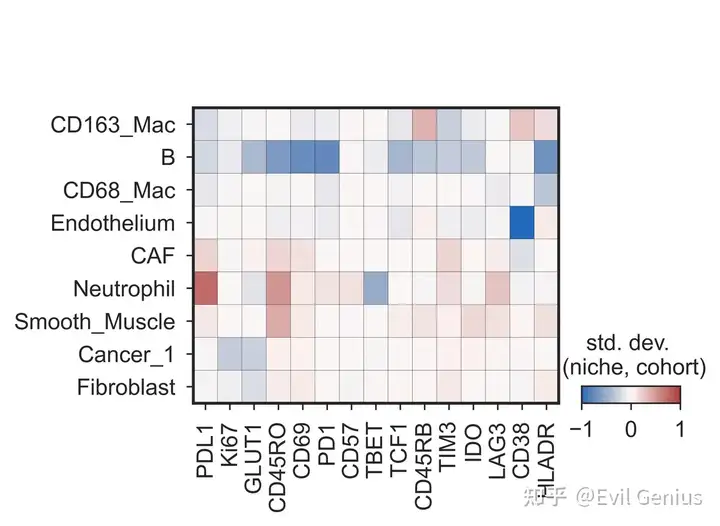

生活很好,有你更好。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
评论
登录后参与评论
推荐阅读
目录

