流程升级---原位捕获数据的无分割分析(Stereo-seq、HD)
原创流程升级---原位捕获数据的无分割分析(Stereo-seq、HD)
原创
追风少年i
发布于 2024-11-11 14:17:00
发布于 2024-11-11 14:17:00
作者,Evil Genius
高分辨率的空间转录组,最好做细胞分割,一般不要用一刀切的bin划分方法,但是我们如果没有条件做细胞分割,就需要改进一下一刀切的bin分割方法。
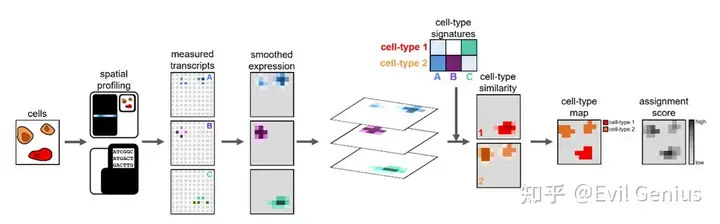
尽可能恢复空间转录组的细胞精度矩阵。
空间转录组学保留了空间信息,能够识别细胞niche,并研究不同细胞类型在其原生微环境中的相互作用。这种空间背景对于理解复杂的生物过程(如发育、疾病进展和对治疗的反应)至关重要。
SRT方法可以大致分为两类。第一类通常被称为基于成像的方法通过原位杂交(ISH)或原位测序(ISS)为选定数量的靶基因提供单分子分辨率,通常在100s-1000s的范围内。第二类被称为原位捕获方法,并在测序前将空间条形码整合到转录物上,允许整个转录组覆盖,但空间分辨率有限(例如,Visium的spot间距为100 um)。原位捕获方法的低空间分辨率使单细胞的空间分析复杂化,需要反卷积、插补和/或整合外部单细胞转录组学资源。然而,空间转录组学的最新进展通过Stereo-seq、Seq-Scope、Open-ST和Nova-ST等定序方法提供了纳米分辨率的全转录组图谱。这些高分辨率技术提供了独特的优势,例如在亚细胞水平上解析转录组全表达的能力,在某些情况下在亚微米范围内。
目前的分析挑战:高分辨率方法产生的数据量巨大,需要强大且可扩展的计算工具进行高效的数据处理和分析。精确的细胞分割仍然难以实现。
改进后的分析框架
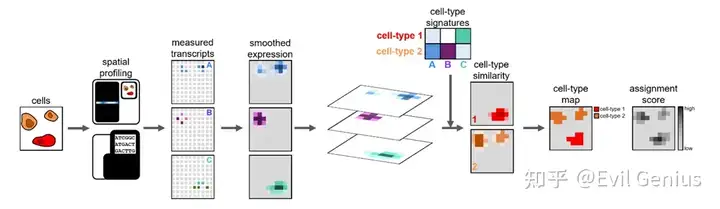
核心分析原理:通过应用核密度估计(KDE)将空间基因表达建模为密度,然后可用于整个组织的稳健和准确的细胞类型分配。计算步骤大致可分为读取数据、质量控制、数据预处理、空间基因表达建模、生成或获取特定细胞类型的基因表达模式,以及将细胞类型分配到像素生成细胞类型映射。
算法的数据验证(模拟数据)
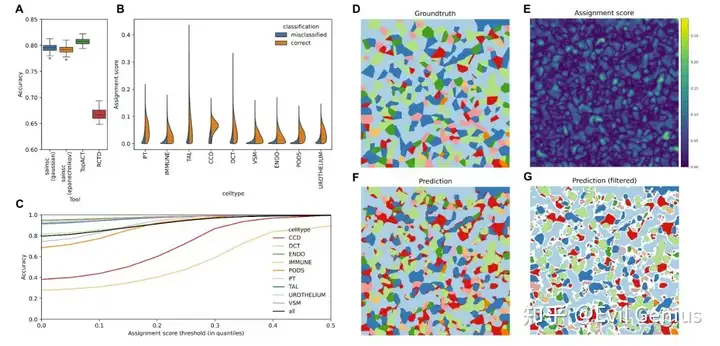
华大小鼠数据(性能验证)
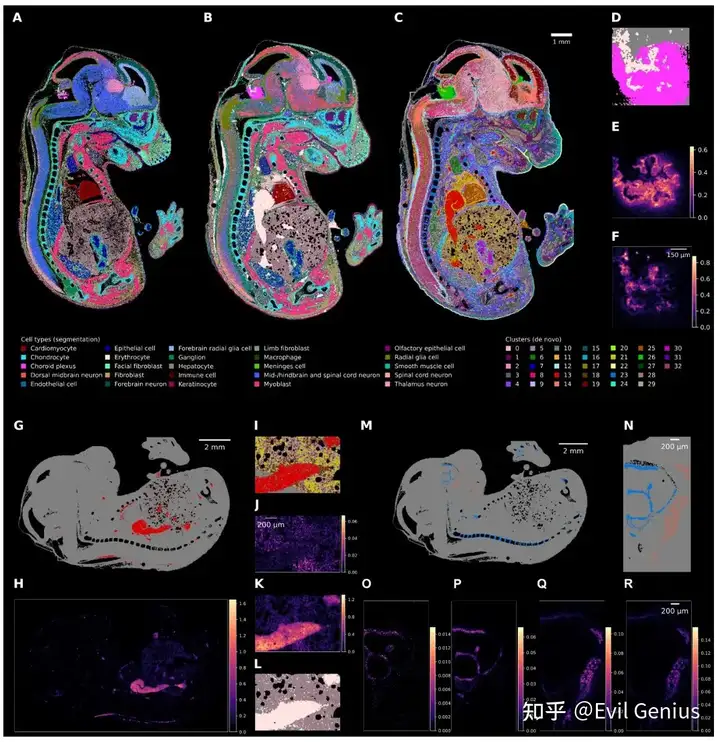
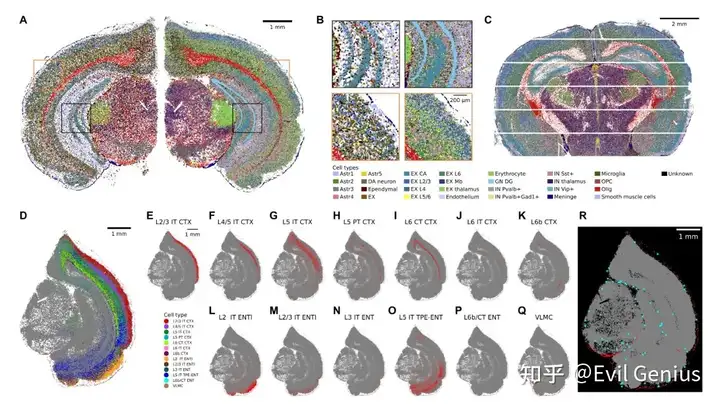
华大数据小鼠脑(性能验证)
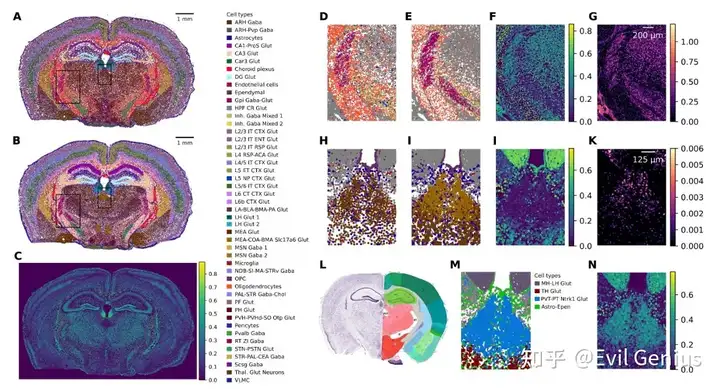
Xenium数据
最后看看示例代码(华大数据)
pip install sainsc
###conda install bioconda::sainsc
from pathlib import Path
import numpy as np
import pandas as pd
from sainsc.io import read_StereoSeq
####加载数据
embryo = read_StereoSeq(stereo_seq_file)
embryo.calculate_total_mRNA()
_ = embryo.plot_genecount(im_kwargs={"vmax": 5})
embryo.counts.crop((5_000, 33_000), (9_000, None))
embryo.calculate_total_mRNA()
_ = embryo.plot_genecount(im_kwargs={"vmax": 5})
from PIL import Image
# export total mRNA as png
u8 = np.uint8
Image.fromarray(
np.clip(embryo.total_mRNA, np.iinfo(u8).min, np.iinfo(u8).max).astype(u8)
).save("embryo_counts.png")
import matplotlib.pyplot as plt
from sainsc.datasets import fetch_embryo_mask
# avoid the DecompressionBombError
Image.MAX_IMAGE_PIXELS = None
mask_file = fetch_embryo_mask()
embryo_mask = np.array(Image.open(mask_file), dtype=bool)
_ = plt.imshow(embryo_mask.T, origin="lower")
embryo.counts.filter_mask(embryo_mask)
embryo.counts.filter_genes_by_count(1)
embryo.calculate_total_mRNA()
_ = embryo.plot_genecount(im_kwargs={"vmax": 5})
###核密度
embryo.gaussian_kernel(4, unit="um")
embryo.calculate_total_mRNA_KDE()
_ = embryo.plot_KDE()
embryo.filter_background(0.9)
_ = embryo.plot_KDE(remove_background=True)
####Cell-type assignment
from sainsc.datasets import fetch_embryo_signatures
signatures = fetch_embryo_signatures()
signatures.iloc[:, :4].head()
embryo.assign_celltype(signatures)
import colorcet as cc
import seaborn as sns
cmap = dict(
zip(embryo.celltypes, sns.color_palette(cc.glasbey, n_colors=len(embryo.celltypes)))
)
_ = embryo.plot_celltype_map(cmap=cmap)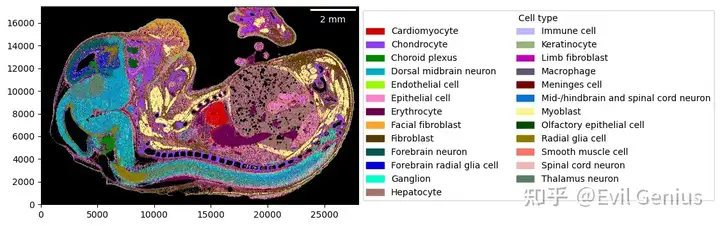
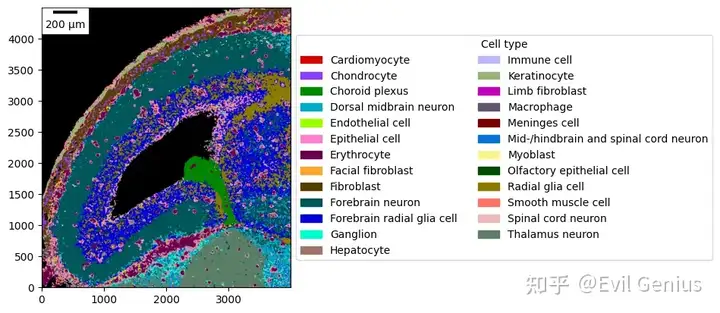
_ = embryo.plot_celltype_map(
crop=((2_000, 6_000), (10_500, 15_000)),
cmap=cmap,
scalebar_kwargs={"location": "upper left"},
)
_ = embryo.plot_assignment_score()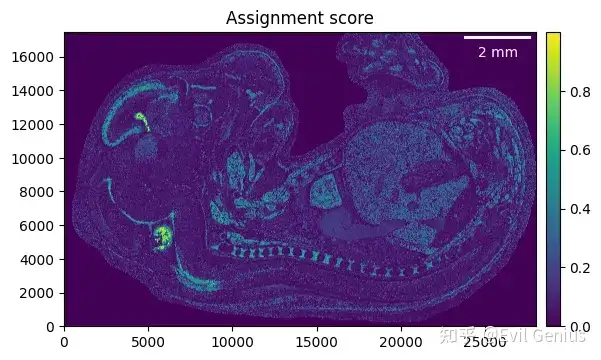
示例代码在GitHub - HiDiHlabs/sainsc: Segmentation-free Analysis of In Situ Capture data
生活很好,有你更好
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
评论
登录后参与评论
推荐阅读
目录

