方法迭代----基于STAMP的空间转录组学可解释的空间aware降维(Stereo-seq)
原创方法迭代----基于STAMP的空间转录组学可解释的空间aware降维(Stereo-seq)
原创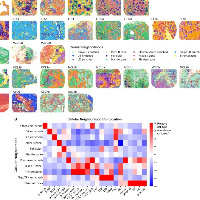
追风少年i
发布于 2024-10-16 09:56:23
发布于 2024-10-16 09:56:23
作者,Evil Genius
时间飞快~~~
如今都是高通量的时代了,无论是Spatial HD,还是华大的Stereo-seq,都是以超高维的数据量著称。单个样本的spot数量就已经达到了十几万(8 um),如果做多样本整合,那么就要面临几十万甚至上百万的spot,R的性能很难处理这个规模的数据量,即使是python,也需要GPU,可见,随着数据量的快速提升,我们需要进行方法的迭代,数据量急剧上升的同时,也要保证对空间数据分析的准确性。
今日参考文献,新加坡的华人。

知识积累
- 空间转录组学产生具有空间背景的高维基因表达测量。获得这些数据的具有生物学意义的低维表示对于有效解释和下游分析至关重要。
- 空间感知分析需要结合基因表达和空间信息
- 经典的降维方法,如主成分分析(PCA)、非负矩阵分解(NMF)和潜在狄利克雷分配(LDA)经常用于单细胞分析。
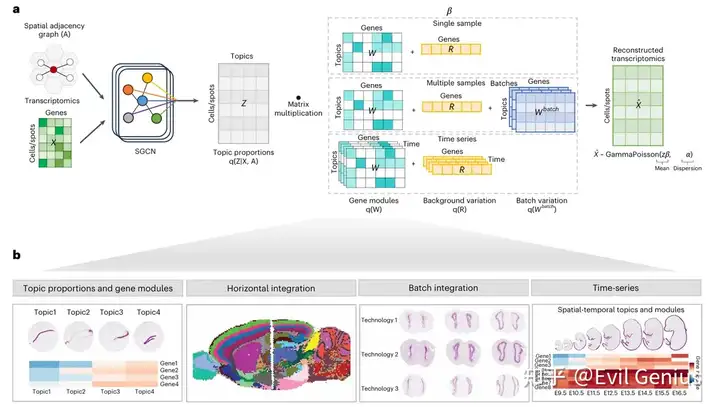
Workflow of STAMP
核心就是需要同时考虑基因表达基因和空间邻域,也要可以处理超高通量的数据。
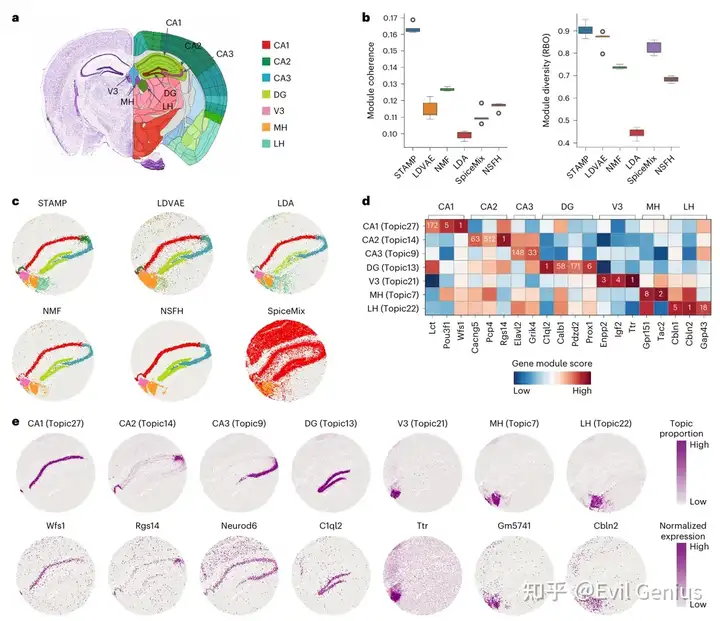
STAMP在小鼠海马空间域的运用(Slide-seq V2)
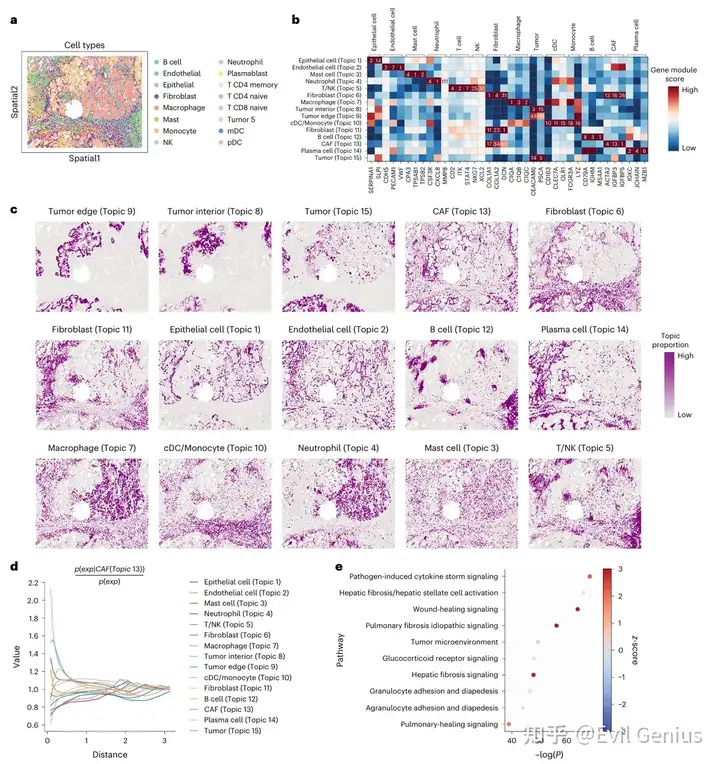
STAMP揭示了癌症相关的成纤维细胞( CosMx SMI,NSCLC)
STAMP整合小鼠前、后脑切片(10x Genomics Visium )

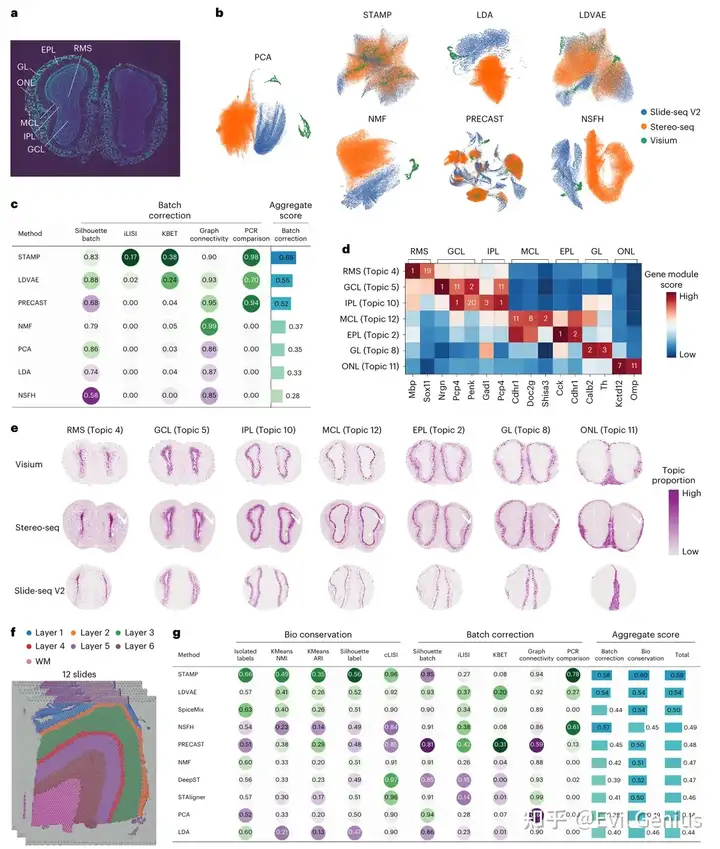
STAMP识别跨不同技术的空间数据(Slide-seq V2, Stereo-seq and 10x Genomics Visium)
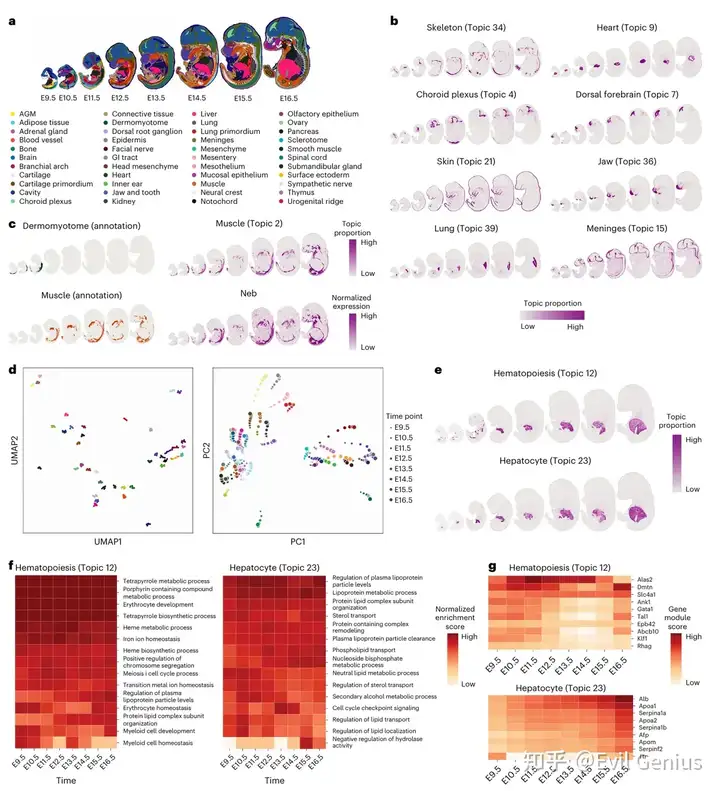
STAMP揭示了胚胎发育的时空主题(Stereo-seq)
示例代码在scTM (jinmiaochenlab.github.io)
我们来简单看一下
pip install scTM
import scanpy as sc
import numpy as np
import sctm
import squidpy as sq
import sklearn
import seaborn as sns
%load_ext autoreload
%autoreload 2
adata = sc.read_h5ad(
"../../../../STAMP/Reproducibility/ProcessedData/Visium_Mousebrain/adata_benchmark.h5ad"
)
adata.var_names_make_unique()
sc.pp.calculate_qc_metrics(adata, inplace=True)
adata.layers["counts"] = adata.X.copy()
sq.gr.spatial_neighbors(adata, library_key="data")
sctm.seed.seed_everything(5)
model = sctm.stamp.STAMP(
adata[:, adata.var.highly_variable],
n_topics=n_topics,
layer = "counts",
categorical_covariate_keys = ["data"],
gene_likelihood = "nb")
model.train(learning_rate = 0.01, min_epochs = 200)
topic_prop = model.get_cell_by_topic()
beta = model.get_feature_by_topic()
for i in topic_prop.columns:
adata.obs[i] = topic_prop[i]
topic_prop = model.get_cell_by_topic()
beta = model.get_feature_by_topic(pseudocount = 0.0)
for i in topic_prop.columns:
adata.obs[i] = topic_prop[i]
top_genes = []
for i in topic_prop.columns:
top_genes += beta.nlargest(1, i).index.tolist()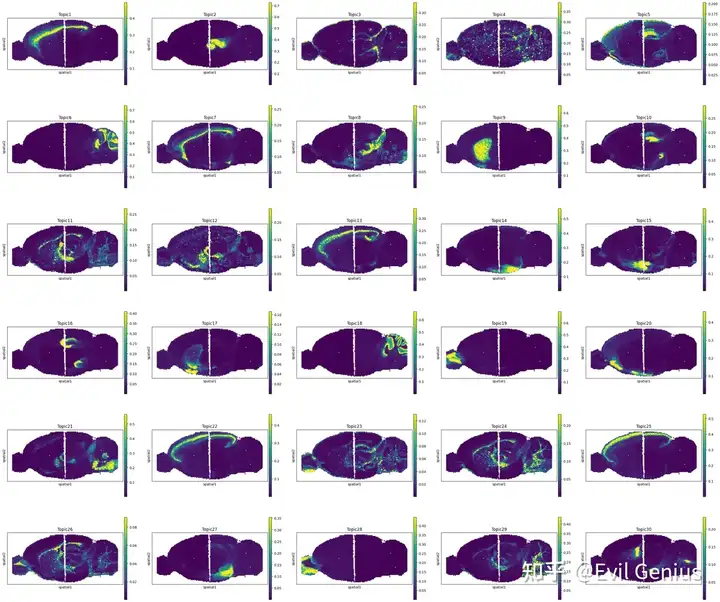
sctm.pl.spatial(adata, color=top_genes, ncols=5, size=40, vmax="p99")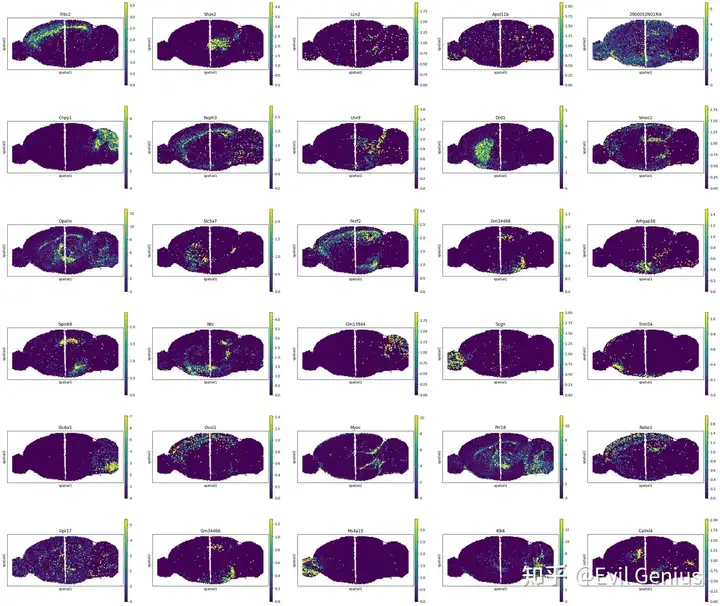 
topics = ["Topic1", "Topic7", "Topic13", "Topic22", "Topic25", "Topic10", "Topic16"]
fig = sctm.pl.plot_spatial(
adata,
topic_prop.loc[:, topics],
spot_size=7,
display_zeros=True,
axis_y_flipped=True,
)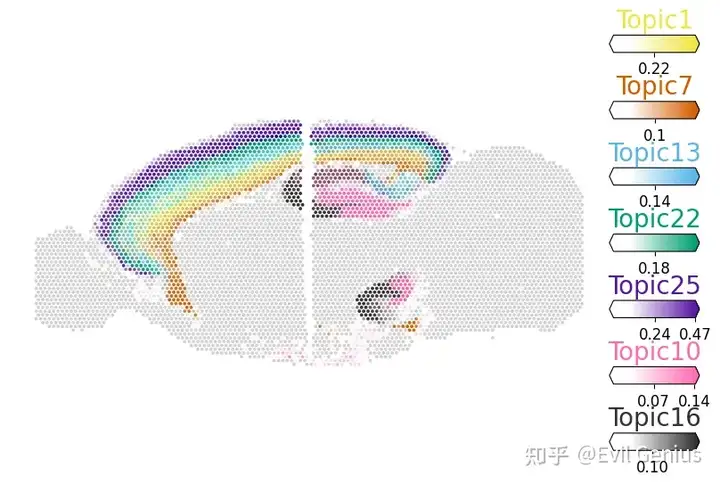
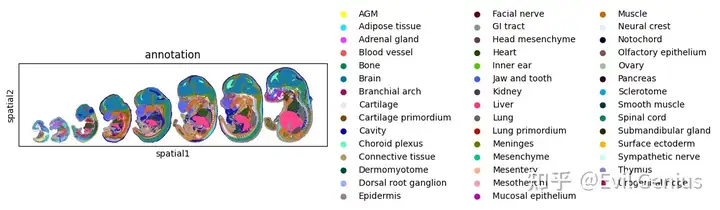
放一张华大的分析图
生活很好,有你更好
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
评论
登录后参与评论
推荐阅读
目录

