带光滑线的Python散点图
带光滑线的Python散点图
提问于 2018-08-24 12:24:51
我有如下所示的数据(经过大量预处理后获得)
请找数据
d = {'token': {361: '180816_031', 119: '180816_031', 101: '180816_031', 135: '180816_031', 292: '180816_031',
133: '180816_031', 99: '180816_031', 270: '180816_031', 19: '180816_031', 382: '180816_031',
414: '180816_031', 267: '180816_031', 218: '180816_031', 398: '180816_031', 287: '180816_031',
155: '180816_031', 392: '180816_031', 265: '180816_031', 239: '180816_031', 237: '180816_031'},
'station': {361: 'deneb', 119: 'callisto', 101: 'callisto', 135: 'callisto', 292: 'callisto', 133: 'deneb',
99: 'callisto', 270: 'callisto', 19: 'deneb', 382: 'callisto', 414: 'deneb', 267: 'callisto',
218: 'deneb', 398: 'callisto', 287: 'deneb', 155: 'deneb', 392: 'deneb', 265: 'callisto',
239: 'callisto', 237: 'callisto'},
'cycle_number': {361: 'cycle09', 119: 'cycle06', 101: 'cycle04', 135: 'cycle01', 292: 'cycle04', 133: 'cycle05',
99: 'cycle06', 270: 'cycle07', 19: 'cycle04', 382: 'cycle08', 414: 'cycle04', 267: 'cycle10',
218: 'cycle07', 398: 'cycle08', 287: 'cycle09', 155: 'cycle08', 392: 'cycle06', 265: 'cycle02',
239: 'cycle09', 237: 'cycle07'},
'variable': {361: 'adj_high_quality_reads', 119: 'short_pass', 101: 'short_pass', 135: 'cell_mask_bilayers_sum',
292: 'adj_active_polymerase', 133: 'cell_mask_bilayers_sum', 99: 'short_pass',
270: 'adj_active_polymerase', 19: 'Unnamed: 0', 382: 'adj_high_quality_reads',
414: 'num_align_high_quality_reads', 267: 'adj_active_polymerase', 218: 'adj_single_pores',
398: 'num_align_high_quality_reads', 287: 'adj_active_polymerase', 155: 'cell_mask_bilayers_sum',
392: 'num_align_high_quality_reads', 265: 'adj_active_polymerase', 239: 'adj_single_pores',
237: 'adj_single_pores'},
'value': {361: 99704.0, 119: 2072785.0, 101: 2061059.0, 135: 1682208.0, 292: 675306.0, 133: 1714292.0,
99: 2072785.0, 270: 687988.0, 19: 19.0, 382: np.nan, 414: 285176.0, 267: 86914.0, 218: 948971.0,
398: 405196.0, 287: 137926.0, 155: 1830032.0, 392: 480081.0, 265: 951689.0, 239: 681452.0,
237: 882671.0}}数据:
token station cycle_number variable \
19 180816_031 deneb cycle04 Unnamed: 0
99 180816_031 callisto cycle06 short_pass
101 180816_031 callisto cycle04 short_pass
119 180816_031 callisto cycle06 short_pass
133 180816_031 deneb cycle05 cell_mask_bilayers_sum
135 180816_031 callisto cycle01 cell_mask_bilayers_sum
155 180816_031 deneb cycle08 cell_mask_bilayers_sum
218 180816_031 deneb cycle07 adj_single_pores
237 180816_031 callisto cycle07 adj_single_pores
239 180816_031 callisto cycle09 adj_single_pores
265 180816_031 callisto cycle02 adj_active_polymerase
267 180816_031 callisto cycle10 adj_active_polymerase
270 180816_031 callisto cycle07 adj_active_polymerase
287 180816_031 deneb cycle09 adj_active_polymerase
292 180816_031 callisto cycle04 adj_active_polymerase
361 180816_031 deneb cycle09 adj_high_quality_reads
382 180816_031 callisto cycle08 adj_high_quality_reads
392 180816_031 deneb cycle06 num_align_high_quality_reads
398 180816_031 callisto cycle08 num_align_high_quality_reads
414 180816_031 deneb cycle04 num_align_high_quality_reads
value
19 19.0
99 2072785.0
101 2061059.0
119 2072785.0
133 1714292.0
135 1682208.0
155 1830032.0
218 948971.0
237 882671.0
239 681452.0
265 951689.0
267 86914.0
270 687988.0
287 137926.0
292 675306.0
361 99704.0
382 NaN
392 480081.0
398 405196.0
414 285176.0 我试图用光滑的线条来创建散点图。
fig,ax = plt.subplots()
fig.set_size_inches(16,4)
#to get different colors for each of the `variable` value assign the variable to hue
g2=sns.lmplot(x='cycle_number',y='value',data=df, hue='variable', size=4, aspect=5)此代码只为散点图提供一个值,但我的预期输出如下所示
预期输出:
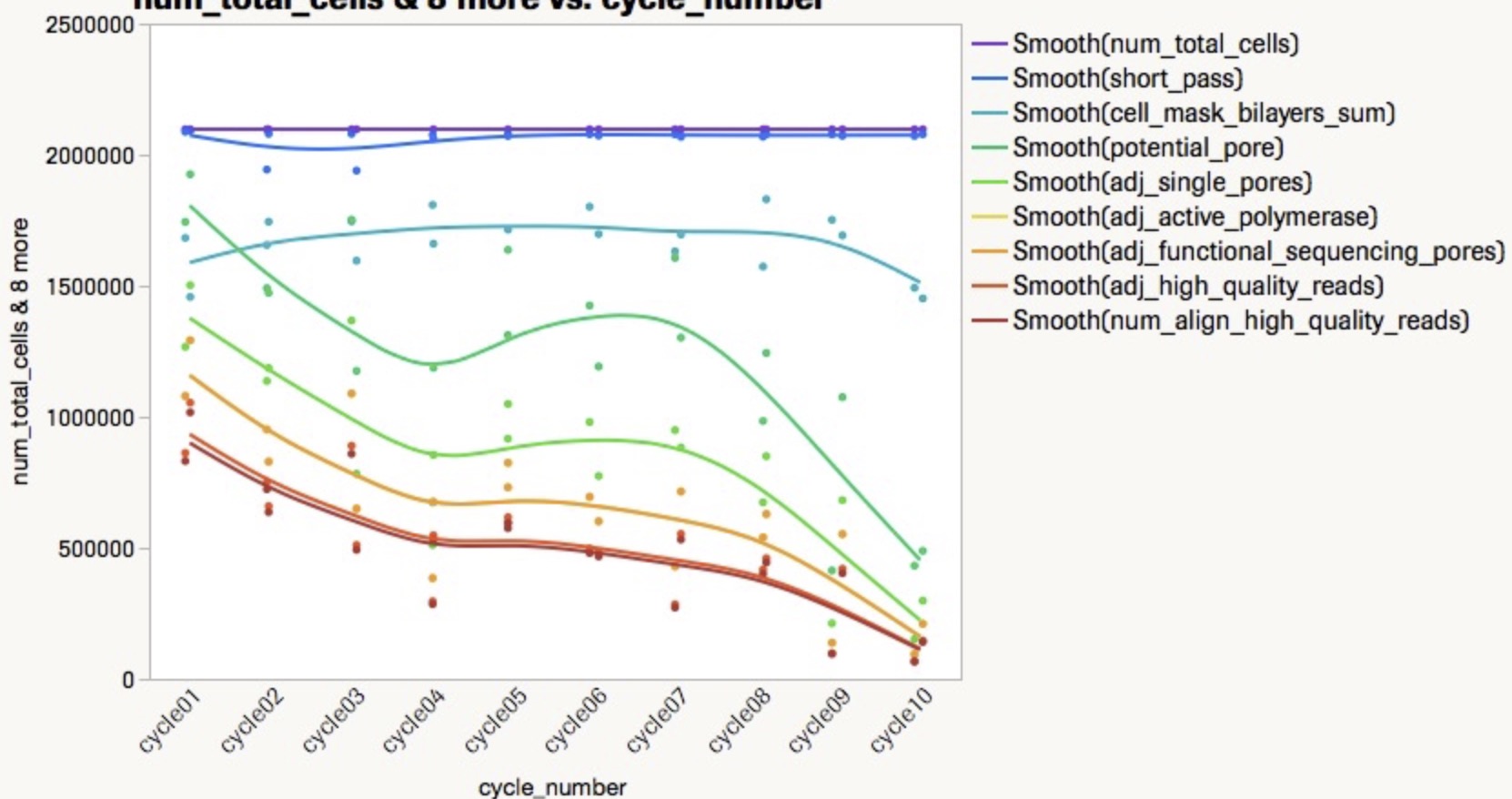
尝试结果
尝试1
我试图创建条形图(在一些帮助下)&我是成功的,但是用散点图我做不到。
下面的代码可将其转换为条形图
df1 = df.groupby(['token','variable']).agg({'value': 'mean'})
df1.reset_index(inplace=True)
df1.sort_values('value',inplace=True,ascending=False)
fig,ax = plt.subplots()
fig.set_size_inches(16,8)
#to get different colors for each of the variable assign the variable to hue
g=sns.barplot(x='token',y='value',data=df1, hue='variable',ax=ax)
#Code for to put legend outside the plot
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
# Put a legend to the right of the current axis
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# Adding respective values to the top of each bar
for p in ax.patches:
ax.annotate("%d" % p.get_height(), (p.get_x() + p.get_width() / 2, p.get_height()),
ha='center', va='center', fontsize=11, color='black', xytext=(0, 10),
textcoords='offset points',fontweight='bold')
plt.show()尝试2
g2=sns.lmplot(x='cycle_number',y='value',data=df), this gives error
ValueError: could not convert string to float: 'cycle10'我知道错误在这里意味着什么,但是试图复制到输出代码时我感到无能为力。
尝试3:
sns.lmplot('cycle_number', 'value', data=df, hue='variable', fit_reg=False)生成的输出:空白网格
回答 1
Stack Overflow用户
发布于 2018-08-24 12:57:06
用途:
sns.pointplot('cycle_number', 'value', data=df, hue='variable')
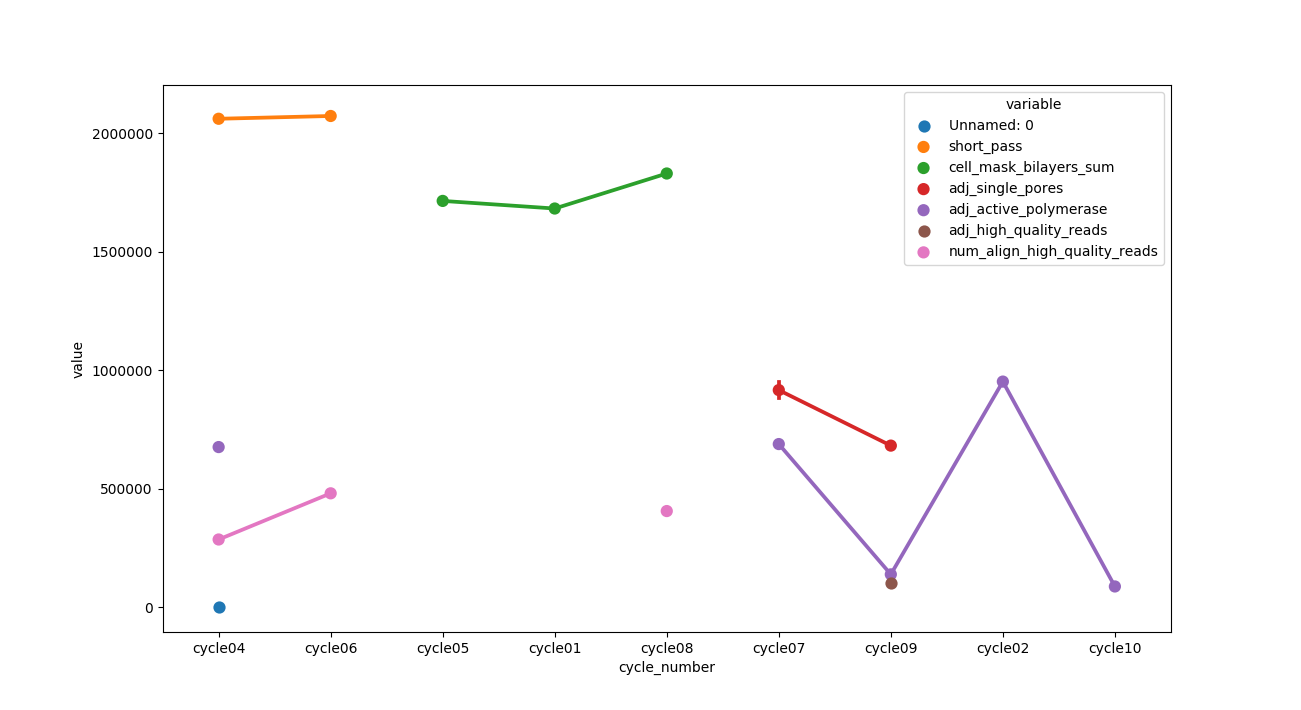
documnetation: https://seaborn.pydata.org/generated/seaborn.pointplot.html
使用此输出与预期输出生成输出。
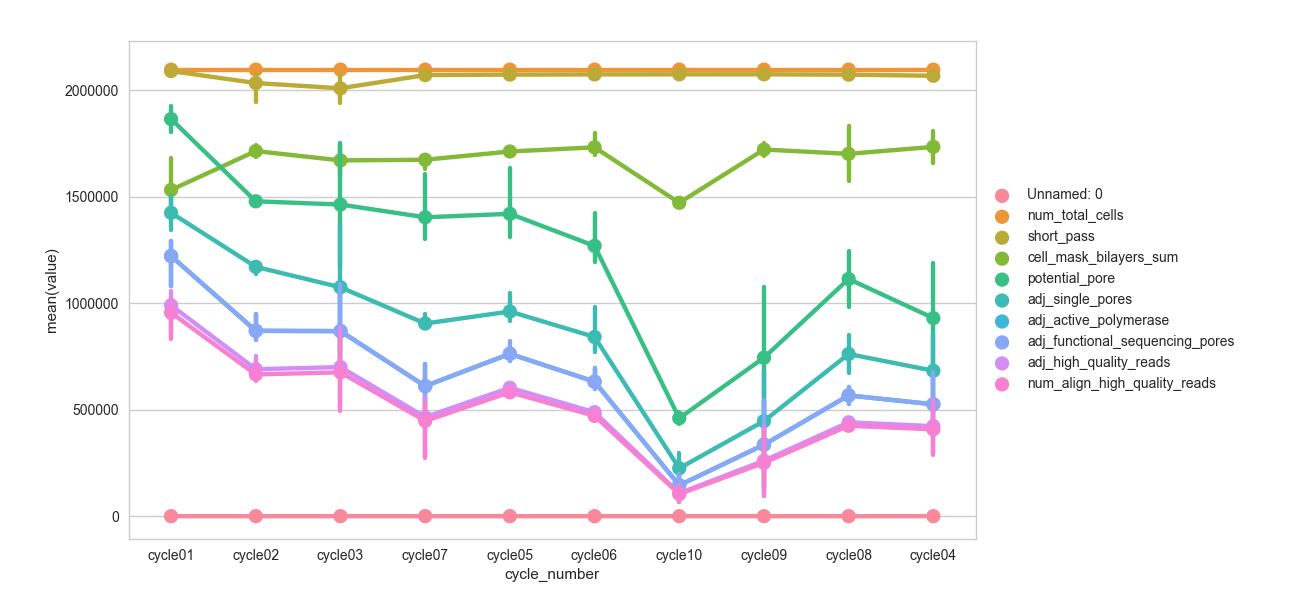
试试这个:
df = pd.DataFrame(d)
df['cycle_number'] = df['cycle_number'].str.replace('cycle', '')
df['cycle_number'] = df['cycle_number'].apply(pd.to_numeric)
print(df)
fig, ax = plt.subplots()
fig.set_size_inches(16, 4)
# sns.pointplot('cycle_number', 'value', data=df, hue='variable', err_style="bars", ci=68)
sns.lmplot('cycle_number', 'value', data=df, hue='variable', ci=None, order=2, truncate=True)
# use order = 5 to see more curveorder=2的输出
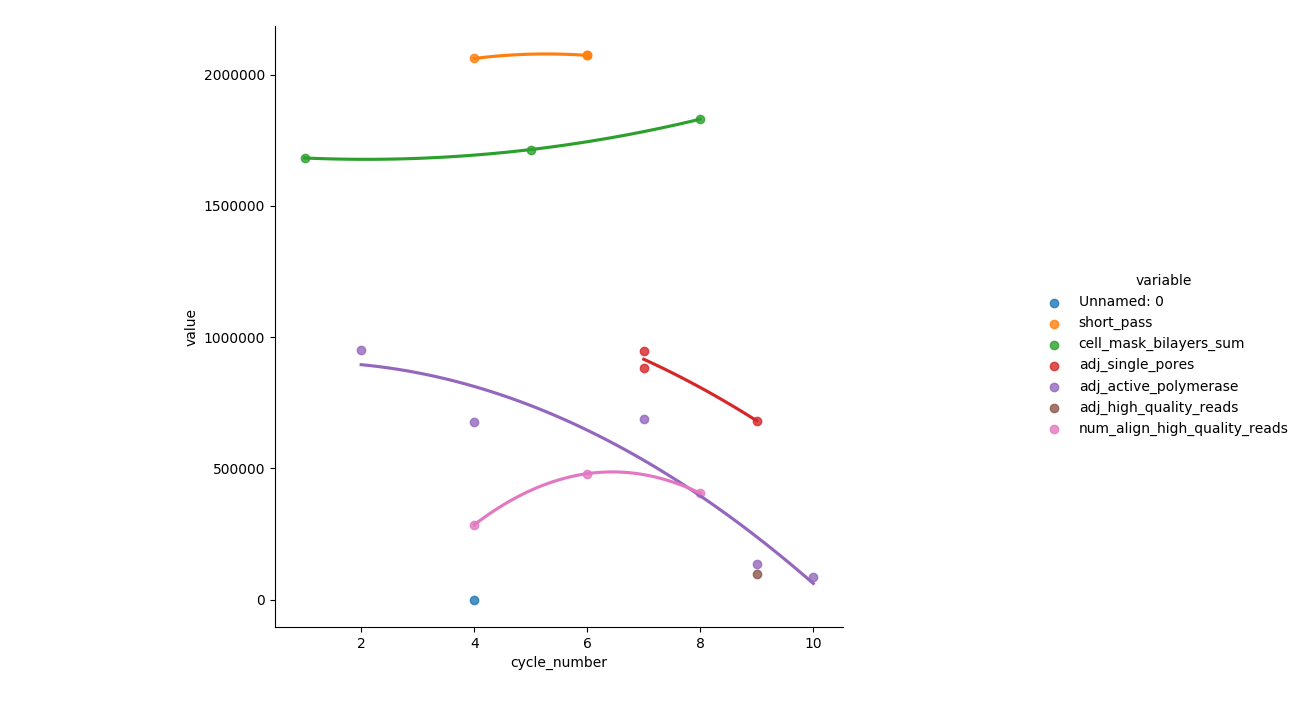
根据最新的共享代码(用于order=2__)输出
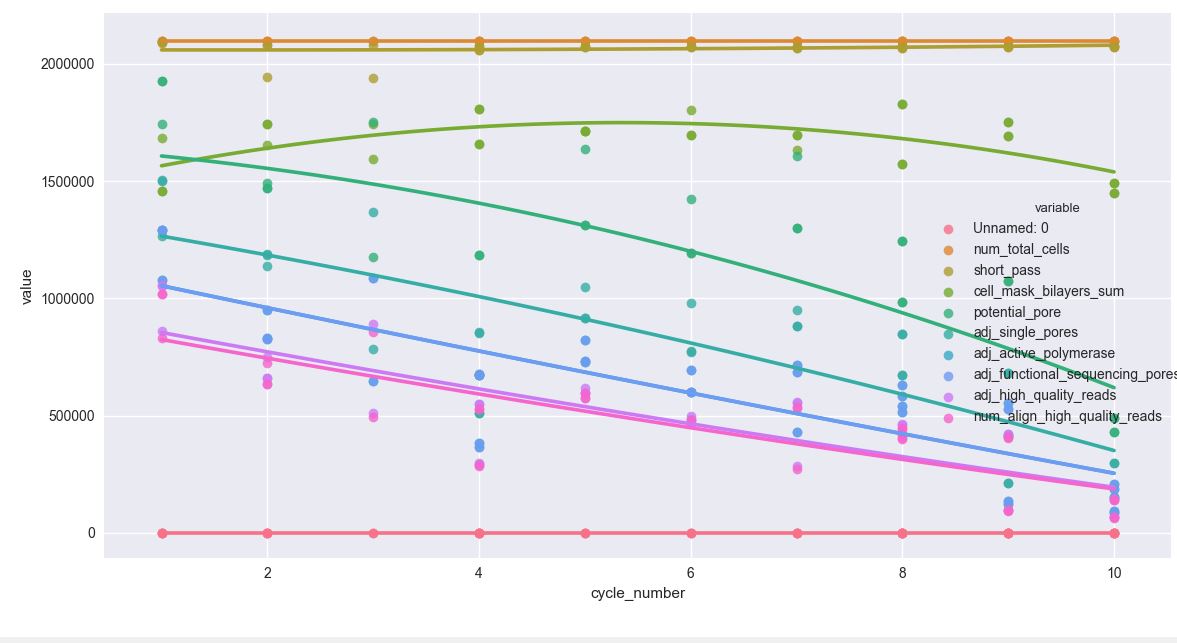
- 图例与图形区域重叠。
产出4(用于order=5)*:
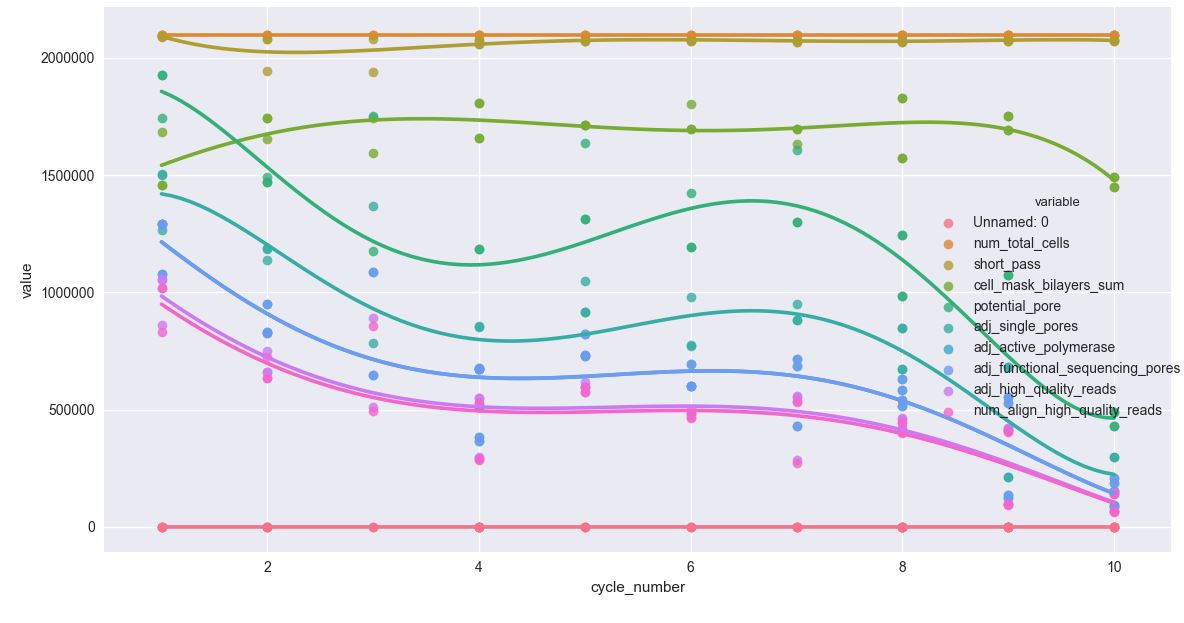
图形曲线非常精细,只是图例与绘图区域重叠。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/52004437
复制相关文章
相似问题

