mlr -如何看到影响目标变量的方向(正或负)特性
让我们从一个简单的线性回归输出(抄袭而来)开始,
Call:
lm(formula = a1 ~ ., data = clean.algae[, 1:12])
Residuals:
Min 1Q Median 3Q Max
-37.679 -11.893 -2.567 7.410 62.190
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 42.942055 24.010879 1.788 0.07537 .
seasonspring 3.726978 4.137741 0.901 0.36892
seasonsummer 0.747597 4.020711 0.186 0.85270
seasonwinter 3.692955 3.865391 0.955 0.34065
sizemedium 3.263728 3.802051 0.858 0.39179
sizesmall 9.682140 4.179971 2.316 0.02166 *
speedlow 3.922084 4.706315 0.833 0.40573
speedmedium 0.246764 3.241874 0.076 0.93941
mxPH -3.589118 2.703528 -1.328 0.18598
mnO2 1.052636 0.705018 1.493 0.13715
Cl -0.040172 0.033661 -1.193 0.23426
NO3 -1.511235 0.551339 -2.741 0.00674 **
NH4 0.001634 0.001003 1.628 0.10516
oPO4 -0.005435 0.039884 -0.136 0.89177
PO4 -0.052241 0.030755 -1.699 0.09109 .
Chla -0.088022 0.079998 -1.100 0.27265
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 17.65 on 182 degrees of freedom
Multiple R-squared: 0.3731, Adjusted R-squared: 0.3215
F-statistic: 7.223 on 15 and 182 DF, p-value: 2.444e-12从这个输出中,我们可以看到模型的适应度以及变量对目标变量的显着影响。另外,我们可以通过查看系数的符号来判断变量是负的还是正的。
现在看看这个例子包装手册,
### Select features
sfeats = selectFeatures(learner = "surv.coxph", task = wpbc.task, resampling = rdesc,
control = ctrl, show.info = FALSE)
sfeats
## FeatSel result:
## Features (14): mean_radius, mean_compactness, mean_concavepoints, mean_symmetry, mean_fractaldim, SE_perimeter, SE_area, SE_concavity, SE_fractaldim, worst_radius, worst_perimeter, worst_concavity, worst_concavepoints, tsize
## cindex.test.mean=0.6718346从上面的输出中,我可以看到一些重要的特性。我的问题是,我怎样才能看到(正或负)特征(自变量)影响目标变量的方向?有人在这问题上帮忙吗?推荐阅读材料将不胜感激。
加法
我试图在示例多标签分类示例中实现您的建议,
library(mlr)
library(mmpf)
yeast <- getTaskData(yeast.task)
labels <- colnames(yeast)[1:14]
yeast.task <- makeMultilabelTask(id = "multi", data = yeast, target = labels)
lrn.br <- makeLearner("classif.rpart", predict.type = "prob")
lrn.br <- makeMultilabelBinaryRelevanceWrapper(lrn.br)
mod <- mlr::train(lrn.br, yeast.task, subset = 1:1500, weights = rep(1/1500, 1500))
pred <- predict(mod, newdata = yeast[1501:1600,])
performance(pred, measures = list(multilabel.subset01, multilabel.hamloss, multilabel.acc,
multilabel.f1, timepredict))
rdesc <- makeResampleDesc(method = "CV", stratify = FALSE, iters = 3)
r <- resample(learner = lrn.br, task = yeast.task, resampling = rdesc, show.info = FALSE)
getMultilabelBinaryPerformances(pred, measures = list(acc, mmce, auc))
getMultilabelBinaryPerformances(r$pred, measures = list(acc, mmce))
getLearnerModel(mod)
pd <- generatePartialDependenceData(mod, yeast.task)
plotPartialDependence(pd)最后三行给出了下面的输出。我不知道这些是否有用。知道我是不是做错什么了吗?
> getLearnerModel(mod)
$label1
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label1; obs = 1500; features = 103
Hyperparameters: xval=0
$label2
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label2; obs = 1500; features = 103
Hyperparameters: xval=0
$label3
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label3; obs = 1500; features = 103
Hyperparameters: xval=0
$label4
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label4; obs = 1500; features = 103
Hyperparameters: xval=0
$label5
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label5; obs = 1500; features = 103
Hyperparameters: xval=0
$label6
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label6; obs = 1500; features = 103
Hyperparameters: xval=0
$label7
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label7; obs = 1500; features = 103
Hyperparameters: xval=0
$label8
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label8; obs = 1500; features = 103
Hyperparameters: xval=0
$label9
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label9; obs = 1500; features = 103
Hyperparameters: xval=0
$label10
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label10; obs = 1500; features = 103
Hyperparameters: xval=0
$label11
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label11; obs = 1500; features = 103
Hyperparameters: xval=0
$label12
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label12; obs = 1500; features = 103
Hyperparameters: xval=0
$label13
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label13; obs = 1500; features = 103
Hyperparameters: xval=0
$label14
Model for learner.id=classif.rpart; learner.class=classif.rpart
Trained on: task.id = label14; obs = 1500; features = 103
Hyperparameters: xval=0
>
> pd <- generatePartialDependenceData(mod, yeast.task)
Error in data.table(preds, design[, vars, drop = FALSE], key = vars) :
column or argument 1 is NULL
> plotPartialDependence(pd)
Error in checkClass(x, classes, ordered, null.ok) : object 'pd' not found回答 2
Stack Overflow用户
发布于 2018-07-19 07:33:20
如果要提取底层学习者模型的系数,则必须在mlr中使用getLearnerModel():
library(mlr)
mod = train(learner = "surv.coxph", task = lung.task)
getLearnerModel(mod)输出:
Call:
survival::coxph(formula = f, data = data)
coef exp(coef) se(coef) z p
inst -3.04e-02 9.70e-01 1.31e-02 -2.31 0.02062
age 1.28e-02 1.01e+00 1.19e-02 1.07 0.28340
sex -5.67e-01 5.67e-01 2.01e-01 -2.81 0.00489
ph.ecog 9.07e-01 2.48e+00 2.39e-01 3.80 0.00014
ph.karno 2.66e-02 1.03e+00 1.16e-02 2.29 0.02223
pat.karno -1.09e-02 9.89e-01 8.14e-03 -1.34 0.18016
meal.cal 2.60e-06 1.00e+00 2.68e-04 0.01 0.99224
wt.loss -1.67e-02 9.83e-01 7.91e-03 -2.11 0.03465
Likelihood ratio test=33.7 on 8 df, p=5e-05
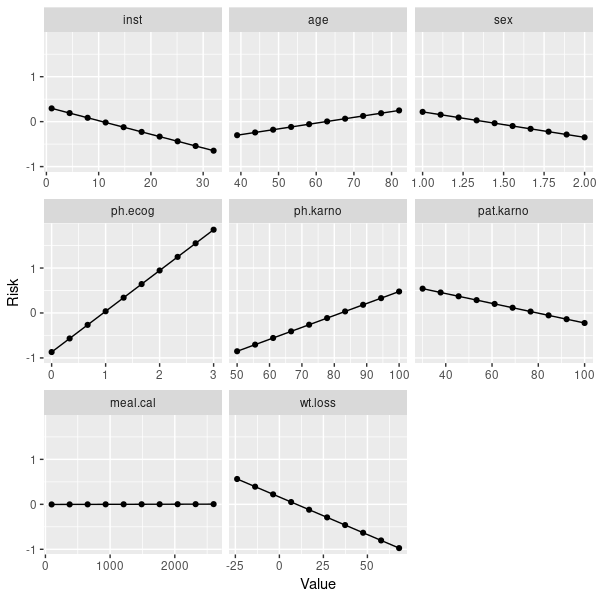
n= 167, number of events= 120如果你对独立于学习者的互动性感兴趣,那么你可以看一看部分依赖情节。对于coxph,它们并不是惊人的线性:
pd = generatePartialDependenceData(mod, lung.task)
plotPartialDependence(pd)
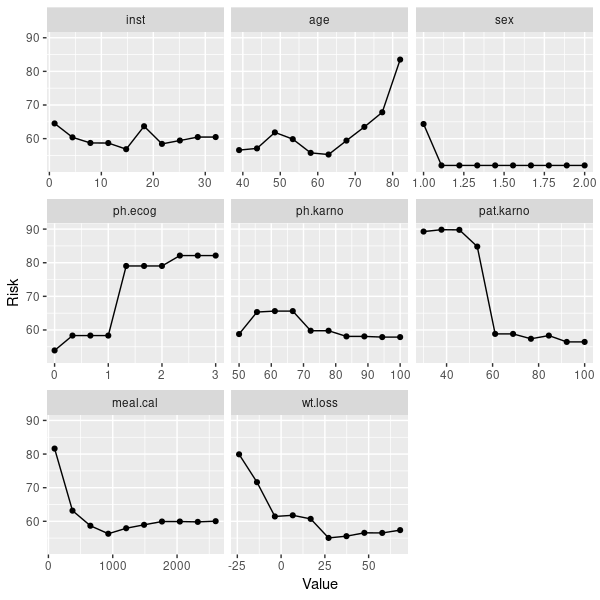
但是,您也可以使用随机森林获取生存数据:
mod2 = train(learner = "surv.randomForestSRC", task = lung.task)
pd2 = generatePartialDependenceData(mod2, lung.task)
plotPartialDependence(pd2)
然而,部分依赖情节也必须谨慎地加以解释。所以你应该读一读它,例如这里。你也可以看看ICE的阴谋。
Stack Overflow用户
发布于 2018-07-19 07:28:43
您可以使用结果的$opt.path成员获得更多关于在特性选择过程中发生的事情的信息,但是通常来说,对单个特性提供正面和负面的相关性/效果是没有意义的。对于大多数模型,您不能像线性回归那样定义方向相关性,因为它对特定的模型没有意义。特征选择函数与模型无关.
即使评估一个特定的特性是提高了模型的性能(正效应)还是降低了模型的性能(负面效应),对于这种类型的选择也没有意义,因为它考虑到了特性的交互作用。对于一个特定的特性,您可能会对其他特性的特定子集产生积极的影响,而对另一个特性则会产生负面的影响。
最后,与输出相关的特性不仅依赖于模型,而且还依赖于特性--它只适用于数字特性。
https://stackoverflow.com/questions/51407895
复制相似问题

