ggfortify不支持超越多个协变量?
下面的示例是通过添加“年龄”从文档示例中修改的
d.coxph <- (survfit(Surv(time, status) ~ sex+age, data = lung))
autoplot(d.coxph)我将得到以下错误:
levels<-(*tmp*,value = if (nl == nL) as.character(labels) level paste0(标签):因子级别41被复制)中的错误 输入一个帧号,或0以退出1:自动绘图(d.coxph)>levels<-中的错误(*tmp*,value = if (nl == nL) as.character(标签)==paste0(标签,:因子级别41被复制) 输入一个帧号,或0以退出1: autoplot(d.coxph) 2: autoplot.survfit(d.coxph) 3: fortify(object,surv.connect = surv.connect,fun = surv.connect ) 4: fortify.survfit(object,surv.connect= surv.connect,fun =surv.connect) 5: factor(groupIDs,model$strata),level=groupID) 2: autoplot.survfit(d.coxph) 3: fortify(object,surv.connect = surv.connect,fun =fortify.survfit) 4: fortify.survfit(object,surv.connect = surv.connect,fun = fun) 5: rep(groupIDs,模型$层),levels = groupIDs)
回答 1
Stack Overflow用户
发布于 2018-05-01 09:49:32
一种可能的解决办法是将连续变量age划分为几个类别,并在覆盖公式中考虑性别和年龄之间的相互作用:
library(survival)
data(lung)
lung$age2cat <- cut(lung$age,breaks=2)
lung$sex <- factor(lung$sex, labels=c("F","M"))
d.coxph <- survfit(Surv(time, status) ~ interaction(sex,age2cat), data = lung)
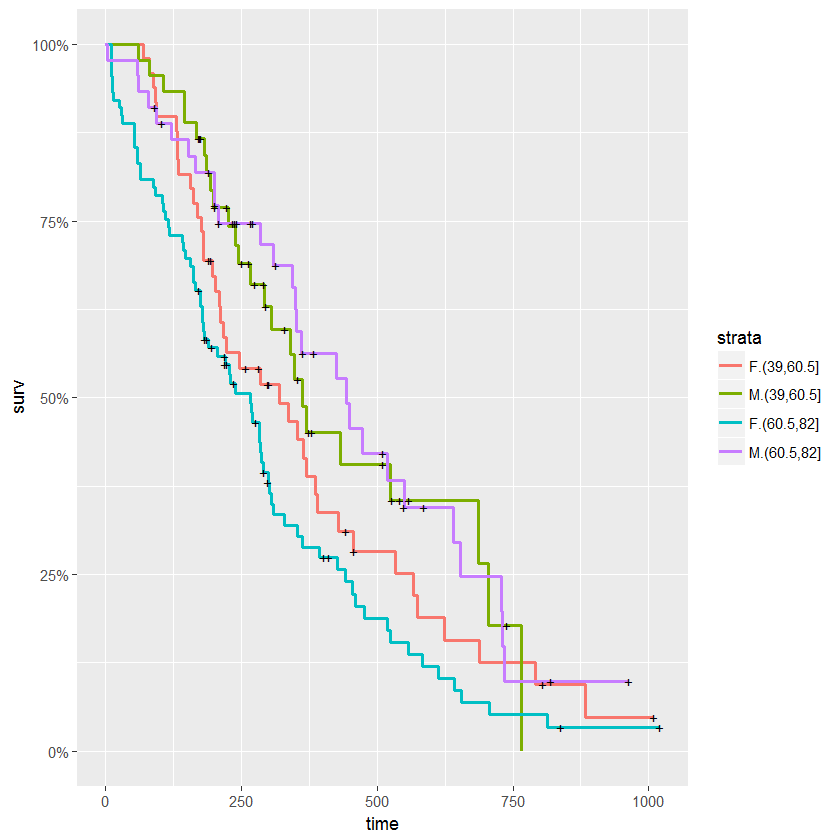
autoplot(d.coxph, conf.int=F, surv.size=1)
https://stackoverflow.com/questions/50110752
复制相似问题

