合并多个hclust对象(或树状图)
合并多个hclust对象(或树状图)
提问于 2018-03-28 18:24:02
是否有一种简单的方法可以在根处合并多个hclust对象(或树状图)?
为了说明我的问题,我把这个例子做得尽可能完整。
假设我想按区域对USArrests进行聚类,然后将所有的hclust对象联合起来,在一个热图中将它们绘制在一起。
USArrests
Northeast <- c("Connecticut", "Maine", "Massachusetts", "New Hampshire", "Rhode Island",
"Vermont", "New Jersey", "New York", "Pennsylvania")
Midwest <- c("Illinois", "Indiana", "Michigan", "Ohio", "Wisconsin",
"Iowa", "Kansas", "Minnesota", "Missouri", "Nebraska", "North Dakota",
"South Dakota")
South <- c("Delaware", "Florida", "Georgia", "Maryland", "North Carolina",
"South Carolina", "Virginia", "West Virginia",
"Alabama", "Kentucky", "Mississippi", "Tennessee", "Arkansas",
"Louisiana", "Oklahoma", "Texas")
West <- c("Arizona", "Colorado", "Idaho", "Montana", "Nevada", "New Mexico",
"Utah", "Wyoming", "Alaska", "California", "Hawaii", "Oregon", "Washington")
h1 <- hclust(dist(USArrests[Northeast,]))
h2 <- hclust(dist(USArrests[Midwest,]))
h3 <- hclust(dist(USArrests[South,]))
h4 <- hclust(dist(USArrests[West,]))现在我有了4个hclust对象(h1通过h4)。我通常是这样合并的:
hc <- as.hclust(merge(merge(merge(
as.dendrogram(h1), as.dendrogram(h2)), as.dendrogram(h3)),
as.dendrogram(h4)))然后,要绘制它们,我必须根据hclust对象重新排序矩阵,然后再绘制(我添加了一些注释以使绘图更清晰):
usarr <- USArrests[c(Northeast, Midwest, South, West),]
region_annotation <- data.frame(Region = c(rep("Northeast", length(Northeast)),
rep("Midwest", length(Midwest)),
rep("South", length(South)),
rep("West", length(West))),
row.names = c(Northeast, Midwest, South, West))
pheatmap(usarr, cluster_rows = hc,
annotation_row = region_annotation)
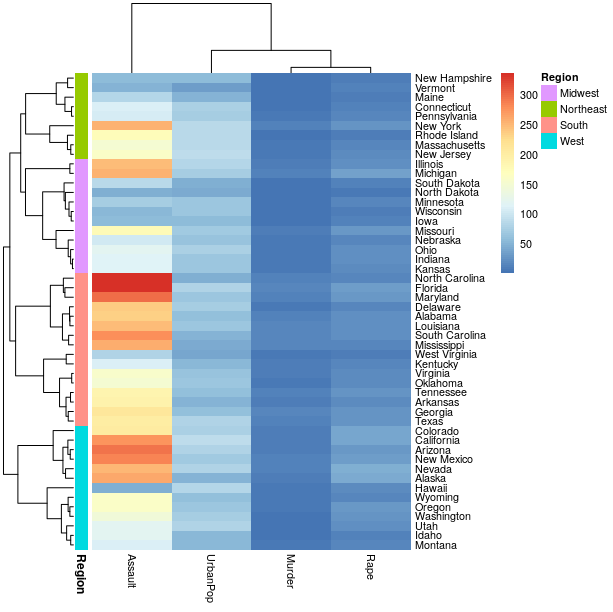
总之:有比合并所有单独的集群更容易做到这一点的方法吗?
回答 2
Stack Overflow用户
回答已采纳
发布于 2018-03-29 15:35:56
最后,我做了几个函数来更自动地完成这个任务。(在我的版本中,我还增加了对相关性“距离”的支持,因此它要大一点)
hclust_semisupervised <- function(data, groups, dist_method = "euclidean",
dist_p = 2, hclust_method = "complete") {
hclist <- lapply(groups, function (group) {
hclust(dist(data[group,], method = dist_method, p = dist_p), method = hclust_method)
})
hc <- .merge_hclust(hclist)
data_reordered <- data[unlist(groups),]
return(list(data = data_reordered, hclust = hc))
}
.merge_hclust <- function(hclist) {
#-- Merge
d <- as.dendrogram(hclist[[1]])
for (i in 2:length(hclist)) {
d <- merge(d, as.dendrogram(hclist[[i]]))
}
as.hclust(d)
}有了USArrests和区域向量,我就这样调用hclust_semisupervised:
semi_hc <- hclust_semisupervised(USArrests, list(Northeast, Midwest, South, West)现在绘制热图:
pheatmap(semi_hc$data, cluster_rows = semi_hc$hclust,
annotation_row = region_annotation)Stack Overflow用户
发布于 2018-03-28 19:56:24
要创建合并的hclust对象,可以在使用new.env创建的自定义环境中安全地使用new.env。
在不使用<<-的情况下,可能有其他方法一次创建两个合并的对象。希望有人能点亮它。
我试着使用do.call('merge', list( dendrograms of h1, h2, h3, h4 )。但是它没有工作,因为hclust需要在顶部有两个分支,而不是4个分支。
代码:
library('pheatmap')
myenv <- new.env()
myenv$hc <- as.dendrogram( hclust( dist(USArrests[Northeast,])))
invisible( lapply( list( Midwest, South, West), function(x){
myenv$hc <<- merge( myenv$hc, as.dendrogram( hclust( dist( USArrests[ x, ]) )) )
NULL
} ) )
myenv$hc <- as.hclust(myenv$hc)图:
pheatmap(usarr, cluster_rows = myenv$hc,
annotation_row = region_annotation)
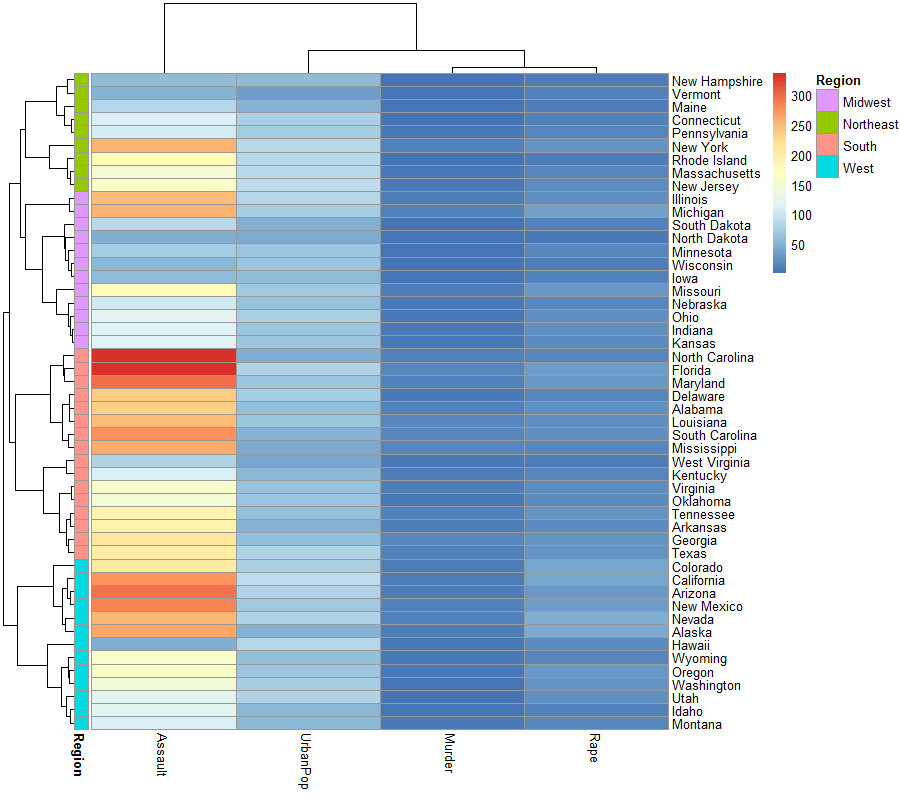
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/49541604
复制相关文章
相似问题

