使用自动绘图(ggfortify)显示非默认的主组件
使用自动绘图(ggfortify)显示非默认的主组件
提问于 2016-03-09 11:33:51
我想使用包PC2的函数autoplot()来绘制针对PC3的ggfortify。默认情况下,只显示PC1和PC2:
library(ggfortify)
myPCA <- prcomp(iris[-5])
autoplot(myPCA)通过重新排序和重命名prcomp对象中的列,我可以得到我想要的:
myPCAtrunc <- myPCA
myPCAtrunc[[1]] <- myPCAtrunc[[1]][c(2,3,1,4)]
myPCAtrunc[[2]] <- myPCAtrunc[[2]][,c(2,3,1,4)]
colnames(myPCAtrunc[[2]]) <- c("PC1","PC2","PC3","PC4") # fake names
myPCAtrunc[[5]] <- myPCAtrunc[[5]][,c(2,3,1,4)]
colnames(myPCAtrunc[[5]]) <- c("PC1","PC2","PC3","PC4") # fake names
autoplot(myPCAtrunc, xlab = "PC2", ylab="PC3")我知道这是正确的,因为它和plot(myPCA$x[, c(2,3)])是一样的。
但必须有一个更清洁的方法来解决这个问题。一些想法?
回答 3
Stack Overflow用户
回答已采纳
发布于 2016-03-09 12:01:53
当查看被调用的方法时,它看起来只被设计为只绘制PC1和PC2:
getS3method("autoplot", class(myPCA) )
> ...
> if (is_derived_from(object, "prcomp")) {
> x.column <- "PC1"
> y.column <- "PC2"
> loadings.column <- "rotation"
> }
> ...如果这是您的选项,我建议您使用ggbiplot包并设置choices参数:
library(ggbiplot)
ggbiplot(myPCA, choices = 2:3 , var.axes =FALSE)
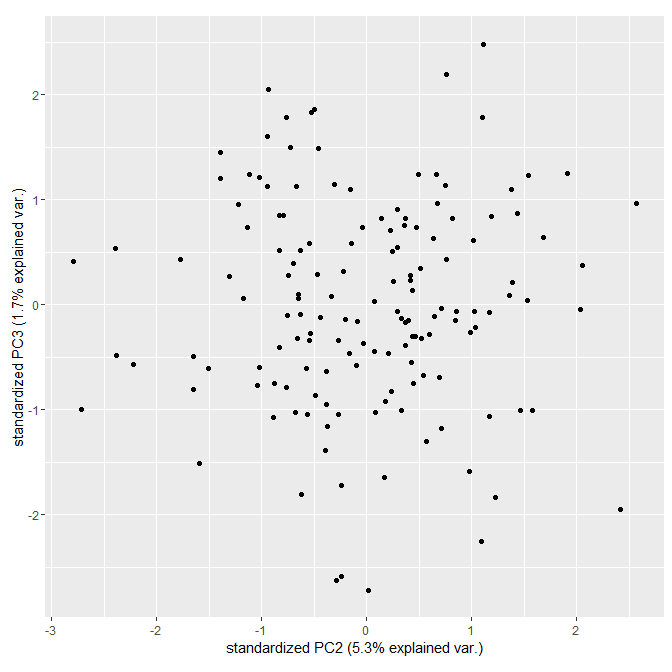
Stack Overflow用户
发布于 2017-04-26 23:35:00
这个问题最近得到了解决(这里)。
autoplot(myPCA, # your prcomp object
x = 2, # PC2
y = 3) # PC3Stack Overflow用户
发布于 2017-02-17 15:59:38
您只能修改prcomp对象。然后像这样改变y标签:
pca_test=pca
pca_test$x=pca_test$x[,c(1,3)]
colnames(pca_test$x)=c("PC1","PC2")
pca_test$rotation=pca_test$rotation[,c(1,3)]
colnames(pca_test$rotation)=c("PC1","PC2")
autoplot(pca_test,data=df,colour='study',shape='species')+scale_color_manual(values=c("Red","Blue","Green","Purple","Brown","Orange","Black"))+scale_fill_manual(values=c("Red","Blue","Green","Purple","Brown","Orange","Black"))+theme_bw()+ylab("PC3")希望能帮上忙!
JC
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/35890459
复制相关文章
相似问题

