PyMC 3的生存分析
PyMC 3的生存分析
提问于 2014-02-25 13:02:23
我试图将简单的生存模型from here (the first one in introduction)从PyMC 2移植到PyMC 3,但是,我没有发现任何与“观察”装饰器等价的东西,我试图编写一个新的发行版也失败了。有人能给我们举个例子吗?在PyMC 3中是如何做到的?
回答 1
Stack Overflow用户
回答已采纳
发布于 2015-08-19 21:12:52
这是一个棘手的端口,需要三个新概念:
theano张量的使用DensityDist的使用- 将
dict传递为observed
此代码提供了与上面链接的PyMC2版本相同的模型:
import pymc3 as pm
from pymc.examples import melanoma_data as data
import theano.tensor as t
times = data.t # not to be confused with the theano tensor t!
failure = (data.censored==0).astype(int)
with pm.Model() as model:
beta0 = pm.Normal('beta0', mu=0.0, tau=0.0001)
beta1 = pm.Normal('beta1', mu=0.0, tau=0.0001)
lam = t.exp(beta0 + beta1*data.treat)
def survival_like(failure, value):
return t.sum(failure * t.log(lam) - lam * value)
survive = pm.DensityDist('survive', survival_like,
observed={'failure': failure, 'value': times})
with model:
start = pm.find_MAP()
step = pm.NUTS(scaling=start)
trace = pm.sample(10000, step=step, start=start)
pm.traceplot(trace);产出如下:
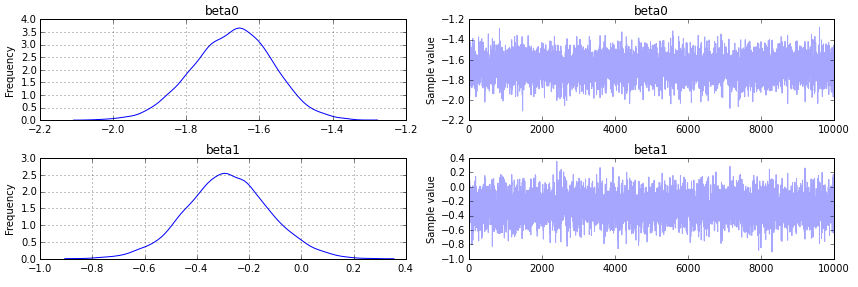
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/22015055
复制相关文章
相似问题

