如何在python中下载彩色KEGG路径图
如何在python中下载彩色KEGG路径图
提问于 2021-06-02 14:26:54
我有一个酶的列表(EC号,代码中的"df“),我想在KEGG路径图中突出显示。我想要制作一个python脚本来自动保存突出显示的映射。到目前为止,我只能手工下载urls:
from bioservices import KEGG
import webbrowser
s = KEGG()
def mapping(df, map_list):
for map in map_list:
names = ""
to_parse = s.get("ec"+map)
parsed = s.parse(to_parse)
for ECs in df["EC_numbers"]:
if str(ECs) in parsed["ENZYME"]:
names=names+"+EC:"+str(ECs)
url = "https://www.genome.jp/pathway/map"+map+names
s.logging.info(url)
webbrowser.open(url)
map_list_example = ["00520", "00010", "00020"]有什么想法吗?
回答 1
Stack Overflow用户
回答已采纳
发布于 2022-04-03 19:58:59
为此,您可以使用函数save_pathway。
from bioservices import KEGG
s = KEGG()
target_map = 'ko00340'
target_kos = ['K00765', 'K01814']
s.save_pathway(target_map, 'test.png', keggid=target_kos)这将产生
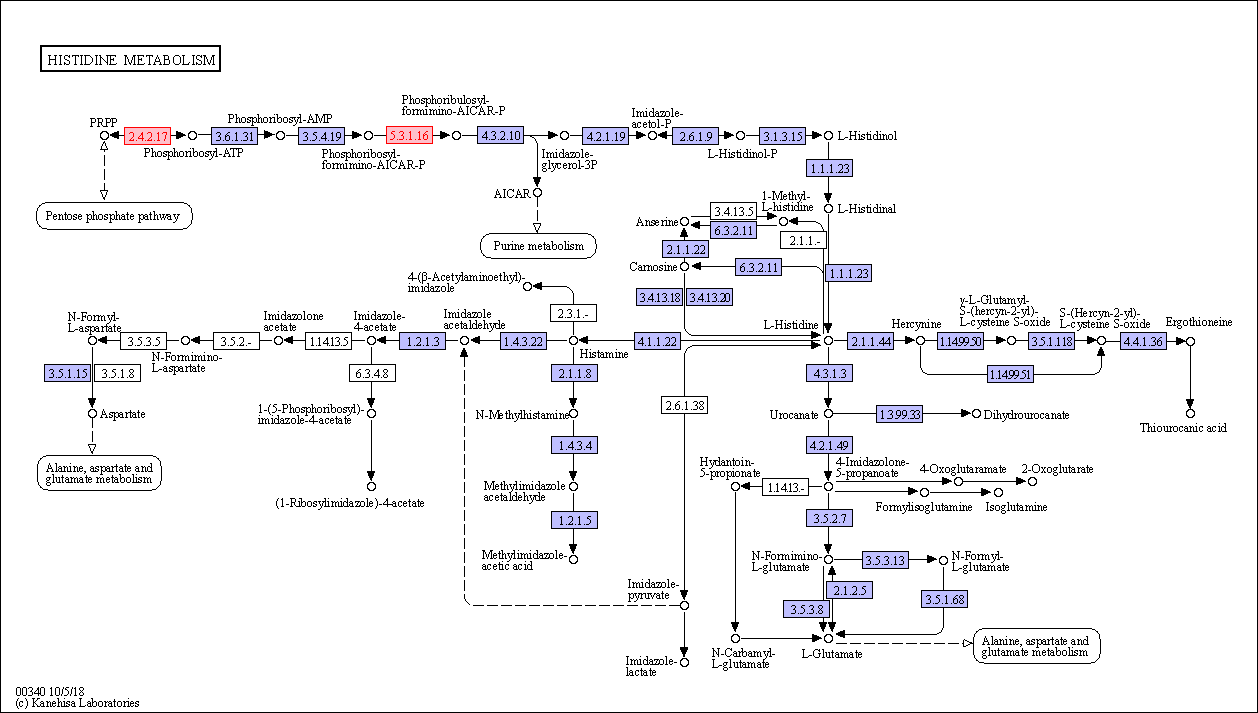
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/67807088
复制相关文章
相似问题

