如何编写文件而不用每次循环迭代覆盖它们
如何编写文件而不用每次循环迭代覆盖它们
提问于 2021-01-03 20:36:30
以下代码用于分析FASTA序列(.faa文件)的氨基酸组成
from Bio import SeqIO
from Bio.SeqUtils.ProtParam import ProteinAnalysis
import fastaparser
import pandas as pd
import matplotlib.pyplot as plt
pd.set_option('display.max_columns', None)
filename = input("Please enter the full path of the amino acid sequence file!: ")
pH_input = input("At which pH should the analysis be conducted? ")
flexibility_ynu = input("Do you wish a flexibility analysis?\n (1) Yes\n (0) No\n")
if pH_input == "":
pH= 7.4
elif pH_input != "":
pH = pH_input
f = open(filename + "_analysis.txt","w+")
for record in SeqIO.parse(filename, 'fasta'):
X = ProteinAnalysis(str(record.seq))
print("ANALYSIS OF", record, "\n ----------- \n -----------", file=f)
#
pd_count_amino_acids = pd.DataFrame(X.count_amino_acids(), index=[1])
print("number of amino acids: \n",pd_count_amino_acids , file=f)
plt_acc = pd_count_amino_acids.plot.bar()
plt.savefig(filename + "_count_amino_acids_plot.pdf")
#
pd_get_amino_acids_percent = pd.DataFrame(X.get_amino_acids_percent(), index=[1])
print("\n percentage of amino acids: \n", pd_get_amino_acids_percent, file=f)
plt_acp = pd_get_amino_acids_percent.plot.bar()
plt.savefig(filename + "amino_acids_percent_plot.pdf")
#
print("\n molecular weight: {:.2f}".format(X.molecular_weight()), file=f)
print("\n aromaticity: {:.2f}".format(X.aromaticity()), file=f)
print("\n instability index: {:.2f}".format(X.instability_index()), file=f)
if flexibility_ynu == "1":
print("\n flexibility: ", X.flexibility(), file=f)
print("\n IEP: ", X.isoelectric_point(), file=f)
print("therefore its charge at pH = ",pH," is {:.2f}".format(X.charge_at_pH(pH)), file=f)
print("secondary structure fraction: (Helix, Turn, Sheet): ", X.secondary_structure_fraction(), "\n\n\n", file=f)
f.close()
print("done")我现在想要的是绝对氨基酸和相对氨基酸的条形图,但是为每一个FASTA ID创建一个独特的图。
例如,011544有5个ID,所以我想生成10个唯一的地块(每个ID 2块,绝对1块,相对数1块)。
有办法这样做吗?
NC_011544.faa
>gi|212671454|ref|YP_002308464.1| replicase [Hosta virus X]
MARLREVFSSFTEPNLKTIVQQETYKLAKAELKTIQTYNPYAQTKDAADLLEDLGINTNPHAVTAHTHAA
AKSIENDLYGITSHYLPKTPITFLFMKRGKLQFFKRGPQHNDLFFYTTHEPKDVIRYQSEDQTADMFRVP
TSTGFIGDTLHFLSLKYLHRLFLKNPNLNTLYATMVLPPEAMYRMASIYPEIYQIQYQEDGFLYIPGGHG
GAAYFHTYDTLTWLRVGQFQAKEFTAHLPKVGDKGANHLFIIQRADLKTPKYRTFVPRRKWVTLPNIFLP
STQANHLFIIQRADLKTPKYRTFVPRRKWVTSNIFLPKHTNARKPILKQTMMQLFLYEKSVKEITFRDVF
AKIRQLIQTKDLEQFDPDELVRLANYVMHTSKLLEKDPYELIEGQGKLQDLVNPIKTWVSEKWQNWFGWK
DYTRLIRALKWVDVDLVLRVMNTRSTPTGIQTSELLPDEAGPPKSKKKRGGKKIPSPEPSRNCRSKSKRT
RGNRAQREKEPHRRKLRWQKENFQRVTVQVHQAPKGDPSPLARFSQSLKELPRRSQPRRLSKFQDFLMSS
TQTRFQIPSSLNRRAGHWRPKQQGTPPTTQEAGTEGPPTTQPGKPTASSPRAAPQPTANAETMEKGSQAS
SATTRGRDPVTDRTREQAPTNLTPEEEALPWKHWLKQLKAVGFKGNETQMDGDGTSISPIEQIKSCPGKP
KSVSKEILETLRSGHAPNFWKPDASRARAYTSDIKNRRTGAAVHMAPQAWKETMDFIAENAERTLHILRH
PWRRRFREEQMSSRDAHKFHFLFDETLVVCPTNELRRDWIDKLPLSEPGSVLTFERALMNPAKGTVIFDD
YTKLPAGFIEAYSICQPNVELVILTGDAKQASHHESNDNAMIAGLDPAAFEFSKFCRYYLNATHRNPRNL
ANALGIYSEKPGNLKVTFTNHLLPEMHILVPSLLKKATLEELGHKCSTYAGCQGVTLSKVQIYLDSNTTL
CSNEVLYTALSRAVEQINFVNSGPFNGPFWAKLEATPYLKTFLRLTREEKINEITPEEPKPKEPEPPKTH
FPVETSAHLYSSITEEMPEKHAREIYNKTHGHTNCVQTDEPLVQMFAHQQAKDEALFWETIEARLRITTS
EANVQELNEKRDIGDLLFHAYHKAMGLPKDPIPFENDLWETCAQEVQQTYLSKPINLIKNGEKRQGPDFD
KNAIMLFLKSQWVKKMEKLGAPTIKPGQTIASFHQITVMLYGTMARYMRRIRDRFCPKHILINCEKTPTQ
ISDFVKAQWDFSDFAYANDFTAFDQSQDGAMLQFEIIKAKFHNIPEDIILGYMDIKTNAKIFLGTLAIMR
LTGEGPTFDANTECNIAYTHLRFNVPENVAQVYAGDDSALSKVCPEKDSFKQFADRLTLKSKPQVFPQTQ
GAWAEFCGLLITPRGIIKDPVKLHASWVLATKLGTLQQIKCVNSYGEDLKLSYDLGDHLQELLSESQCRT
HQVTVRELVKFAGKVEKHQAEIRSVANGNIRQLPFFY
>gi|212671455|ref|YP_002308465.1| 26 kDa protein [Hosta virus X]
MATFASFLSSTRPDFERTNTPLTKPLVIHAVAGAGKTTLLRDFLRANPLTNAQTLGTPDCPTLDGAYIRP
FSGPVANLVNILDEYTAHRHGSWDVLIADPLQHYERAKLPHYICKRSHRLCPATARLLRKLGLDIHSYRE
DESEISFSDIFSGQLEGTVLPLTPLCKDLLERHSCPFKCPSEFIGEQDDIITVVSEIPLSKHPDKTALYR
ALTRHTRRLNVLAPPPYPTP
>gi|212671456|ref|YP_002308466.1| 13 kDa protein [Hosta virus X]
MSSPHRLTPPPNYTPVLLAVVIGVGLAVVTNQLTRSTLPHVGDNIHSLPHGGNYKDGTKSVIYRGPAPFQ
RSHSTAPPFNAVLLLTFAIWFLSCRTRRAAIGIHVCHTCSQTREQQ
>gi|212671457|ref|YP_002308467.1| 8 kDa protein [Hosta virus X]
MQSFCSHLRSGSFPVVLGALLLAFTCATLVLRLGNNNSNNCLIYVDGARAFLEGNCAGISAEVVAALRPH
SHAG
>gi|212671458|ref|YP_002308468.1| coat protein [Hosta virus X]
MASDAPTPPAAPSPVTFTAPTQEQLTSLALPIISTRLPSPDVLNQISVKWQELGVPTASISSTAIALCMA
CYHSGSSGSTLIPGLAPGTTVNYTSLAAAVKSLATLREFARYFAPIIWNYAIEHKIPPANWAAMGYKENT
KYAAFDTFDSILNPAALQPTGGLIRQPTEEELLAHQANSALHIFDSLRNDFASTDGRVTRGHITSNVNSL
NYLPAPEGSS回答 1
Stack Overflow用户
回答已采纳
发布于 2021-01-04 06:48:56
- 代码已经为每条记录创建了一个图,但是,所有的记录都使用相同的文件名,所以每个新的绘图和
analysis.txt都要覆盖前一个记录。 - 将
id从record中提取为record_id,并使用它为每个循环创建唯一的文件名。 - 下面的代码现在将为每个
percent_plot.pdf、count_plot.pdf和analysis.txt生成一个id - 使用
with open,它将自动关闭文件f .T用于在绘图时转换数据,这将氨基酸名称放置在轴上,从而使绘图更容易阅读。
import pandas
import matplotlib.pyplot as plt
from Bio import SeqIO
from Bio.SeqUtils.ProtParam import ProteinAnalysis
# set pandas display option
pd.set_option('display.max_columns', None)
# input selections were removed for testing
filename = 'NC_011544.faa'
flexibility_ynu = '1'
pH = 7.4
for record in SeqIO.parse(filename, 'fasta'):
# get each record id to be used for unique file names
record_id = record.id.split('|')[1]
print(record)
# open the file and use the record id as part of the name, so there will be a unique file for each id
with open(f"{filename}_{record_id}_analysis.txt","w+") as f:
X = ProteinAnalysis(str(record.seq))
print("ANALYSIS OF", record, "\n ----------- \n -----------", file=f)
#
pd_count_amino_acids = pd.DataFrame(X.count_amino_acids(), index=[1])
display(pd_count_amino_acids) # display is for jupyter notebook, otherwise use print
print("number of amino acids: \n",pd_count_amino_acids , file=f)
plt_acc = pd_count_amino_acids.T.plot.bar(legend=False, figsize=(7, 5))
plt.title(f'Number of Amino Acids for {record_id} in {filename}')
plt.xticks(rotation=0)
# add the record id as part of the file name
plt.savefig(f'{filename}_{record_id}_count_amino_acids_plot.pdf')
plt.show()
#
pd_get_amino_acids_percent = pd.DataFrame(X.get_amino_acids_percent(), index=[1])
display(pd_get_amino_acids_percent.round(2)) # display is for jupyter notebook, otherwise use print
print("\n percentage of amino acids: \n", pd_get_amino_acids_percent, file=f)
plt_acp = pd_get_amino_acids_percent.T.plot.bar(legend=False, figsize=(7, 5))
plt.title(f'Percentage of Amino Acids for {record_id} in {filename}')
plt.xticks(rotation=0)
# add the record id as part of the file name
plt.savefig(f'{filename}_{record_id}_amino_acids_percent_plot.pdf')
plt.show()
#
print("\n molecular weight: {:.2f}".format(X.molecular_weight()), file=f)
print("\n aromaticity: {:.2f}".format(X.aromaticity()), file=f)
print("\n instability index: {:.2f}".format(X.instability_index()), file=f)
if flexibility_ynu == "1":
print("\n flexibility: ", X.flexibility(), file=f)
print("\n IEP: ", X.isoelectric_point(), file=f)
print("therefore its charge at pH = ",pH," is {:.2f}".format(X.charge_at_pH(pH)), file=f)
print("secondary structure fraction: (Helix, Turn, Sheet): ", X.secondary_structure_fraction(), "\n\n\n", file=f)
print('\n')第一ID
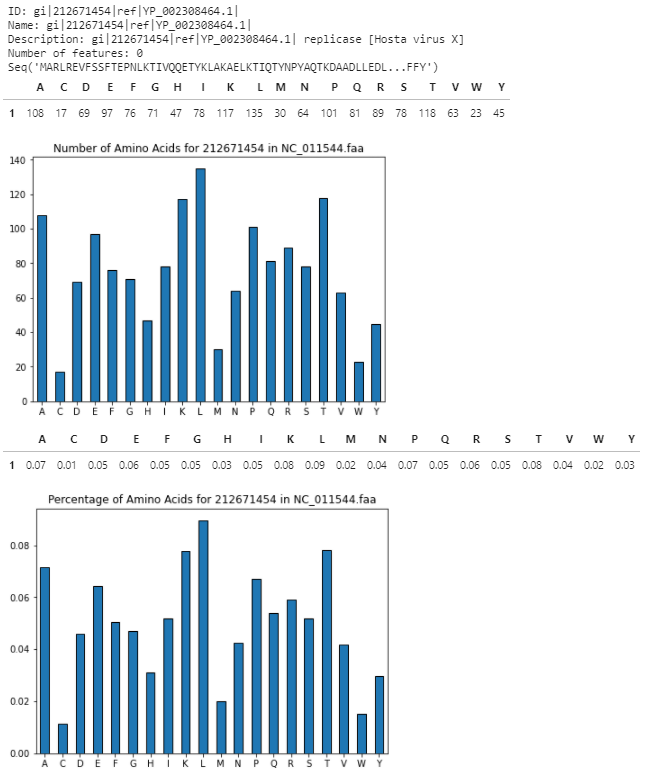
第二ID
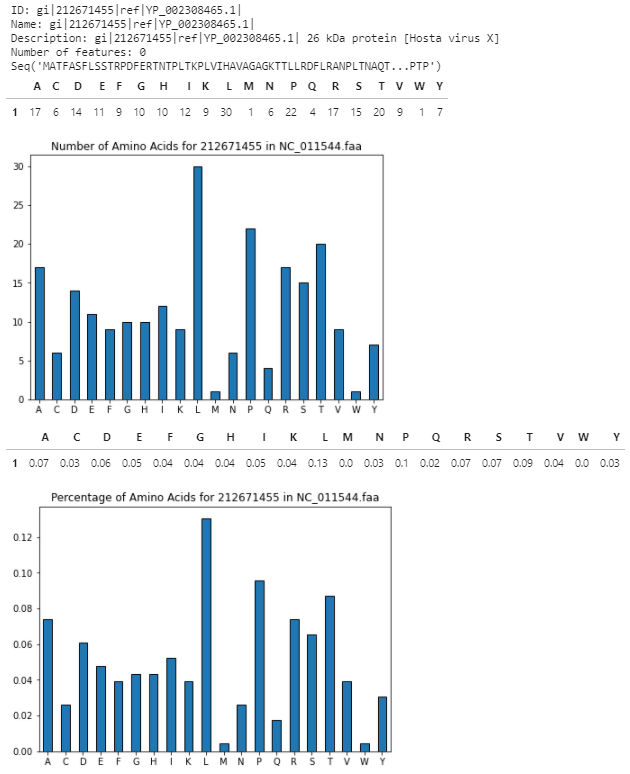
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/65554669
复制相关文章
相似问题

