瓷砖地块的整形表
瓷砖地块的整形表
提问于 2020-01-22 18:59:15
我有一个像这样的数据框架(我简化了它,它在现实中很长)。
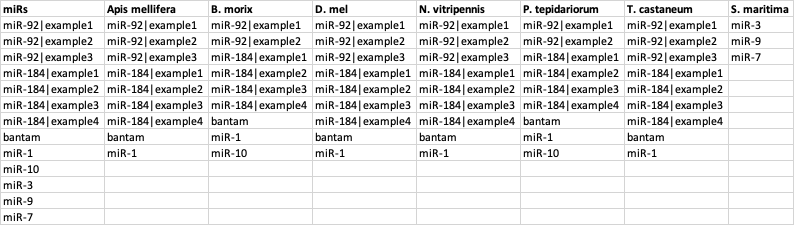
data <- structure(list(miRs = structure(c(10L, 11L, 12L, 3L, 4L, 5L,
6L, 1L, 2L, 7L, 9L, 8L), .Label = c("bantam", "miR-1", "miR-184|example1",
"miR-184|example2", "miR-184|example3", "miR-184|example4", "miR-3",
"miR-7", "miR-9", "miR-92|example1", "miR-92|example2", "miR-92|example3"
), class = "factor"), Apis.mellifera = structure(c(8L, 9L, 10L,
4L, 5L, 6L, 7L, 2L, 3L, 1L, 1L, 1L), .Label = c("", "bantam",
"miR-1", "miR-184|example1", "miR-184|example2", "miR-184|example3",
"miR-184|example4", "miR-92|example1", "miR-92|example2", "miR-92|example3"
), class = "factor"), B..morix = structure(c(9L, 10L, 5L, 6L,
7L, 8L, 2L, 3L, 4L, 1L, 1L, 1L), .Label = c("", "bantam", "miR-1",
"miR-10", "miR-184|example1", "miR-184|example2", "miR-184|example3",
"miR-184|example4", "miR-92|example1", "miR-92|example2"), class = "factor"),
D..mel = structure(c(8L, 9L, 10L, 4L, 5L, 6L, 7L, 2L, 3L,
1L, 1L, 1L), .Label = c("", "bantam", "miR-1", "miR-184|example1",
"miR-184|example2", "miR-184|example3", "miR-184|example4",
"miR-92|example1", "miR-92|example2", "miR-92|example3"), class = "factor"),
N..vitripennis = structure(c(8L, 9L, 10L, 4L, 5L, 6L, 7L,
2L, 3L, 1L, 1L, 1L), .Label = c("", "bantam", "miR-1", "miR-184|example1",
"miR-184|example2", "miR-184|example3", "miR-184|example4",
"miR-92|example1", "miR-92|example2", "miR-92|example3"), class = "factor"),
P..tepidariorum = structure(c(9L, 10L, 5L, 6L, 7L, 8L, 2L,
3L, 4L, 1L, 1L, 1L), .Label = c("", "bantam", "miR-1", "miR-10",
"miR-184|example1", "miR-184|example2", "miR-184|example3",
"miR-184|example4", "miR-92|example1", "miR-92|example2"), class = "factor"),
T..castaneum = structure(c(8L, 9L, 10L, 6L, 7L, 4L, 5L, 2L,
3L, 1L, 1L, 1L), .Label = c("", "bantam|LQNS02278082.1_33125",
"miR-1", "miR-184|LQNS02000211.1_1795", "miR-184|LQNS02000211.1_1950",
"miR-184|LQNS02000211.1_1952", "miR-184|LQNS02000211.1_1954",
"miR-92|example1", "miR-92|example2", "miR-92|example3"), class = "factor"),
S..maritima = structure(c(2L, 4L, 3L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L), .Label = c("", "miR-3", "miR-7", "miR-9"
), class = "factor")), class = "data.frame", row.names = c(NA,
-12L))理想情况下,我希望绘制一个平铺图,其中只显示每个miR的缺位/存在。然而,我有很多旅行,以重塑这张表的R,甚至是规划它。预期的产出将是:
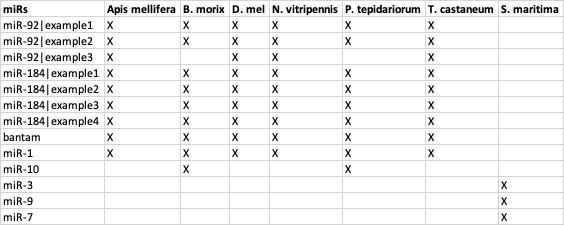
我想把它画出来。对此问题的任何帮助都将不胜感激。谢谢
回答 2
Stack Overflow用户
回答已采纳
发布于 2020-01-22 19:11:12
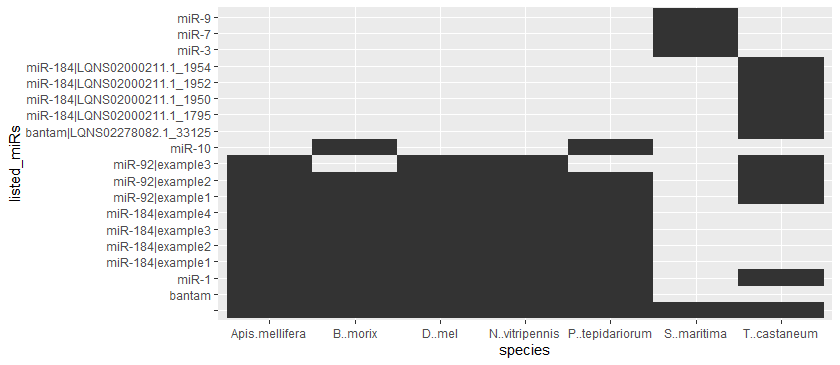
下面是一种使用tidyr::pivot_longer和tidyverse中的ggplot的方法。
library(tidyverse)
data %>%
pivot_longer(cols = -miRs, names_to = "species", values_to = "listed_miRs") %>%
ggplot(aes(species, listed_miRs)) +
geom_tile()
Stack Overflow用户
发布于 2020-01-22 19:08:46
我们可以用pivot_longer将其重塑为“long”格式,然后做一个count并将其重塑为“wide”
library(dplyr)
library(tidyr)
data %>%
type.convert(as.is = TRUE) %>%
pivot_longer(cols = -miRs) %>%
filter(value != "") %>%
count(name, value) %>%
pivot_wider(names_from = name, values_from = n, values_fill = list(n = 0))或者使用来自base R的base R
+(table(unlist(data), rep(names(data), each = nrow(data))) != 0)页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/59866566
复制相关文章
相似问题

