and在模型规格和置信区间方面的意外行为
我的数据是包含许多零的整数。我想用二项式广义线性模型分别对零点进行建模。在模型语句中,我在倾斜体的左侧指定了Y>0,这给了我一个二进制(TRUE,FALSE)向量。我使用emmeans包指定(type = "response")进一步分析了数据。然后,我意识到(根据我的实际数据),信任间隔似乎消失了。我尝试解决这个问题,并决定在我的数据框架中分别创建一个包含TRUE和FALSE值的新变量。这个解决了问题。为什么会发生这种情况?
下面是再现这种行为的代码(尽管效果并不像我原来的数据集那样明显):
require(emmeans)
# example data
d <- structure(list(X = structure(c(1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 3L, 3L, 3L, 3L,
3L, 3L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L
), .Label = c("A", "B", "C", "D"), class = "factor"), Y = c(0L,
4L, 4L, 5L, 6L, 5L, 6L, 7L, 8L, 9L, 0L, 0L, 3L, 4L, 1L, 5L, 2L,
3L, 2L, 1L, 0L, 0L, 0L, 0L, 0L, 12L, 11L, 6L, 8L, 11L, 0L, 0L,
0L, 0L, 0L, 12L, 13L, 11L, 12L, 16L)), class = "data.frame", row.names = c(NA,
-40L))
# add additional variable - set every value > 0 to TRUE, otherwise FALSE
d$no0 <- d$Y>0 下面是使用模型中的关系运算符>的第一个模型:
# binomial GLM using `Y>0` on the left side
m1 <- glm(Y>0 ~ X, family = binomial(), d)
summary(m1)
Call:
glm(formula = Y > 0 ~ X, family = binomial(), data = d)
Deviance Residuals:
Min 1Q Median 3Q Max
-2.1460 -1.1774 0.4590 0.7954 1.1774
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 2.1972 1.0540 2.085 0.0371 *
XB -0.8109 1.3175 -0.615 0.5382
XC -2.1972 1.2292 -1.788 0.0739 .
XD -2.1972 1.2292 -1.788 0.0739 .
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 50.446 on 39 degrees of freedom
Residual deviance: 44.236 on 36 degrees of freedom
AIC: 52.236
Number of Fisher Scoring iterations: 4下面是使用新变量的第二个模型:
# binomial GLM using variable no0
m2 <- glm(no0 ~ X, family = binomial(), d)
summary(m2)
Call:
glm(formula = no0 ~ X, family = binomial(), data = d)
Deviance Residuals:
Min 1Q Median 3Q Max
-2.1460 -1.1774 0.4590 0.7954 1.1774
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 2.1972 1.0540 2.085 0.0371 *
XB -0.8109 1.3175 -0.615 0.5382
XC -2.1972 1.2292 -1.788 0.0739 .
XD -2.1972 1.2292 -1.788 0.0739 .
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 50.446 on 39 degrees of freedom
Residual deviance: 44.236 on 36 degrees of freedom
AIC: 52.236
Number of Fisher Scoring iterations: 4到目前为止,输出是相同的。然后,我继续运行模型1和模型2的emmeans()函数,不使用-- type = "response"参数:
(em1 <- emmeans(m1, ~ X))
X emmean SE df asymp.LCL asymp.UCL
A 2.20 1.054 Inf 0.131 4.26
B 1.39 0.791 Inf -0.163 2.94
C 0.00 0.632 Inf -1.240 1.24
D 0.00 0.632 Inf -1.240 1.24
Results are given on the logit (not the response) scale.
Confidence level used: 0.95
(em2 <- emmeans(m2, ~ X))
X emmean SE df asymp.LCL asymp.UCL
A 2.20 1.054 Inf 0.131 4.26
B 1.39 0.791 Inf -0.163 2.94
C 0.00 0.632 Inf -1.240 1.24
D 0.00 0.632 Inf -1.240 1.24
Results are given on the logit (not the response) scale.
Confidence level used: 0.95 一切都很好。但是,当我添加type = response参数时,除了置信区间不同(比较下面的两个输出)外,一切看起来都很好:
(em3 <- emmeans(m1, ~ X, type = "response"))
X response SE df asymp.LCL asymp.UCL
A 0.9 0.0949 Inf 0.714 1.09
B 0.8 0.1265 Inf 0.552 1.05
C 0.5 0.1581 Inf 0.190 0.81
D 0.5 0.1581 Inf 0.190 0.81
Unknown transformation ">": no transformation done
Confidence level used: 0.95
(em4 <- emmeans(m2, ~ X, type = "response"))
X prob SE df asymp.LCL asymp.UCL
A 0.9 0.0949 Inf 0.533 0.986
B 0.8 0.1265 Inf 0.459 0.950
C 0.5 0.1581 Inf 0.225 0.775
D 0.5 0.1581 Inf 0.225 0.775
Confidence level used: 0.95
Intervals are back-transformed from the logit scale 我看到第一个输出(Unknown transformation ">": no transformation done)中有一个警告,但是为什么它只影响置信区间?
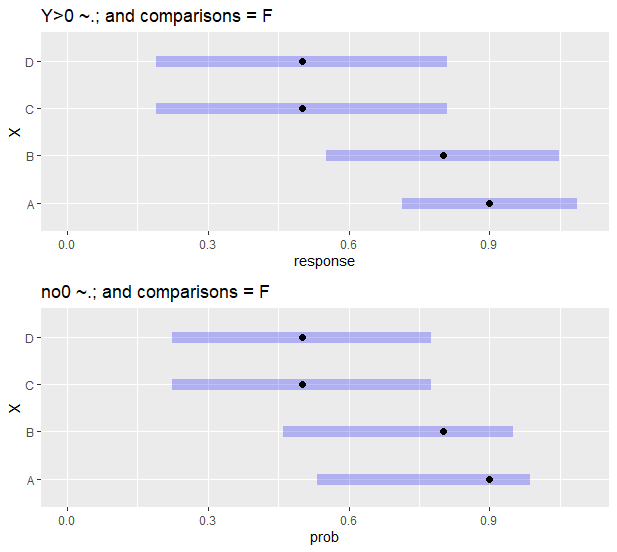
另一个有趣的观察是,当我绘制is表示对象时,没有的,plot()函数中的comparisons = T参数,它用不同的置信区间匹配上面的em3和em4输出:
p1 <- plot(em3, comparisons = F) + scale_x_continuous(limits = c(0,1.1)) + ggtitle("Y>0 ~.; and comparisons = F")
p2 <- plot(em4, comparisons = F) + scale_x_continuous(limits = c(0,1.1)) + ggtitle("no0 ~.; and comparisons = F")
gridExtra::grid.arrange(p1, p2, nrow = 2)
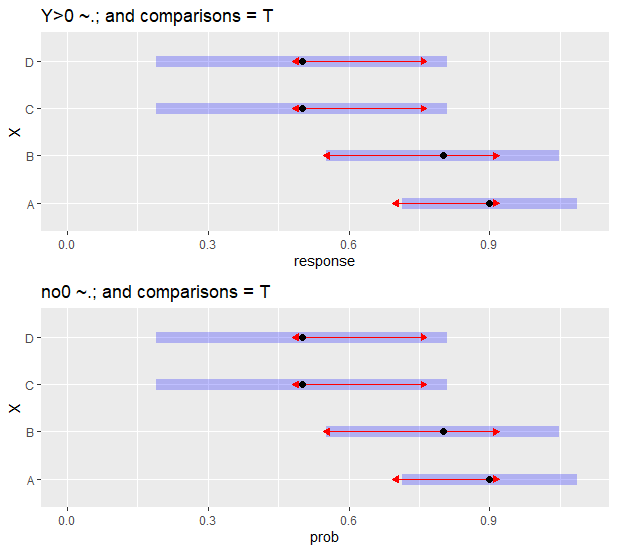
但是当我添加comparisons = T参数时,置信区间现在是相同的,但是两者都匹配基于模型中Y>0规范的模型(参见m3和em3)。
p3 <- plot(em3, comparisons = T) + scale_x_continuous(limits = c(0,1.1)) + ggtitle("Y>0 ~.; and comparisons = T")
p4 <- plot(em4, comparisons = T) + scale_x_continuous(limits = c(0,1.1))+ ggtitle("no0 ~.; and comparisons = T")
gridExtra::grid.arrange(p3, p4, nrow = 2)
这是有点冗长,但我的问题归结为:
在使用Y>0 ~ X emmeans**,时,可以组合使用模型规范,还是应该首先为此创建一个单独的变量?**
回答 1
Stack Overflow用户
发布于 2019-12-13 16:19:19
正在发生的情况是, is意味着允许同时存在响应转换和链接函数的情况。例如,当你用伽马族、逆链接和平方根响应变换来拟合一个模型时,这是很方便的。但是,在这种情况下,>被视为响应转换:
> emm1 <- emmeans(m1, "X")
> str(emm1)
'emmGrid' object with variables:
X = A, B, C, D
Transformation: “logit”
Additional response transformation: “>” 当您指定type = "response"时,summary.emmGrid()尝试撤消这两个转换--也就是说,尝试将其放到Y规模上。您可以撤消链接函数,如下所示:
> confint(emm1, type = "unlink")
X response SE df asymp.LCL asymp.UCL
A 0.9 0.0949 Inf 0.533 0.986
B 0.8 0.1265 Inf 0.459 0.950
C 0.5 0.1581 Inf 0.225 0.775
D 0.5 0.1581 Inf 0.225 0.775
Confidence level used: 0.95
Intervals are back-transformed from the logit scale ..。或者通过删除第二个转换:
> emm1a <- update(emm1, tran2 = NULL)
> confint(emm1a, type = "response")
X response SE df asymp.LCL asymp.UCL
A 0.9 0.0949 Inf 0.533 0.986
B 0.8 0.1265 Inf 0.459 0.950
C 0.5 0.1581 Inf 0.225 0.775
D 0.5 0.1581 Inf 0.225 0.775
Confidence level used: 0.95
Intervals are back-transformed from the logit scale 在这两种情况下,这里的置信区间都是在链接标度上计算的,然后再进行反变换.您在这里看到的其他置信限是在以下步骤反转后得到的,即使用反变换结果的标准误差:
> confint(regrid(emm1, transform = "unlink"))
X response SE df asymp.LCL asymp.UCL
A 0.9 0.0949 Inf 0.714 1.09
B 0.8 0.1265 Inf 0.552 1.05
C 0.5 0.1581 Inf 0.190 0.81
D 0.5 0.1581 Inf 0.190 0.81
Results are given on the > (not the response) scale.
Confidence level used: 0.95 我将考虑是否可以进行更改,从而可靠地确定响应转换显然不是有意的。
https://stackoverflow.com/questions/59316920
复制相似问题

