使用heatmaply创建不需要转换的原始值/数据的热图?
使用heatmaply创建不需要转换的原始值/数据的热图?
提问于 2020-04-25 02:27:00
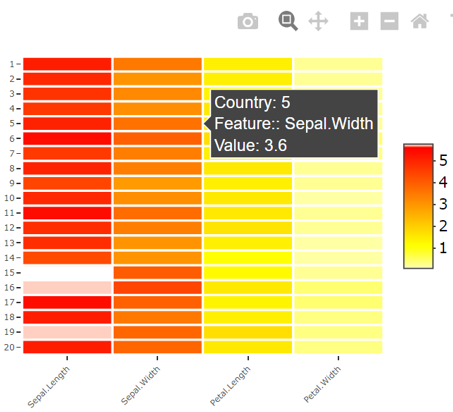
使用heatmaply::heatmaply()创建热图时,对原始数据执行一些转换。这一点在下面的示例中很明显,其中提供了一些虹膜数据集,但是在热图上徘徊显示了一些负值(这些值不在提供给heatmaply()的原始数据中),而旁边的颜色条也显示了负值。
https://i.stack.imgur.com/Mt0vGm.png
library(heatmaply)
mat <- iris[1:20, 1:4] %>% as.matrix
p <- heatmaply(mat,
dendrogram = "none",
xlab = "", ylab = "",
main = "",
scale = "column",
margins = c(60,100,40,20),
grid_color = "white",
grid_width = 0.00001,
titleX = FALSE,
hide_colorbar = FALSE,
branches_lwd = 0.1,
label_names = c("Country", "Feature:", "Value"),
fontsize_row = 5, fontsize_col = 5,
labCol = colnames(mat),
labRow = rownames(mat),
heatmap_layers = theme(axis.line=element_blank()),
colors = rev(c("000000", heat.colors(30)[1:28]))
)
p对iris[1:20, 1:4]的不同规模
https://i.stack.imgur.com/nuMcn.png
问题
如何使heatmaply::heatmaply()完全基于提供的原始值(不进行转换)生成基于热图的heatmaply::heatmaply()?
注:开放使用其他软件包/功能/建议
回答 1
Stack Overflow用户
回答已采纳
发布于 2020-04-25 03:27:07
删除scale参数-默认为“无”。或者改为“无”。
heatmaply(mat,
dendrogram = "none",
xlab = "", ylab = "",
main = "",
# scale = "column", # <-- remove this line. Or change to "none"
margins = c(60,100,40,20), ...
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/61420232
复制相关文章
相似问题

