形象化多组事件存在/缺席的最佳方法
我有一个数据集,记录了40种特定基因突变的存在/缺失,并将正常组织(如肺组织)与肿瘤(如肺肿瘤)的20种组织类型进行了比较。我很难找到最好的方法来可视化这些数据。
数据的一个子集:
Gene Lung_Normal Lung_Cancer Skin_Normal Skin_Cancer Brain_Normal Brain_Cancer
Gene_1 TRUE TRUE TRUE TRUE TRUE TRUE
Gene_2 TRUE TRUE TRUE TRUE TRUE TRUE
Gene_3 FALSE TRUE FALSE FALSE FALSE FALSE
Gene_4 FALSE FALSE FALSE FALSE FALSE FALSE
Gene_5 FALSE TRUE FALSE FALSE FALSE TRUE
Gene_6 FALSE FALSE TRUE TRUE TRUE TRUE
Gene_7 FALSE FALSE FALSE TRUE FALSE FALSE
Gene_8 FALSE FALSE FALSE TRUE FALSE TRUE
Gene_9 FALSE TRUE FALSE FALSE FALSE FALSE
Gene_10 FALSE FALSE FALSE TRUE FALSE TRUE我们要传达的关键信息是,虽然相同的3-4个基因在正常组织中经常发生突变,但每个肿瘤都有更多的附加基因发生突变,肿瘤中也有更多的多样性。我可以把它放在这样的桌子上,但我很想找到一种很好的方法来清晰地将信息可视化。
我想试着制作一个数字,就像一个马戏团的情节,一个圆圈有两个代表所有数据的圆环。内环为正常组织,外环为癌组织,每一节段均包含内环上的相关正常组织,外环上相应的癌组织。每个基因都有颜色编码,只有在变异时才能显示出来。因此,对于所有正常组织,该片段将显示2-3个突变基因的颜色,而外部癌症片段将显示更多的颜色片段,代表更多的突变。
然而,我还没有找到一个绘图软件,可以创建这样的可视化。有谁知道有什么方法可以让你像这样想象吗?即使把我指向一个R包也是很有帮助的。我已经查看过circos和雷达图,但我没有找到一个可以使我想到的可视化类型的包,只显示在每种情况下发生的事件。
如果有人认为另一种可视化方式可以代表这些数据,请告诉我,我很乐意考虑清楚地代表数据的替代方案。
提前谢谢你。
回答 3
Stack Overflow用户
发布于 2020-09-28 14:06:56
不知道这是不是你要找的,但我试了一下。另外,从上面的描述中我也不完全确定你想对不同类型的细胞做什么--肺,皮肤,大脑?如果这不是您想要的,也许您可以发布一张预期输出应该是什么样子的绘图。
在下图中,内环是正常细胞,外环是癌细胞。我在这里的回答得益于这个职位。
## Make the data
tib <- tibble::tribble(
~Gene, ~Lung_Normal, ~Lung_Cancer, ~Skin_Normal, ~Skin_Cancer, ~Brain_Normal, ~Brain_Cancer,
"Gene_1", TRUE , TRUE , TRUE , TRUE , TRUE , TRUE,
"Gene_2", TRUE, TRUE, TRUE, TRUE, TRUE, TRUE,
"Gene_3", FALSE , TRUE , FALSE , FALSE , FALSE , FALSE,
"Gene_4", FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
"Gene_5", FALSE , TRUE , FALSE , FALSE , FALSE , TRUE,
"Gene_6", FALSE, FALSE, TRUE, TRUE, TRUE, TRUE,
"Gene_7", FALSE , FALSE , FALSE , TRUE , FALSE , FALSE,
"Gene_8", FALSE, FALSE, FALSE, TRUE, FALSE, TRUE,
"Gene_9", FALSE , TRUE , FALSE , FALSE , FALSE , FALSE,
"Gene_10", FALSE, FALSE, FALSE, TRUE, FALSE, TRUE)
library(tidyr)
library(dplyr)
## Re-arrange into long format
tib <- tib %>%
pivot_longer(cols=-Gene, names_pattern="(.*)_(.*)", names_to=c("type", ".value")) %>%
pivot_longer(c(Normal, Cancer), names_to = "diag", values_to="val") %>%
# code colors as the gene if it's mutated, otherwise Unmutated
mutate(f = case_when(val ~ Gene, TRUE ~ "Unmutated")) %>%
group_by(Gene, f, diag) %>%
summarise(s = n()) %>%
mutate(diag = factor(diag, levels=c("Normal", "Cancer")),
f = factor(f, levels=c(paste("Gene", c(1,2,6,3,5,7,8,9,10,4), sep="_"), "Unmutated")))
library(ggplot2)
library(RColorBrewer)
ggplot(tib, aes(x=diag,
y = s,
fill=f)) +
geom_bar(stat="identity") +
coord_polar("y") +
theme_void() +
scale_fill_manual(values=c(brewer.pal(9, "Paired"), "gray75")) +
labs(fill = "Mutations")
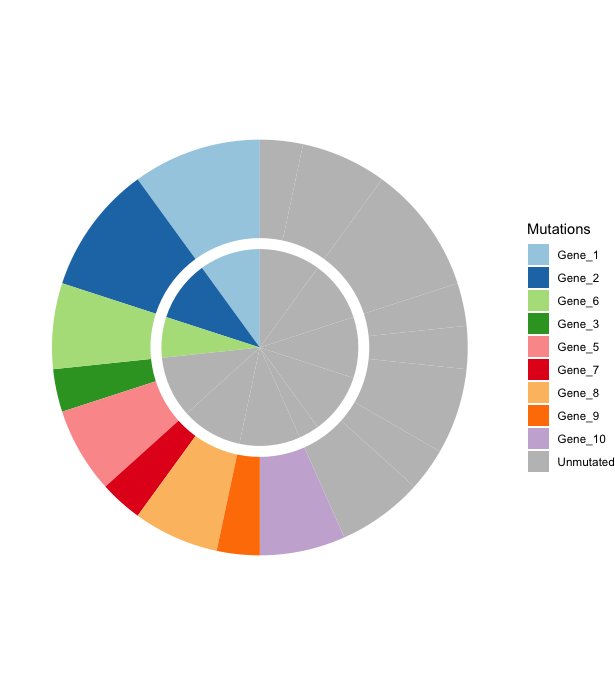
编辑
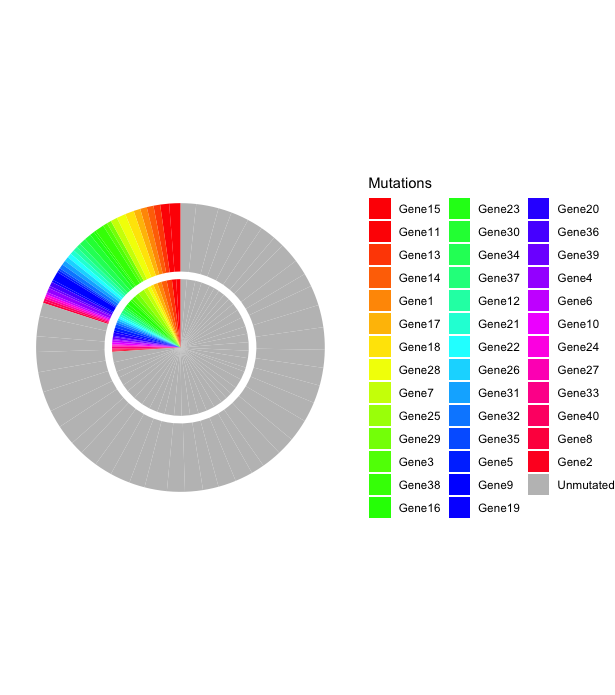
以下是Allan建议的数据的样子。这种方法不太适合有很多颜色的需要,这将降低情节的可读性。
df <- structure(list(genes = c("Gene1", "Gene2", "Gene3", "Gene4",
"Gene5", "Gene6", "Gene7", "Gene8", "Gene9", "Gene10", "Gene11",
"Gene12", "Gene13", "Gene14", "Gene15", "Gene16", "Gene17", "Gene18",
"Gene19", "Gene20", "Gene21", "Gene22", "Gene23", "Gene24", "Gene25",
"Gene26", "Gene27", "Gene28", "Gene29", "Gene30", "Gene31", "Gene32",
"Gene33", "Gene34", "Gene35", "Gene36", "Gene37", "Gene38", "Gene39",
"Gene40"), bone_cancer = c(FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, FALSE,
TRUE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE), bone_normal = c(FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
TRUE, FALSE, TRUE), brain_cancer = c(TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE), brain_normal = c(FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE,
FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
TRUE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE), breast_cancer = c(FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, TRUE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE), breast_normal = c(TRUE,
FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, TRUE, FALSE,
FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE,
FALSE), colon_cancer = c(FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE), colon_normal = c(FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
TRUE, TRUE, FALSE), kidney_cancer = c(FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE),
kidney_normal = c(FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, TRUE,
TRUE, FALSE, TRUE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, TRUE, TRUE, TRUE, FALSE, FALSE, TRUE,
TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE), liver_cancer = c(FALSE,
FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE), liver_normal = c(TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE,
TRUE, TRUE, FALSE, TRUE, FALSE, FALSE, TRUE, TRUE, FALSE,
TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE,
FALSE), lung_cancer = c(TRUE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE),
lung_normal = c(FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE, TRUE, TRUE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE), prostate_cancer = c(TRUE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, TRUE, FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, TRUE,
FALSE, TRUE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, TRUE), prostate_normal = c(TRUE, FALSE, FALSE,
FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE), skin_cancer = c(FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, FALSE, FALSE), skin_normal = c(TRUE,
FALSE, TRUE, TRUE, TRUE, FALSE, TRUE, FALSE, FALSE, TRUE,
FALSE, FALSE, TRUE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, TRUE, FALSE, TRUE, TRUE, TRUE, TRUE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE,
FALSE, FALSE, FALSE), thyroid_cancer = c(FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, TRUE, TRUE, FALSE, TRUE, TRUE,
FALSE, FALSE, TRUE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE), thyroid_normal = c(FALSE, FALSE, TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE)),
class = "data.frame", row.names = c(NA, 40L))
names(df)[1] <- "Gene"
tib <- df %>%
pivot_longer(cols=-Gene, names_pattern="(.*)_(.*)", names_to=c("type", ".value")) %>%
pivot_longer(c(normal, cancer), names_to = "diag", values_to="val") %>%
# code colors as the gene if it's mutated, otherwise Unmutated
mutate(f = case_when(val ~ Gene, TRUE ~ "Unmutated")) %>%
group_by(Gene, f, diag) %>%
summarise(s = n()) %>%
ungroup() %>%
group_by(Gene) %>%
mutate(diag = factor(diag, levels=c("normal", "cancer")))
levs <- tib %>%
dplyr::select(f, s) %>%
summarise(pct_mutated = sum(s*(f!= "Unmutated"))/sum(s)) %>%
arrange(-pct_mutated) %>%
dplyr::select(Gene) %>%
pull()
tib<- tib %>%
mutate(f = factor(f, levels=c(levs, "Unmutated")))
library(ggplot2)
library(RColorBrewer)
ggplot(tib, aes(x=diag,
y = s,
fill=f)) +
geom_bar(stat="identity") +
coord_polar("y") +
theme_void() +
scale_fill_manual(values=c(rainbow(length(levels(tib$f))-1), "gray75")) +
labs(fill = "Mutations")
Stack Overflow用户
发布于 2020-09-28 14:38:24
另一种选择是热图。你可以通过改变癌症与正常的比例来做到这一点,或者调整填充来反映癌症的突变,只有正常的组织,两者都不能。
对于这两种情况,首先需要对数据进行整形:
选项1
library(tidyr)
library(dplyr)
library(ggplot2)
df %>%
pivot_longer(-1) %>%
separate(name, into = c("tissue", "state"), sep = "_") %>%
mutate(genes = factor(genes, paste0("Gene", 1:40))) %>%
ggplot(aes(tissue, genes, fill = value)) + geom_tile(color = "black") +
facet_grid(.~state) +
scale_x_discrete(guide = guide_axis(n.dodge = 2))
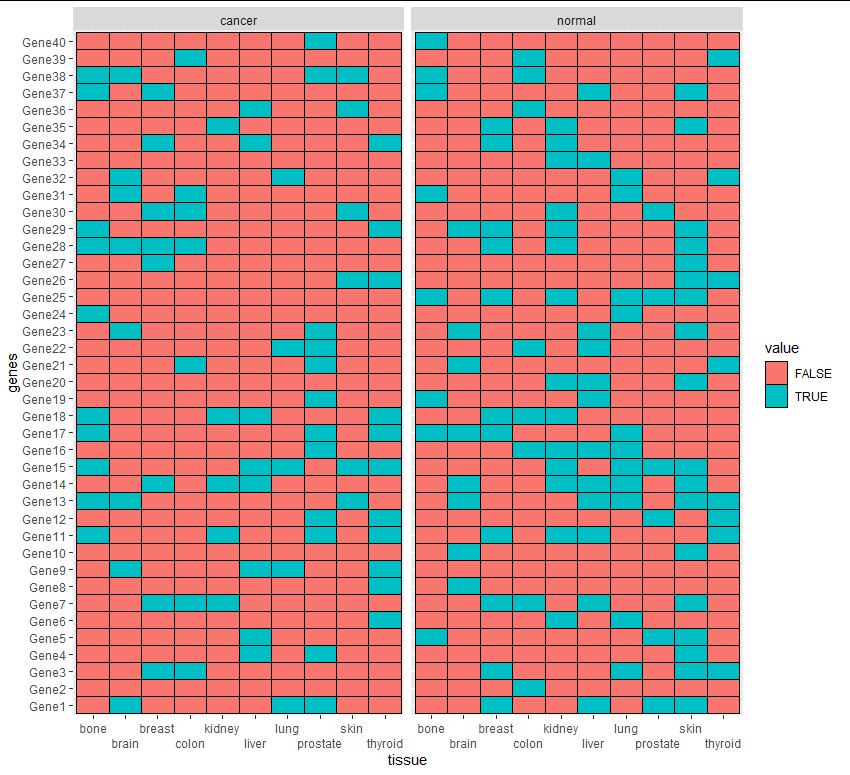
选项2
df %>%
pivot_longer(-1) %>%
separate(name, into = c("tissue", "state"), sep = "_") %>%
mutate(genes = factor(genes, paste0("Gene", 1:40))) %>%
group_by(genes, tissue) %>%
summarize(mutations = factor(2 + diff(value) + 2 * all(value))) %>%
ggplot(aes(tissue, genes, fill = mutations)) + geom_tile(color = "black") +
scale_x_discrete(guide = guide_axis(n.dodge = 2)) +
scale_fill_discrete(labels = c("Neither", "Cancer only", "Healthy only", "Both"))
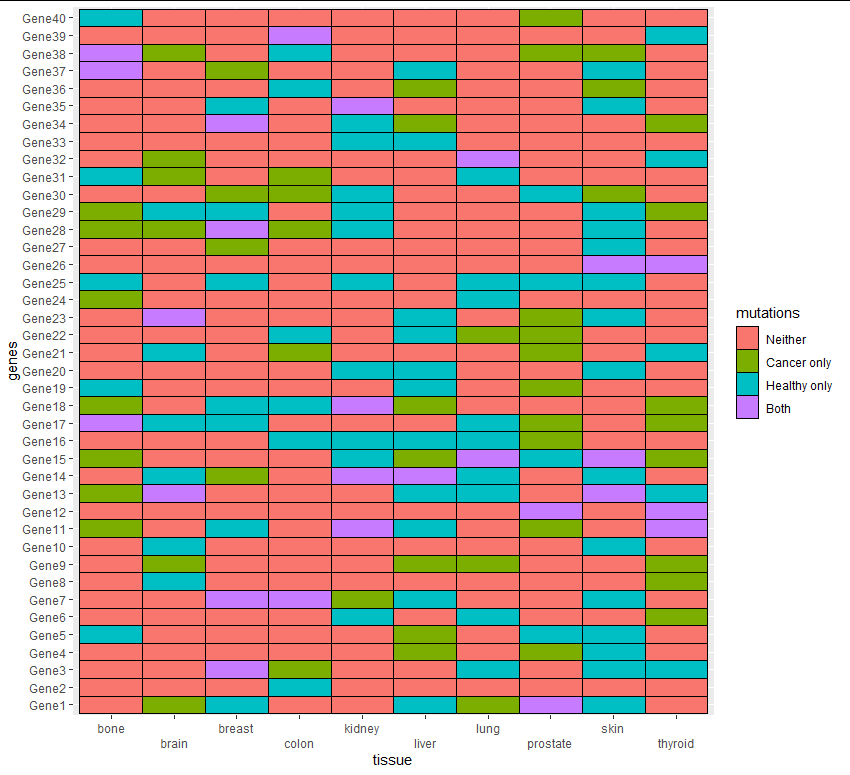
对于这些数据,我使用了类似于您的数据结构的示例数据:
df <- structure(list(genes = c("Gene1", "Gene2", "Gene3", "Gene4",
"Gene5", "Gene6", "Gene7", "Gene8", "Gene9", "Gene10", "Gene11",
"Gene12", "Gene13", "Gene14", "Gene15", "Gene16", "Gene17", "Gene18",
"Gene19", "Gene20", "Gene21", "Gene22", "Gene23", "Gene24", "Gene25",
"Gene26", "Gene27", "Gene28", "Gene29", "Gene30", "Gene31", "Gene32",
"Gene33", "Gene34", "Gene35", "Gene36", "Gene37", "Gene38", "Gene39",
"Gene40"), bone_cancer = c(FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, FALSE,
TRUE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE), bone_normal = c(FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
TRUE, FALSE, TRUE), brain_cancer = c(TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE), brain_normal = c(FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE,
FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
TRUE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE), breast_cancer = c(FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, TRUE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE), breast_normal = c(TRUE,
FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, TRUE, FALSE,
FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE,
FALSE), colon_cancer = c(FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE), colon_normal = c(FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
TRUE, TRUE, FALSE), kidney_cancer = c(FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE),
kidney_normal = c(FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, TRUE,
TRUE, FALSE, TRUE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, TRUE, TRUE, TRUE, FALSE, FALSE, TRUE,
TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE), liver_cancer = c(FALSE,
FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE), liver_normal = c(TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE,
TRUE, TRUE, FALSE, TRUE, FALSE, FALSE, TRUE, TRUE, FALSE,
TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE,
FALSE), lung_cancer = c(TRUE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE),
lung_normal = c(FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE, TRUE, TRUE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE), prostate_cancer = c(TRUE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, TRUE, FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, TRUE,
FALSE, TRUE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, TRUE), prostate_normal = c(TRUE, FALSE, FALSE,
FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE), skin_cancer = c(FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, TRUE, FALSE, FALSE), skin_normal = c(TRUE,
FALSE, TRUE, TRUE, TRUE, FALSE, TRUE, FALSE, FALSE, TRUE,
FALSE, FALSE, TRUE, TRUE, TRUE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, TRUE, FALSE, TRUE, TRUE, TRUE, TRUE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, TRUE,
FALSE, FALSE, FALSE), thyroid_cancer = c(FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, TRUE, TRUE, FALSE, TRUE, TRUE,
FALSE, FALSE, TRUE, FALSE, TRUE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, TRUE, FALSE,
FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE), thyroid_normal = c(FALSE, FALSE, TRUE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, TRUE, TRUE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, TRUE, FALSE)), class = "data.frame", row.names = c(NA,
40L))Stack Overflow用户
发布于 2020-09-28 16:11:04
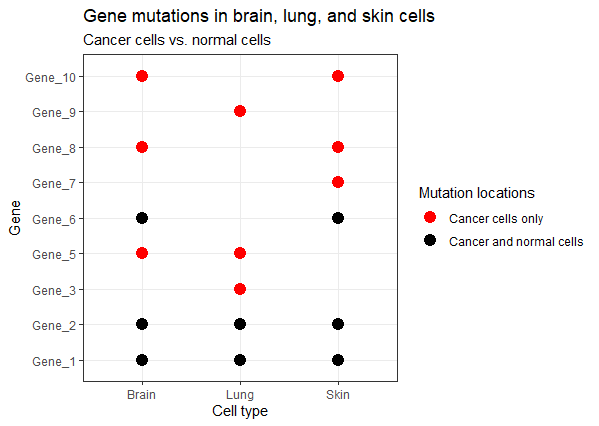
这是另一种选择。类似于热图,而是使用彩色点。我也选择隐藏非突变,因为我认为我们更感兴趣的可视化突变发生的地方。如果FALSE值也是有色的,它会增加认知负荷,为我们的大脑提供一个额外的解释重点。
# Creating the data.
gene_data <- structure(list(Gene = c("Gene_1", "Gene_2", "Gene_3", "Gene_4",
"Gene_5", "Gene_6", "Gene_7", "Gene_8", "Gene_9", "Gene_10"),
Lung_Normal = c(TRUE, TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE), Lung_Cancer = c(TRUE, TRUE, TRUE, FALSE,
TRUE, FALSE, FALSE, FALSE, TRUE, FALSE), Skin_Normal = c(TRUE,
TRUE, FALSE, FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE
), Skin_Cancer = c(TRUE, TRUE, FALSE, FALSE, FALSE, TRUE,
TRUE, TRUE, FALSE, TRUE), Brain_Normal = c(TRUE, TRUE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, FALSE), Brain_Cancer = c(TRUE,
TRUE, FALSE, FALSE, TRUE, TRUE, FALSE, TRUE, FALSE, TRUE)), row.names = c(NA,
-10L), class = c("tbl_df", "tbl", "data.frame"))稍微处理一下数据:
mutation_data <- gene_data %>%
pivot_longer(-Gene, values_to = "Mutation") %>% # making long form
separate(name, into = c("Type", "Status")) %>% # splitting cell type and cancer status
mutate(Mutation = as.numeric(Mutation), # cleaning data for plotting
Type = factor(Type, levels = c("Cancer", "Normal")),
Gene = factor(Gene, levels = paste0("Gene_", 1:10))) %>% #genes appear in order
filter(Mutation == 1) # Removing FALSE for cleaner plot建造地块
ggplot() +
# plotting all cancer mutations in red
geom_point(data = mutation_data %>% filter(Status == "Cancer"),
aes(x = Type, y = Gene, color = Status), size = 4) +
# plotting all mutations present in normal and cancer in black over top of the red
geom_point(data = mutation_data %>% filter(Status == "Normal"),
aes(x = Type, y = Gene, color = Status), size = 4) +
# formatting the color scale and legend
scale_color_manual(name = "Mutation locations", values = c("Cancer" = "Red", "Normal" = "Black"),
labels = c("Cancer cells only", "Cancer and normal cells")) +
# providing title, subtitle, and x label
labs(title = "Gene mutations in brain, lung, and skin cells",
subtitle = "Cancer cells vs. normal cells",
x = "Cell type") +
theme_bw() # setting a theme I like
https://stackoverflow.com/questions/64103097
复制相似问题

