在R中从非选定的行名中删除NAs
在R中从非选定的行名中删除NAs
提问于 2021-03-31 18:29:24
我有一组我想要画的基因,我已经重新命名了。但当我策划他们的时候,我也会得到那些没有被选中的人的NAs。我怎样才能删除只显示我所选基因的NAs?
library(pheatmap)
set.seed(2020)
mat = matrix(rnorm(200),20,10)
rownames(mat) = paste0("g",1:20)
rownames(mat)[rownames(mat) == "g2"] <- "abc"
rownames(mat)[rownames(mat) == "g4"] <- "pqr"
rownames(mat)[rownames(mat) == "g6"] <- "zzz"
selected_genes <- c("abc","pqr", "zzz")
obj = pheatmap(mat,cluster_rows = T, scale = 'row', show_rownames = T, labels_row = selected_genes)
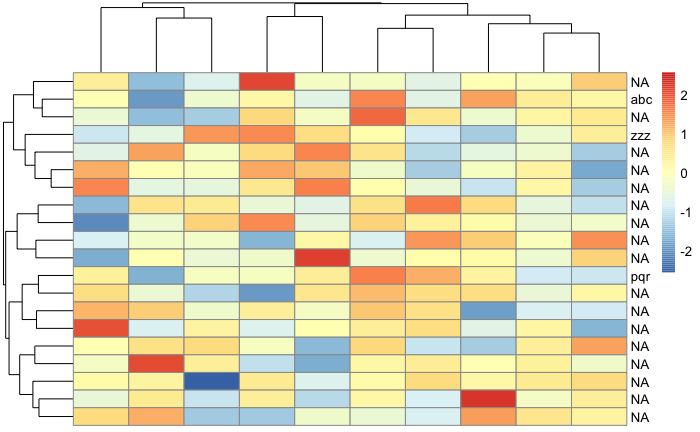
我见过你也可以做这样的事:
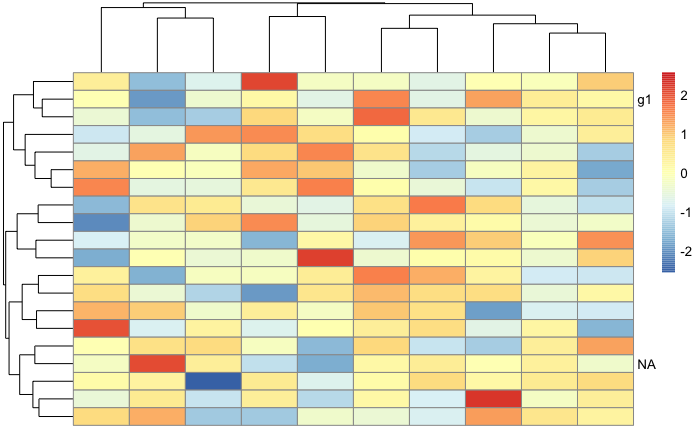
labRow <- c(row.names(mat)[1], rep('', length(row.names(mat))-2)) # Selected row2 as example
pheatmap(mat,cluster_rows = T, scale = 'row', show_rownames = T, labels_row = labRow)
回答 1
Stack Overflow用户
回答已采纳
发布于 2021-11-05 00:03:22
由于您已经发现将label_row参数填充为非打印值,所以您应该尝试:
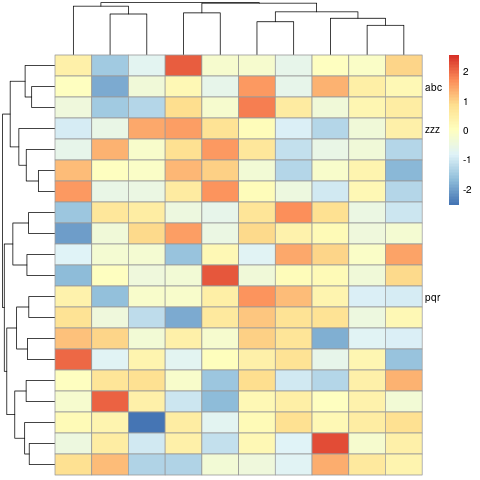
obj = pheatmap(mat,cluster_rows = T, scale = 'row', show_rownames = T,
labels_row = c(selected_genes,rep(" ",17) ) )
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/66893332
复制相关文章
相似问题

