Keras并不包括所有的类
我已经做了一个模型,它被训练来预测一个从34-63 (无十进制数)的数字,总共是30个潜在的输出。
当我运行模型时,它会抱怨并希望我在我的最后一层中放入15,我已经理解这应该是输出的数量。
经过培训后,我还会在终端中得到以下输出:
ValueError: y_true and y_pred contain different number of classes 7, 16. Please provide the true labels explicitly through the labels argument. Classes found in y_true: [51 52 53 54 56 59 63]然后,当我运行模型时:
prediction = model.predict(test)
print(model.predict(test))
print(np.argmax(model.predict(test), axis=-1))我得到:
警告:tensorflow:在过去19次对的调用中,有6次调用触发了tf.function回溯。跟踪非常昂贵,过多的跟踪可能是由于(1)在循环中反复创建@tf.function,(2)传递形状不同的张量,(3)传递Python对象而不是张量。对于(1),请在循环之外定义您的@tf.function。对于(2),@tf.function有experimental_relax_shapes=True选项,它可以放松参数形状,从而避免不必要的回溯。有关(3)的详细信息,请参阅https://www.tensorflow.org/guide/function#controlling_retracing和https://www.tensorflow.org/api_docs/python/tf/function。
[[0.00006836 0.33038142 0.22732003 0.03764497 0.22742009 0.01213347
0.16344884 0.00000338 0.0012028 0.00014862 0.00000717 0.00017032
0.00000437 0.00001909 0.00002712]]
[1]我猜想矩阵应该是所有的大小,但只有15个。我查看了我的数据集,所有的类至少有3个实例,所以它们应该包括在培训中。
#更新我已经将模型包含在下面
np.set_printoptions(suppress=True)
pd.set_option("display.max_rows", None, "display.max_columns", None)
config = ConfigProto()
config.gpu_options.allow_growth = True
session = InteractiveSession(config=config)
Labeldata = ['Output_Label']
RelevantFeatures = ['column A','column B','column X']
RelevantFeaturesandlabel = ['column A','column B','column X','Output_Label']
PATH = 'Training_Data.xlsx'
PATHVa = 'Validation_Data.xlsx'
Full_Data = pd.read_excel(PATH)
ValidationFull = pd.read_excel(PATHVa)
# Which range of outputs should be included
Full_Data = Full_Data[(Full_Data['Output_Label'] >= 34) & (Full_Data['Output_Label'] <= 70)]
ValidationFull = ValidationFull[(ValidationFull['Output_Label'] >= 34) & (ValidationFull['Output_Label'] <= 70)]
FeatureDatadf = Full_Data.filter(items = RelevantFeatures, axis = 1)
Validation = ValidationFull.filter(items = RelevantFeatures, axis = 1)
ValidationLabel = ValidationFull.filter(items = Labeldata, axis = 1)
FeatureData = pd.DataFrame(StandardScaler().fit_transform(FeatureDatadf))
Validation = pd.DataFrame(StandardScaler().fit_transform(Validation))
FeatureData = FeatureData.apply(pd.to_numeric, errors='coerce')
FeatureData = FeatureData.to_numpy()
Validation = Validation.to_numpy()
#Standardisation
LabelData = Full_Data.filter(items = Labeldata, axis=1)
LabelData = LabelData.apply(pd.to_numeric, errors='coerce')
dummies = pd.get_dummies(['34','35','36','37','38','39','40','41','42','43','44','45','46','47','48','49','50','51','52','53','54','55','56','57','58','59','60','61','62','63'], prefix = 'Size')
dummies = pd.get_dummies(LabelData['Output_Label'],prefix = 'Size')
LabelData = dummies.to_numpy()
# Split the sets up
Feature_train, Feature_test, Label_train, Label_test = train_test_split(FeatureData, LabelData, test_size=0.2)
# Model
model = Sequential()
model.add(Dense(26, activation = LeakyReLU(alpha=1), input_dim = 26,activity_regularizer=regularizers.l1(1e-4),use_bias=False))#209))
model.add(Dense(26,RandomFourierFeatures(output_dim=1024, scale=10.0, kernel_initializer="gaussian"),use_bias=False))
model.add(Dense(26,RandomFourierFeatures(output_dim=1024, scale=10.0, kernel_initializer="gaussian"),use_bias=False))
model.add(Dense(26,RandomFourierFeatures(output_dim=1024, scale=10.0, kernel_initializer="gaussian"),use_bias=False))
model.add(Dense(50,RandomFourierFeatures(output_dim=1024, scale=10.0, kernel_initializer="gaussian"),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26,Dropout(0.4),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26,Dropout(0.4),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26,Dropout(0.4),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26,Dropout(0.4),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1),use_bias=False))
model.add(Dense(26,GaussianNoise(stddev = 0.5),use_bias=False))
model.add(Dense(26,GaussianNoise(stddev = 0.5),use_bias=False))
model.add(Dense(26,GaussianNoise(stddev = 0.5),use_bias=False))
model.add(Dense(26,GaussianNoise(stddev = 0.5),use_bias=False))
model.add(Dense(26,GaussianNoise(stddev = 0.5),use_bias=False))
#model.add(Dense(25,Normalization(),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1), activity_regularizer=regularizers.l1(1e-4),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1), activity_regularizer=regularizers.l1(1e-4),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1), activity_regularizer=regularizers.l1(1e-4),use_bias=False))
model.add(Dense(26, activation = LeakyReLU(alpha=1), activity_regularizer=regularizers.l1(1e-4),use_bias=False))
model.add(Dense(15, activation = 'softmax',use_bias=False))#Output is the number of classes
#optimisation
opt = SGD(lr=0.001, momentum=0.9)
# Compile
model.compile(loss='categorical_crossentropy', optimizer=opt, metrics=['categorical_accuracy'])
history = model.fit(Feature_train, Label_train, validation_data=(Feature_test, Label_test), epochs=500, verbose=1)
# evaluate the model
_, train_acc = model.evaluate(Feature_train, Label_train, verbose=1)
_, test_acc = model.evaluate(Feature_test, Label_test, verbose=1)
print('Train: %.3f, Test: %.3f' % (train_acc, test_acc))
# plot loss during training
pyplot.subplot(211)
pyplot.title('Loss')
pyplot.plot(history.history['loss'], label='train')
pyplot.plot(history.history['val_loss'], label='test')
pyplot.legend()
# plot accuracy during training
pyplot.subplot(212)
pyplot.title('categorical_accuracy')
pyplot.plot(history.history['categorical_accuracy'], label='train')
pyplot.plot(history.history['val_categorical_accuracy'], label='test')
pyplot.legend()
pyplot.show()
prediction = model.predict(Validation)
Predictionsdf = pd.DataFrame(prediction, columns = dummies.columns)
Predictionsdf.to_excel('Preditions.xlsx', index = False)
#Save model
model.summary()
model.save(os.path.join('.', 'Output_Label.h5'))
score = metrics.log_loss(ValidationLabel, prediction)
print("Log loss score: {}".format(score))目前的数据结构如下:
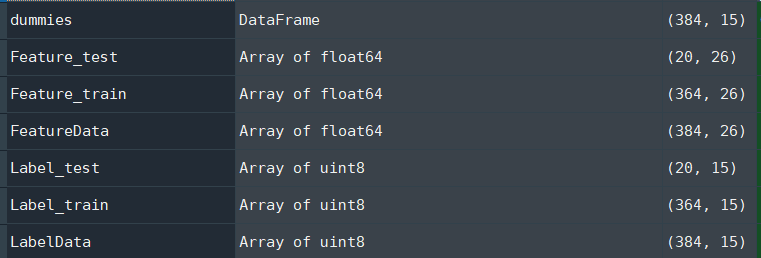
- 如何修复模型错误并使网络包含所有类
更新2)如何打印前三大类的预测:它们的准确性和名称?
更新后,我有以下代码用于打印预测及其准确性:
prediction = model_Chest.predict(test)
print(model_Chest.predict(test))
y_pred = model_Chest.predict(test)
# top_k has shape (N, k)
K=18
dummies = pd.get_dummies(['44','45', '46','47', '48','49', '50','51', '52','53', '54','55', '56','57', '58','59', '60', '61'], prefix = 'Size')
top_K = np.argsort(y_pred, -1)[:, :K]
names = dummies.columns.to_numpy()[top_K]
probs = np.take_along_axis(y_pred, top_K, -1)
print(names)
print(probs)应该是:
从44-61 smallest
- 打印出acc预测,然后从到打印出acc预测从最大到最小的。
但我明白:
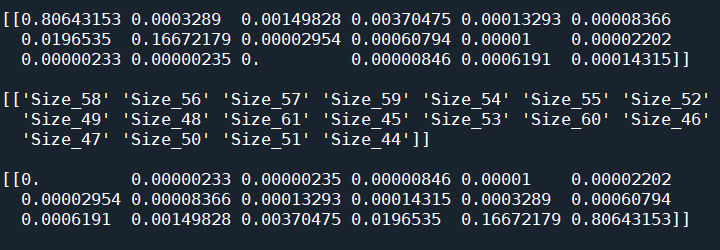
回答 1
Stack Overflow用户
发布于 2021-04-19 07:43:35
正如the comments中提到的,数据集中的类数只有15个,因此,为什么输出15个值是合适的。
要获得最高的k类概率,可以使用numpy.argsort,然后使用dataframe列获取类名:
y_pred = model.predict(x)
# top_k has shape (N, k)
top_k = np.argsort(y_pred, -1)[:, :k]
names = dummies.columns.to_numpy()[top_k]
probs = np.take_along_axis(y_pred, top_k, -1)然后,names包含x中每个实例的顶级k类的名称,probs包含相应的概率。
https://stackoverflow.com/questions/67140570
复制相似问题

