更改摘要函数通过浏览打印的顺序。
更改摘要函数通过浏览打印的顺序。
提问于 2019-03-18 15:15:36
我使用skimr,并在函数skim的摘要函数列表中添加了两个摘要函数(iqr_na_rm和median_na_rm)。但是,默认情况下,这些新的摘要函数(在skimmers文档中称为skimr文档)出现在表的末尾。相反,我希望median和iqr出现在mean和sd之后。
最后的目标是在这样的.Rmd报告中显示结果:
---
title: "Test"
output: html_document
---
```{r setup, include=FALSE}Knitr::opts_chunk$set(警告=假,
message = FALSE, echo = FALSE)## Test
```{r test, results = 'asis'}图书馆(Skimr)
图书馆(Dplyr)
图书馆(Ggplot2)
iqr_na_rm <-函数(X) IQR(x,na.rm = TRUE)
median_na_rm <-函数(X)中值(x,na.rm = TRUE)
Skim_with(p50= NULL,中位数= median_na_rm,iqr = iqr_na_rm),
integer = list(p50 = NULL, median = median_na_rm, iqr = iqr_na_rm))睡眠%>%
group_by(vore) %>%
脱脂(Sleep_total) %>%
凯布尔()
呈现的HTML:
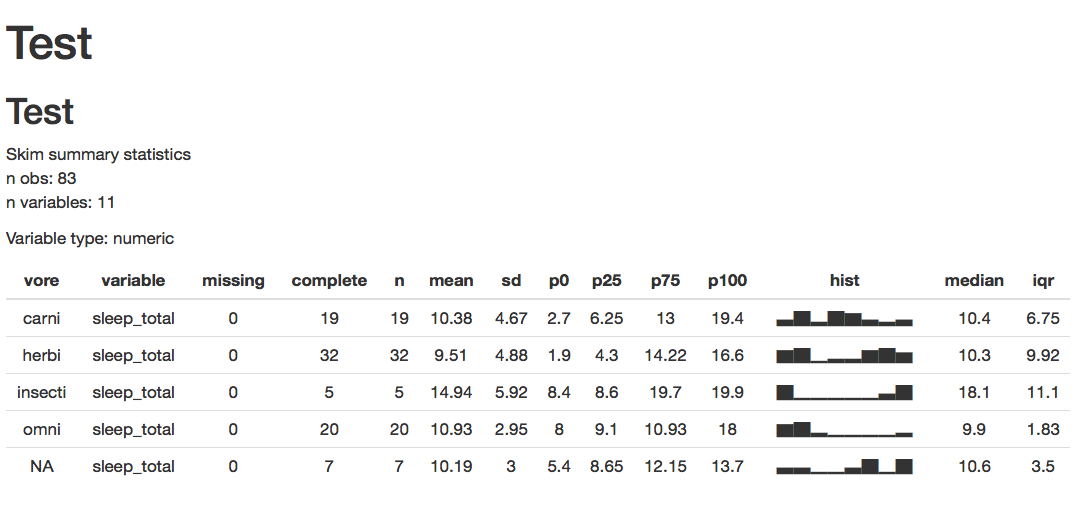
如您所见,median和iqr都是打印出来的,表的末尾是火花线直方图之后。我想把它们印在sd之后和p0之前。有可能吗?
回答 2
Stack Overflow用户
回答已采纳
发布于 2019-03-18 16:48:03
skim()输出中有两个部分。如果您想要控制数字部分,可以像这样使用skim_to_list。以另一种格式导出也更容易。
msleep %>%
group_by(vore) %>%
skim_to_list(sleep_total)%>%
.[["numeric"]]%>%
dplyr::select(vore,variable,missing,complete,n,mean,sd,
median,iqr,p0,p25,p75,p100,hist)
# A tibble: 5 x 14
vore variable missing complete n mean sd median iqr p0 p25 p75 p100 hist
* <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
1 carni sleep_total 0 19 19 10.38 4.67 10.4 " 6.75" 2.7 6.25 "13 " 19.4 ▃▇▂▇▆▃▂▃
2 herbi sleep_total 0 32 32 " 9.51" 4.88 10.3 " 9.92" 1.9 "4.3 " 14.22 16.6 ▆▇▁▂▂▆▇▅
3 insecti sleep_total 0 5 5 14.94 5.92 18.1 "11.1 " 8.4 "8.6 " "19.7 " 19.9 ▇▁▁▁▁▁▃▇
4 omni sleep_total 0 20 20 10.93 2.95 " 9.9" " 1.83" "8 " "9.1 " 10.93 "18 " ▆▇▂▁▁▁▁▂
5 NA sleep_total 0 7 7 10.19 "3 " 10.6 " 3.5 " 5.4 8.65 12.15 13.7 ▃▃▁▁▃▇▁▇编辑
根据注释中的要求添加kable()。
msleep %>%
group_by(vore) %>%
skim_to_list(sleep_total)%>%
.[["numeric"]]%>%
dplyr::select(vore,variable,missing,complete,n,mean,sd,median,iqr,p0,p25,p75,p100,hist)%>%
kable()
| vore | variable | missing | complete | n | mean | sd | median | iqr | p0 | p25 | p75 | p100 | hist |
|---------|-------------|---------|----------|----|-------|------|--------|------|-----|------|-------|------|----------|
| carni | sleep_total | 0 | 19 | 19 | 10.38 | 4.67 | 10.4 | 6.75 | 2.7 | 6.25 | 13 | 19.4 | ▃▇▂▇▆▃▂▃ |
| herbi | sleep_total | 0 | 32 | 32 | 9.51 | 4.88 | 10.3 | 9.92 | 1.9 | 4.3 | 14.22 | 16.6 | ▆▇▁▂▂▆▇▅ |
| insecti | sleep_total | 0 | 5 | 5 | 14.94 | 5.92 | 18.1 | 11.1 | 8.4 | 8.6 | 19.7 | 19.9 | ▇▁▁▁▁▁▃▇ |
| omni | sleep_total | 0 | 20 | 20 | 10.93 | 2.95 | 9.9 | 1.83 | 8 | 9.1 | 10.93 | 18 | ▆▇▂▁▁▁▁▂ |
| NA | sleep_total | 0 | 7 | 7 | 10.19 | 3 | 10.6 | 3.5 | 5.4 | 8.65 | 12.15 | 13.7 | ▃▃▁▁▃▇▁▇ |Stack Overflow用户
发布于 2019-03-21 03:15:04
下面是另一个使用append=FALSE选项的选项。
library(skimr)
library(dplyr)
library(ggplot2)
iqr_na_rm <- function(x) IQR(x, na.rm = TRUE)
median_na_rm <- function(x) median(x, na.rm = TRUE)
my_skimmers <- list(n = length, missing = n_missing, complete = n_complete,
mean = mean.default, sd = purrr::partial(sd, na.rm = TRUE),
median = median_na_rm, iqr = iqr_na_rm
)
skim_with(numeric = my_skimmers,
integer = my_skimmers, append = FALSE)
msleep %>%
group_by(vore) %>%
skim(sleep_total) %>%
kable()我没有放置所有的统计数据,但是您可以查看函数s.R和stats.R文件来查看如何定义各种统计信息。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/55224543
复制相关文章
相似问题

