factoextra::fviz_gap_stat()与factoextra::fviz_nbclust(df,method = "gap_stat")
factoextra::fviz_gap_stat()与factoextra::fviz_nbclust(df,method = "gap_stat")
提问于 2019-01-28 01:52:07
我试图弄清楚为什么这两个函数来自具有看似相似的参数的factoextra包(例如kmeans、kmeans)产生不同的结果。
library(cluster)
library(cluster.datasets)
library(tidyverse)
library(factoextra)
# load data and scale it
data("all.mammals.milk.1956")
mammals <- all.mammals.milk.1956 %>% select(-name)
mammals_scaled <- scale(mammals)第一种方法使用factoextra::clusGap()和factoextra::fviz_gap_stat()。
gap_stat <- clusGap(mammals_scaled, FUN = kmeans, K.max = 24, B = 50)
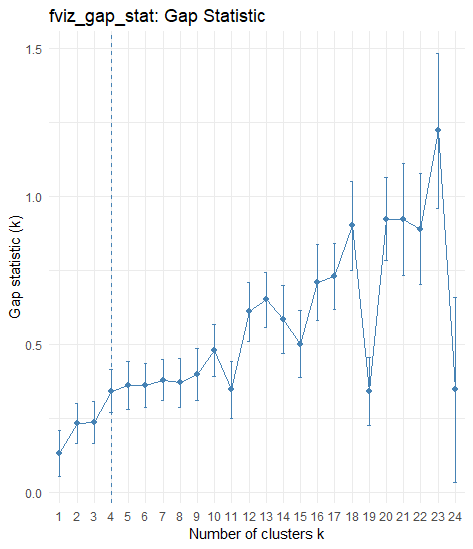
fviz_gap_stat(gap_stat) + theme_minimal() + ggtitle("fviz_gap_stat: Gap Statistic")
第二种方法使用factoextra::fviz_nbclust(),其中
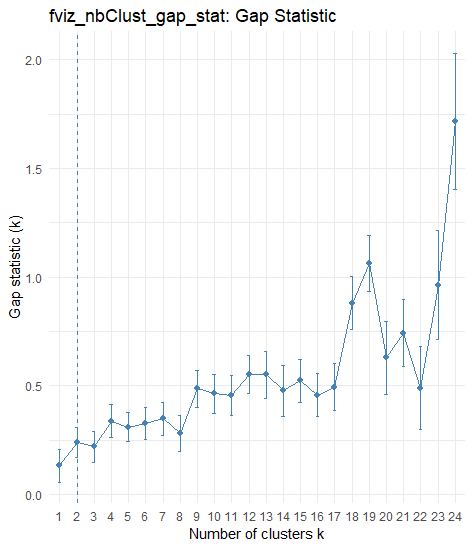
fviz_nbclust(mammals_scaled, kmeans, method = "gap_stat", k.max = 24, nboot = 50) + theme_minimal() + ggtitle("fviz_nbClust_gap_stat: Gap Statistic")
我认为这可能是来自clusGap()的clusGap()选项,但是当我使用jimhester/lookup用下面的代码读取fviz_nbclust()的源代码时,我找不到问题所在:
devtools::install_github("jimhester/lookup")
lookup::lookup(fviz_nbclust)
function (x, FUNcluster = NULL, method = c("silhouette", "wss",
"gap_stat"), diss = NULL, k.max = 10, nboot = 100, verbose = interactive(),
barfill = "steelblue", barcolor = "steelblue", linecolor = "steelblue",
print.summary = TRUE, ...)
{
set.seed(123)
if (k.max < 2)
stop("k.max must bet > = 2")
method = match.arg(method)
if (!inherits(x, c("data.frame", "matrix")) & !("Best.nc" %in%
names(x)))
stop("x should be an object of class matrix/data.frame or ",
"an object created by the function NbClust() [NbClust package].")
if (inherits(x, "list") & "Best.nc" %in% names(x)) {
best_nc <- x$Best.nc
if (class(best_nc) == "numeric")
print(best_nc)
else if (class(best_nc) == "matrix")
.viz_NbClust(x, print.summary, barfill, barcolor)
}
else if (is.null(FUNcluster))
stop("The argument FUNcluster is required. ", "Possible values are kmeans, pam, hcut, clara, ...")
else if (method %in% c("silhouette", "wss")) {
if (is.data.frame(x))
x <- as.matrix(x)
if (is.null(diss))
diss <- stats::dist(x)
v <- rep(0, k.max)
if (method == "silhouette") {
for (i in 2:k.max) {
clust <- FUNcluster(x, i, ...)
v[i] <- .get_ave_sil_width(diss, clust$cluster)
}
}
else if (method == "wss") {
for (i in 1:k.max) {
clust <- FUNcluster(x, i, ...)
v[i] <- .get_withinSS(diss, clust$cluster)
}
}
df <- data.frame(clusters = as.factor(1:k.max), y = v)
ylab <- "Total Within Sum of Square"
if (method == "silhouette")
ylab <- "Average silhouette width"
p <- ggpubr::ggline(df, x = "clusters", y = "y", group = 1,
color = linecolor, ylab = ylab, xlab = "Number of clusters k",
main = "Optimal number of clusters")
if (method == "silhouette")
p <- p + geom_vline(xintercept = which.max(v), linetype = 2,
color = linecolor)
return(p)
}
else if (method == "gap_stat") {
extra_args <- list(...)
gap_stat <- cluster::clusGap(x, FUNcluster, K.max = k.max,
B = nboot, verbose = verbose, ...)
if (!is.null(extra_args$maxSE))
maxSE <- extra_args$maxSE
else maxSE <- list(method = "firstSEmax", SE.factor = 1)
p <- fviz_gap_stat(gap_stat, linecolor = linecolor,
maxSE = maxSE)
return(p)
}
}回答 1
Stack Overflow用户
回答已采纳
发布于 2019-07-23 16:01:40
区别就在fviz_nbclust函数的开头。在第6行中,设置了随机种子:
set.seed(123)
由于kmeans算法使用的是随机启动,所以在重复运行时,结果可能会有所不同。例如,我用你的数据和两个不同的随机种子得出了稍微不同的结果。
set.seed(123)
gap_stat <- cluster::clusGap(mammals_scaled, FUN = kmeans, K.max = 24, B = 50)
fviz_gap_stat(gap_stat) + theme_minimal() + ggtitle("fviz_gap_stat: Gap Statistic")set.seed(42)
gap_stat <- cluster::clusGap(mammals_scaled, FUN = kmeans, K.max = 24, B = 50)
fviz_gap_stat(gap_stat) + theme_minimal() + ggtitle("fviz_gap_stat: Gap Statistic")我不完全确定种子123的结果为什么不一样,但我认为它与这样一个事实有关:在我的代码中,它就在clusGap函数的上方执行,而在Fviz_nbclust中,其他几个命令是在两者之间进行计算的。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/54394604
复制相关文章
相似问题

