用scatter3D错误再现pca图
用scatter3D错误再现pca图
提问于 2018-09-12 09:02:30
我对mtcars数据集进行了主成分分析,并使用下面给出的代码使用scatter3D绘制了它:
require(rgl)
require(SciViews)
require(plotrix)
require(ggplot2)
require(reshape)
require("gridExtra")
library(plot3D)
cars.pca <- pcomp(~mpg+cyl+disp+hp+drat+wt+qsec, data = mtcars)#, subset = -(8:14))
mtcars_pca = cbind(cbind(mtcars, cars.pca$scores), car = rownames(mtcars))
k <- kmeans(mtcars, 5, nstart=25, iter.max=1000)
new = cbind(mtcars_pca,cluster = k$cluster)
with(new,scatter3D(PC1,PC2,PC3, col=k$cluster, size=10, type='s', bty = "g", pch = 20, cex = 1.5,xlab="PC1",ylab="PC2",zlab="PC3",lwd = 4,pch = 20,colkey = FALSE))
with(new,text3D(PC1, PC2, PC3, labels = car,add = TRUE, colkey = FALSE, cex = 0.5,adj=-0.15))因此,获得了以下情节:
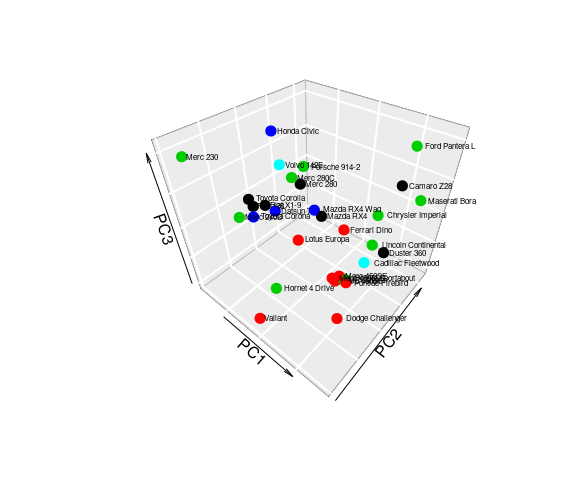
但是,当我试图为下面给出的数据集复制相同的地块时:
"NB1" "NB2" "NB3" "NF1" "NF2" "NF3" "NG1" "NG2" "NG3" "NH1" "NH2" "NH" "NL1" "NL2" "NL3" "NM1" "NM2" "NM3" "NN1" "NN2" "NN3" "NP1" "NP2" "NP3"
"Ca" 1.153 1.29013 1.221565 1.091 1.167239 1.1291195 1.156 1.28064 1.21832 1.108 1.19312 1.15056 1.134 1.013 1.0735 1.308 1.002 1.155 0.898 0.96566 0.93183 0.963 0.773 0.868
"Po" 0.842 0.91382 0.87791 0.602 0.615158 0.608579 0.544 0.55236 0.54818 0.606 0.62084 0.61342 0.689 0.616 0.6525 0.878 0.748 0.813 0.72 0.7574 0.7387 0.701 0.782 0.7415
"Gs" 1.023 1.13283 1.077915 0.853 0.898537 0.8757685 0.96 1.0474 1.0037 0.94 1.0016 0.9708 0.886 0.854 0.87 1.138 0.959 1.0485 1.008 1.09436 1.05118 0.946 0.834 0.89
"Ap" 0.549 0.55929 0.554145 0.525 0.528225 0.5266125 0.655 0.68445 0.669725 0.59 0.6026 0.5963 0.535 0.519 0.527 0.65 0.609 0.6295 0.542 0.54914 0.54557 0.664 0.554 0.609
"So" 1.74518207282913 2.00667030812325 1.87592619047619 2.15723163841808 2.37101451977401 2.26412307909605 2.32591160220995 2.67283480662983 2.49937320441989 2.73303797468354 3.04566329113924 2.88935063291139 1.14326086956522 1.84277044854881 1.49301565905701 2.0930303030303 2.49391061452514 2.29347045877772 4.03489795918367 4.6358306122449 4.33536428571429 2.64142857142857 2.20642228739003 2.4239254294093
"MDC" 0.5531148 0.5524648636 0.5527898318 0.5413232 0.541223504 0.541273352 0.562528 0.5613016 0.5619148 0.5451037 0.545458885 0.5452812925 0.5433668 0.5432226 0.5432947 0.5453266 0.5419368 0.5436317 0.5590145 0.55859421 0.558804355 0.5416098 0.5419624 0.5417861
"Elt" 72.4806763285024 72.4806763285024 72.4806763285024 74.5671641791045 74.5671641791045 74.5671641791045 51.4309309309309 51.4309309309309 51.4309309309309 83.1568265682657 83.1568265682657 83.1568265682657 73.8212341197822 90.0522388059701 77.368529286626 73.1224783861671 105.763157894737 86.6952861952862 81.1878306878307 81.1878306878307 81.1878306878307 83.2403482210447 86.8753213367609 84.5876607908528
"Hmfwt" 618.357142857143 631.321428571428 624.839285714286 654.071428571429 644.214285714286 649.142857142857 807.642857142857 799.785714285714 803.714285714286 1050.5 1078.78571428571 1064.64285714286 1000.5 793.357142857143 896.928571428571 743.357142857143 746.928571428571 745.142857142857 614.785714285714 597.785714285714 606.285714285714 821.928571428571 721.928571428571 771.928571428571
"Can" 3.10751 3.5507867 2.9510594 7.12473 8.2509341 6.7272462 1.40903 1.5635651 1.3272173 2.80357 3.1951769 2.99937345 4.17967 6.2562 5.217935 5.8482 6.25014 6.04917 3.03144 3.3605272 3.1959836 2.96378 7.63304 5.29841
"Cab" 1.5809 1.764653 1.516046 2.6063 2.964371 2.479922 1.0004 1.085468 0.955364 1.4213 1.577921 1.4996105 2.0747 2.6 2.33735 2.2589 2.3777 2.3183 2.798 3.09674 2.94737 1.886 2.6195 2.25275
"Tc" 4.18841 4.8154397 3.9671054 9.23103 10.7153051 8.7071682 1.90943 2.1490331 1.7825813 3.72487 4.2730979 3.99898395 5.75437 8.3562 7.055285 7.6071 8.12784 7.86747 5.32944 5.9572672 5.6433536 4.34978 9.75254 7.05116
"Crt" 2.40583946363636 2.72983217245455 2.29148909581818 3.79729982525253 4.35784079554545 3.59946183573737 1.36768463333333 1.515191021 1.28959301633333 1.9468614979798 2.19282795263636 2.06984472530808 2.69910104343434 3.56556422222222 3.13233263282828 3.49860908080808 3.51460955151515 3.50660931616162 3.00008494545455 3.32509598836364 3.16259046690909 2.36201111313131 3.9031210969697 3.1325661050505
"Nol" 6.5 5.5 7.5 5.5 7.5 11.5 10.5 9.5 8.5 13.5 11.5 15.5 10.5 14.5 22.5 16.5 13.5 11.5 12.5 17.5 15.5 13.5 15.5 19.5
"Sl" 1.74 1.68 1.62 7.06 9.9 9.1 6.4 5.34 4.28 6.295 6.74 5.85 5.95 8.85 7.45 8.4 7.75 8.1 6.9 6.73 6.815 4.74 3.9 11.73
"Rl" 2.8 2.6735 2.547 5.153 5.576 4.73 5.8 5.48 5.16 7.85 5.4 10.3 6.37 10.37 9.17 6.7 8.75 8.5 8.3 6.06 7.18 7.3 5.4 13.1
"Lfa" 1.7775 1.41389285714286 1.05028571428571 1.25 1.41028571428571 2.67818181818182 1.967 1.836625 1.70625 2.65254545454545 1.95309090909091 3.352 1.5857 5.34785714285714 5.99945454545455 6.7570625 4.603 4.64545454545454 3.13916666666667 1.84729411764706 2.49323039215686 3.848 3.03866666666667 2.50152631578947
"Stfwt" 0.5825 0.5835 0.5812 0.5422 0.5392 0.5412 0.5472 0.5434 0.5396 0.6444 0.5535 0.7353 0.5515 1.3233 0.8082 1.0485 0.8833 0.7783 0.5752 0.6337 0.60445 0.5877 0.685 0.6978
"Stdwt" 0.563 0.573 0.561 0.536054368932039 0.533491262135922 0.5352 0.5208 0.52015 0.5195 0.6105 0.525 0.696 0.5271 1.1622 0.7424 0.8148 0.8211 0.7366 0.5351 0.5437 0.5394 0.565 0.6598 0.556
"Lfwt" 0.5645 0.5607 0.5569 0.5372 0.5575 1.0131 0.6626 0.65245 0.6423 1.1657 0.675 1.6564 0.6694 2.3376 4.1303 2.6067 2.1981 1.598 1.0212 0.8315 0.92635 1.5245 1.2483 1.0622
"Lfdwt" 0.5398 0.53745519379845 0.535110387596899 0.5152 0.5314 0.565804852686308 0.606123723041998 0.596166677744066 0.586209632446134 0.668353675538858 0.58896499238965 0.747742358688067 0.570931034482759 0.739456110950592 1.10839018138801 1.39596683191146 0.739585612262672 0.88201750547046 0.847367447173044 0.725077490774908 0.786222468973976 0.620431922351038 0.57650553443177 0.928141916859122
"Flfwt" 0.6841 0.6732 0.6641 0.57455 0.6038 0.5453 0.5803 0.58005 0.5798 0.5619 0.5594 0.5644 0.5766 0.9186 0.6897 1.2573 0.6885 0.8109 0.7289 0.6962 0.71255 0.9158 0.598 0.6235
"Fldwt" 0.5488 1.2031 0.7833 0.5601 0.5838 0.5364 0.5692 0.56885 0.5685 0.54875 0.5487 0.5488 0.564 0.8768 0.6708 1.2031 0.6592 0.7833 0.7073 0.6771 0.6922 0.8571 0.5654 0.5944
"Rfwt" 0.5345 0.5325 0.5355 0.5522 0.5543 0.5527 0.5316 0.5315 0.5314 0.58015 0.5339 0.6264 0.532 1.1512 0.6641 1.6326 0.8613 0.8158 0.6577 0.7232 0.69045 0.8303 0.5377 0.6956
"rdw" 0.516 0.513 0.519 0.518819734345351 0.519576850094877 0.519 0.5185 0.5178 0.5171 0.5391 0.5189 0.5593 0.5147 0.8079 0.5672 0.8905 0.6732 0.6135 0.5582 0.5788 0.5685 0.5695 0.5127 0.6144使用上面给出的相同代码,我得到以下错误:
require(rgl)
require(SciViews)
require(plotrix)
require(ggplot2)
require(reshape)
require("gridExtra")
final_df <- read.table("dataset")
mydata.pca <-prcomp(~ NB1+ NB2+ NB3+ NF1+ NF2+ NF3+ NG1+ NG2+ NG3+NH1+NH2+NH+ NL1+ NL2+NL3+ NM1+ NM2+ NM3+ NN1+ NN2+ NN3+ NP1+ NP2+NP3,data=final_df)我的数据集上的pca没有生成任何pca分数,如下所示:
> mydata.pca$scores
NULL也许这就是下一条语句中错误的原因:
mydata_pca = cbind(cbind(final_df, mydata.pca$scores), car = rownames(final_df))Data.frame中的错误(.,check.names = FALSE):参数意味着不同的行数: 24,0
如何消除这一错误和上面的情节可以再现?
下面显示的是数据集的dput()输出
dput(final)
structure(list(NB1 = c(1.153, 0.842, 1.023, 0.549, 1.74518207282913,
0.5531148, 72.4806763285024, 618.357142857143, 3.10751, 1.5809,
4.18841, 2.40583946363636, 6.5, 1.74, 2.8, 1.7775, 0.5825, 0.563,
0.5645, 0.5398, 0.6841, 0.5488, 0.5345, 0.516), NB2 = c(1.29013,
0.91382, 1.13283, 0.55929, 2.00667030812325, 0.5524648636, 72.4806763285024,
631.321428571428, 3.5507867, 1.764653, 4.8154397, 2.72983217245455,
5.5, 1.68, 2.6735, 1.41389285714286, 0.5835, 0.573, 0.5607, 0.53745519379845,
0.6732, 1.2031, 0.5325, 0.513), NB3 = c(1.221565, 0.87791, 1.077915,
0.554145, 1.87592619047619, 0.5527898318, 72.4806763285024, 624.839285714286,
2.9510594, 1.516046, 3.9671054, 2.29148909581818, 7.5, 1.62,
2.547, 1.05028571428571, 0.5812, 0.561, 0.5569, 0.535110387596899,
0.6641, 0.7833, 0.5355, 0.519), NF1 = c(1.091, 0.602, 0.853,
0.525, 2.15723163841808, 0.5413232, 74.5671641791045, 654.071428571429,
7.12473, 2.6063, 9.23103, 3.79729982525253, 5.5, 7.06, 5.153,
1.25, 0.5422, 0.536054368932039, 0.5372, 0.5152, 0.57455, 0.5601,
0.5522, 0.518819734345351), NF2 = c(1.167239, 0.615158, 0.898537,
0.528225, 2.37101451977401, 0.541223504, 74.5671641791045, 644.214285714286,
8.2509341, 2.964371, 10.7153051, 4.35784079554545, 7.5, 9.9,
5.576, 1.41028571428571, 0.5392, 0.533491262135922, 0.5575, 0.5314,
0.6038, 0.5838, 0.5543, 0.519576850094877), NF3 = c(1.1291195,
0.608579, 0.8757685, 0.5266125, 2.26412307909605, 0.541273352,
74.5671641791045, 649.142857142857, 6.7272462, 2.479922, 8.7071682,
3.59946183573737, 11.5, 9.1, 4.73, 2.67818181818182, 0.5412,
0.5352, 1.0131, 0.565804852686308, 0.5453, 0.5364, 0.5527, 0.519
), NG1 = c(1.156, 0.544, 0.96, 0.655, 2.32591160220995, 0.562528,
51.4309309309309, 807.642857142857, 1.40903, 1.0004, 1.90943,
1.36768463333333, 10.5, 6.4, 5.8, 1.967, 0.5472, 0.5208, 0.6626,
0.606123723041998, 0.5803, 0.5692, 0.5316, 0.5185), NG2 = c(1.28064,
0.55236, 1.0474, 0.68445, 2.67283480662983, 0.5613016, 51.4309309309309,
799.785714285714, 1.5635651, 1.085468, 2.1490331, 1.515191021,
9.5, 5.34, 5.48, 1.836625, 0.5434, 0.52015, 0.65245, 0.596166677744066,
0.58005, 0.56885, 0.5315, 0.5178), NG3 = c(1.21832, 0.54818,
1.0037, 0.669725, 2.49937320441989, 0.5619148, 51.4309309309309,
803.714285714286, 1.3272173, 0.955364, 1.7825813, 1.28959301633333,
8.5, 4.28, 5.16, 1.70625, 0.5396, 0.5195, 0.6423, 0.586209632446134,
0.5798, 0.5685, 0.5314, 0.5171), NH1 = c(1.108, 0.606, 0.94,
0.59, 2.73303797468354, 0.5451037, 83.1568265682657, 1050.5,
2.80357, 1.4213, 3.72487, 1.9468614979798, 13.5, 6.295, 7.85,
2.65254545454545, 0.6444, 0.6105, 1.1657, 0.668353675538858,
0.5619, 0.54875, 0.58015, 0.5391), NH2 = c(1.19312, 0.62084,
1.0016, 0.6026, 3.04566329113924, 0.545458885, 83.1568265682657,
1078.78571428571, 3.1951769, 1.577921, 4.2730979, 2.19282795263636,
11.5, 6.74, 5.4, 1.95309090909091, 0.5535, 0.525, 0.675, 0.58896499238965,
0.5594, 0.5487, 0.5339, 0.5189), NH = c(1.15056, 0.61342, 0.9708,
0.5963, 2.88935063291139, 0.5452812925, 83.1568265682657, 1064.64285714286,
2.99937345, 1.4996105, 3.99898395, 2.06984472530808, 15.5, 5.85,
10.3, 3.352, 0.7353, 0.696, 1.6564, 0.747742358688067, 0.5644,
0.5488, 0.6264, 0.5593), NL1 = c(1.134, 0.689, 0.886, 0.535,
1.14326086956522, 0.5433668, 73.8212341197822, 1000.5, 4.17967,
2.0747, 5.75437, 2.69910104343434, 10.5, 5.95, 6.37, 1.5857,
0.5515, 0.5271, 0.6694, 0.570931034482759, 0.5766, 0.564, 0.532,
0.5147), NL2 = c(1.013, 0.616, 0.854, 0.519, 1.84277044854881,
0.5432226, 90.0522388059701, 793.357142857143, 6.2562, 2.6, 8.3562,
3.56556422222222, 14.5, 8.85, 10.37, 5.34785714285714, 1.3233,
1.1622, 2.3376, 0.739456110950592, 0.9186, 0.8768, 1.1512, 0.8079
), NL3 = c(1.0735, 0.6525, 0.87, 0.527, 1.49301565905701, 0.5432947,
77.368529286626, 896.928571428571, 5.217935, 2.33735, 7.055285,
3.13233263282828, 22.5, 7.45, 9.17, 5.99945454545455, 0.8082,
0.7424, 4.1303, 1.10839018138801, 0.6897, 0.6708, 0.6641, 0.5672
), NM1 = c(1.308, 0.878, 1.138, 0.65, 2.0930303030303, 0.5453266,
73.1224783861671, 743.357142857143, 5.8482, 2.2589, 7.6071, 3.49860908080808,
16.5, 8.4, 6.7, 6.7570625, 1.0485, 0.8148, 2.6067, 1.39596683191146,
1.2573, 1.2031, 1.6326, 0.8905), NM2 = c(1.002, 0.748, 0.959,
0.609, 2.49391061452514, 0.5419368, 105.763157894737, 746.928571428571,
6.25014, 2.3777, 8.12784, 3.51460955151515, 13.5, 7.75, 8.75,
4.603, 0.8833, 0.8211, 2.1981, 0.739585612262672, 0.6885, 0.6592,
0.8613, 0.6732), NM3 = c(1.155, 0.813, 1.0485, 0.6295, 2.29347045877772,
0.5436317, 86.6952861952862, 745.142857142857, 6.04917, 2.3183,
7.86747, 3.50660931616162, 11.5, 8.1, 8.5, 4.64545454545454,
0.7783, 0.7366, 1.598, 0.88201750547046, 0.8109, 0.7833, 0.8158,
0.6135), NN1 = c(0.898, 0.72, 1.008, 0.542, 4.03489795918367,
0.5590145, 81.1878306878307, 614.785714285714, 3.03144, 2.798,
5.32944, 3.00008494545455, 12.5, 6.9, 8.3, 3.13916666666667,
0.5752, 0.5351, 1.0212, 0.847367447173044, 0.7289, 0.7073, 0.6577,
0.5582), NN2 = c(0.96566, 0.7574, 1.09436, 0.54914, 4.6358306122449,
0.55859421, 81.1878306878307, 597.785714285714, 3.3605272, 3.09674,
5.9572672, 3.32509598836364, 17.5, 6.73, 6.06, 1.84729411764706,
0.6337, 0.5437, 0.8315, 0.725077490774908, 0.6962, 0.6771, 0.7232,
0.5788), NN3 = c(0.93183, 0.7387, 1.05118, 0.54557, 4.33536428571429,
0.558804355, 81.1878306878307, 606.285714285714, 3.1959836, 2.94737,
5.6433536, 3.16259046690909, 15.5, 6.815, 7.18, 2.49323039215686,
0.60445, 0.5394, 0.92635, 0.786222468973976, 0.71255, 0.6922,
0.69045, 0.5685), NP1 = c(0.963, 0.701, 0.946, 0.664, 2.64142857142857,
0.5416098, 83.2403482210447, 821.928571428571, 2.96378, 1.886,
4.34978, 2.36201111313131, 13.5, 4.74, 7.3, 3.848, 0.5877, 0.565,
1.5245, 0.620431922351038, 0.9158, 0.8571, 0.8303, 0.5695), NP2 = c(0.773,
0.782, 0.834, 0.554, 2.20642228739003, 0.5419624, 86.8753213367609,
721.928571428571, 7.63304, 2.6195, 9.75254, 3.9031210969697,
15.5, 3.9, 5.4, 3.03866666666667, 0.685, 0.6598, 1.2483, 0.57650553443177,
0.598, 0.5654, 0.5377, 0.5127), NP3 = c(0.868, 0.7415, 0.89,
0.609, 2.4239254294093, 0.5417861, 84.5876607908528, 771.928571428571,
5.29841, 2.25275, 7.05116, 3.1325661050505, 19.5, 11.73, 13.1,
2.50152631578947, 0.6978, 0.556, 1.0622, 0.928141916859122, 0.6235,
0.5944, 0.6956, 0.6144)), .Names = c("NB1", "NB2", "NB3", "NF1",
"NF2", "NF3", "NG1", "NG2", "NG3", "NH1", "NH2", "NH", "NL1",
"NL2", "NL3", "NM1", "NM2", "NM3", "NN1", "NN2", "NN3", "NP1",
"NP2", "NP3"), class = "data.frame", row.names = c("Ca", "Po",
"Gs", "Ap", "So", "MDC", "Elt", "Hmfwt", "Can", "Cab", "Tc",
"Crt", "Nol", "Sl", "Rl", "Lfa", "Stfwt", "Stdwt", "Lfwt", "Lfdwt",
"Flfwt", "Fldwt", "Rfwt", "rdw"))回答 1
Stack Overflow用户
回答已采纳
发布于 2018-09-12 13:07:49
我们可以试试这个。首先,分数不在mydata.pca$scores中,因为正如您可以看到的那样,键入?prcomp没有它,但是您可以拥有$x,并使用它。
所以你的模型:
require(rgl)
require(car)
mydata.pca <-prcomp(~ NB1+ NB2+ NB3+ NF1+ NF2+ NF3+ NG1+ NG2+ NG3+NH1+NH2+NH+ NL1+ NL2+NL3+ NM1+ NM2+ NM3+ NN1+ NN2+ NN3+ NP1+ NP2+NP3,data=final)
mydata_pc <- mydata.pca$x[,1:3]前三台个人电脑。请注意,如果您发送以下内容,您似乎有一些不好的东西:
summary(mydata.pca)因为你只需要一台PC来解释所有的可变性,而另一台电脑则让另一台电脑很难去关心(但这是我的观点,也许你不应该同意,但它指出你应该去看它,也要关注cor(final))。
现在您有了k表示输出:
k <- kmeans(final, 5, nstart=25, iter.max=1000)
# bind together the outputs
new <- data.frame(mydata_pc,cluster = k$cluster)再一次,它似乎是一个输出。
summary(k)最后,您可以绘制所有内容:
scatter3d(PC1~PC2+PC3, data=new, group=as.factor(new$cluster), size=10, type='s', bty = "g", pch = 20, cex = 1.5,xlab="PC1",ylab="PC2",zlab="PC3",lwd = 4,pch = 20,colkey = FALSE)
正如这里所说,您可以找到一种更好的方法来绘制它们。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/52291358
复制相关文章
相似问题

