条形图分类数据综述
条形图分类数据综述
提问于 2018-04-27 11:12:00
I ^有一个如下所示的数据集:
typestudy dloop cytb coi other microsat SNP
methods no no no no yes no
methods yes no no no no yes
methods no no no no yes no
methods no no no no yes no
wildcrime no no no yes no no
taxonomy no no no no yes no
methods yes no no no no no
methods no no no no yes no
taxonomy no no no no yes no
wildcrime yes no no no no no
methods yes no no no no no
taxonomy no no no no yes yes
taxonomy no no no no yes no但它有10列是/否,与进一步的遗传元素相对应,有200多行。
在Excel中,图形摘要选项给了我一个很好的堆叠条形图,但我需要能够在R中重新创建它,以满足我的报告的大学标准
> summary(dframe1$type.of.study)
methods development other
49 5
population genetic structure taxonomy
91 86
wildlife crime
6
> barplot(as.matrix(dframe1))
There were 11 warnings (use warnings() to see them)
> warnings()
Warning messages:
1: In apply(height, 2L, cumsum) : NAs introduced by coercion
2: In apply(height, 2L, cumsum) : NAs introduced by coercion
3: In apply(height, 2L, cumsum) : NAs introduced by coercion
4: In apply(height, 2L, cumsum) : NAs introduced by coercion
5: In apply(height, 2L, cumsum) : NAs introduced by coercion
6: In apply(height, 2L, cumsum) : NAs introduced by coercion
7: In apply(height, 2L, cumsum) : NAs introduced by coercion
8: In apply(height, 2L, cumsum) : NAs introduced by coercion
9: In apply(height, 2L, cumsum) : NAs introduced by coercion
10: In apply(height, 2L, cumsum) : NAs introduced by coercion
11: In apply(height, 2L, cumsum) : NAs introduced by coercion这给了我这个

我也成功地制作了这个,但是找不到我所用的脚本
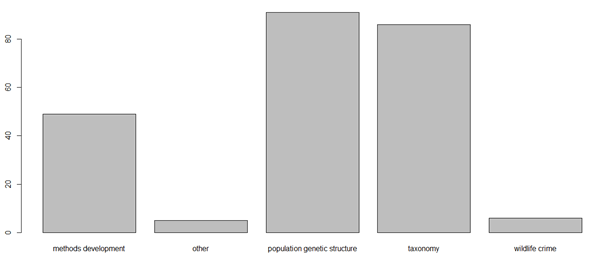
我的目标与此类似:
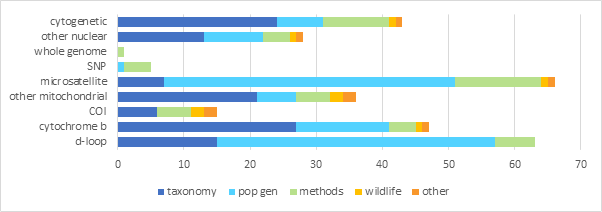
这是相当可悲的,但我花了将近一个星期的故障排除基于在线资源和其他问题在这里达到这一点。我不知道如何计算研究的类型,所以他们被计算出来,使不同的遗传标记对应的条形图的高度。我知道这对于堆栈溢出的标准来说太模糊了,但是我很绝望,所以我不想再提这个了,以防有人有任何建议
^ (我会尽量简洁,但你会看到我在R方面的流利程度太差了,我不会寻求帮助,但我花了好几天时间处理这些数据,我被吓呆了,我永远找不到解决办法,也没有人在我的研究职位上寻求帮助)
回答 3
Stack Overflow用户
发布于 2018-04-27 11:24:20
我们可以通过循环获得每一列的table,然后执行barplot。
barplot(sapply(df1[-1], function(x) table(factor(x,
levels = c("yes", "no")))), col = c("red", "blue"))
legend("topright", legend = c("yes", "no"), fill = c("red", "blue"))Stack Overflow用户
发布于 2018-04-27 11:29:08
以防万一你想要的东西看起来像这个例子(有点):
df <- read.table(text = "typestudy dloop cytb coi other microsat SNP
methods no no no no yes no
methods yes no no no no yes
methods no no no no yes no
methods no no no no yes no
wildcrime no no no yes no no
taxonomy no no no no yes no
methods yes no no no no no
methods no no no no yes no
taxonomy no no no no yes no
wildcrime yes no no no no no
methods yes no no no no no
taxonomy no no no no yes yes
taxonomy no no no no yes no",
header = T, stringsAsFactors = F)
library(tidyr)
library(ggplot2)
library(dplyr)
df %>% gather(key = key, value = value, -typestudy) %>%
filter(value == "yes") %>%
ggplot(aes(x = key, fill = typestudy)) +
geom_bar() +
coord_flip() +
theme_minimal() +
theme(legend.position = "bottom",
panel.grid.minor = element_blank(),
panel.grid.major.y = element_blank()) +
xlab(NULL) +
ylab(NULL)Stack Overflow用户
发布于 2018-04-27 12:25:33
我不知道您是否只是在yeses之后,但是这里有一种可能,可以让您使用no,以防您想要按响应类型(是/否)的巴图。
df %>%
gather(var, value, -typestudy) %>%
group_by(typestudy, var, value) %>%
count() %>%
filter(value == "yes") %>%
ggplot(aes(var, n, group = typestudy, fill = typestudy)) +
geom_bar(stat = "identity") +
scale_fill_brewer(palette = "Dark2", direction = -1) +
coord_flip() +
theme(
axis.title.x=element_blank(),
axis.title.y=element_blank(),
legend.position = "bottom",
panel.grid.minor = element_blank(),
panel.grid.major.y = element_blank(),
legend.title=element_blank())
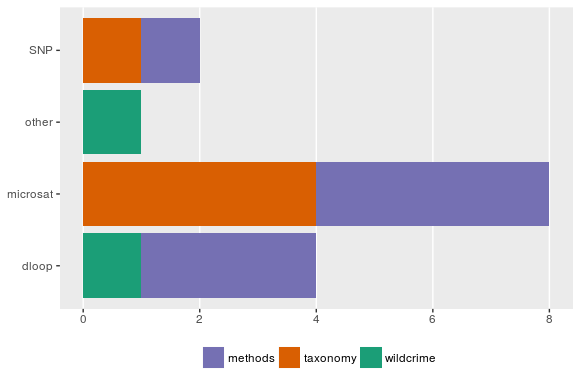
数据
df <- structure(list(typestudy = c("methods", "methods", "methods",
"methods", "wildcrime", "taxonomy", "methods", "methods", "taxonomy",
"wildcrime", "methods", "taxonomy", "taxonomy"), dloop = c("no",
"yes", "no", "no", "no", "no", "yes", "no", "no", "yes", "yes",
"no", "no"), cytb = c("no", "no", "no", "no", "no", "no", "no",
"no", "no", "no", "no", "no", "no"), coi = c("no", "no", "no",
"no", "no", "no", "no", "no", "no", "no", "no", "no", "no"),
other = c("no", "no", "no", "no", "yes", "no", "no", "no",
"no", "no", "no", "no", "no"), microsat = c("yes", "no",
"yes", "yes", "no", "yes", "no", "yes", "yes", "no", "no",
"yes", "yes"), SNP = c("no", "yes", "no", "no", "no", "no",
"no", "no", "no", "no", "no", "yes", "no")), .Names = c("typestudy",
"dloop", "cytb", "coi", "other", "microsat", "SNP"), class = "data.frame", row.names = c(NA,
-13L))页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/50061165
复制相关文章
相似问题

