表示on图上“`lsmeans()”中的多对比较p值
表示on图上“`lsmeans()”中的多对比较p值
提问于 2018-04-22 02:15:17
我已经通过使用lsmeans()对lme模型对象进行临时对对比较(更正)生成了许多p-值。我有一个来自ggplot2的图解,它本质上是多个躲避条形图。我现在要显示那些p-值,上面有星星和线,以显示比较。有什么简单的方法吗?
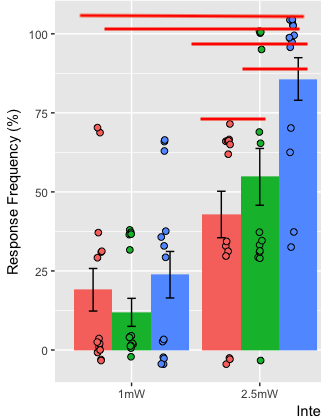
甚至更好。有没有一种更优雅的方式来展示这些两两比较呢?
回答 1
Stack Overflow用户
发布于 2022-04-12 22:42:17
派对有点晚了,但这里有个可能对你有用的答案。甚至还能很好地处理小情节。我将在序言中说,rstatix包有一些选项,可以使数据帧很容易地传递到ggprism::add_pvalue()函数。在这里,我使用来自rstatix::wilcoox_test()的坐标映射输出,并将它与我感兴趣的测试的输出结合起来。我的方法可能看起来非常复杂,但它是有效的!
我的数据片段:
- 参与者样本的样本->样本ID是从(纵向研究中的依赖抽样)中收集的。
- 时间点->从V0到V5的字符向量,识别每个样本的收集时间(被绑定到特定的时间点)
- 从很低到很高的描述抗体-抗原相互作用强度的字符载体(基于Kd值)。
- 亲和力->:每个“范围”的浓度的量
首先,从在rstatix中运行未配对Wilcox测试开始,并使用add_xy_position,但我们只提取组比较(group1和group2)、因式分解变量(Range)以及x和y定位。
library(tidyverse)
library(here)
# Perform Wilcox test just to get y.position and xmin/max for lsmeans
comp <- avw_abs %>%
rstatix::group_by(Range) %>%
rstatix::wilcox_test(Avidity ~ Timepoint, paired = F) %>%
rstatix::add_y_position(fun = "max",step.increase = 0.001) %>%
rstatix::add_x_position() %>%
select(c(group1, group2, Range, y.position, xmin,xmax)) 接下来,我们将使用lme4::lmer()函数和lsmeans::lsmeans()执行实际建模和比较。根据对比度(组比较)输出的不同,您需要使用separate()参数。分离和联合函数将分组变量分解为ggprism::add_pvalue()函数所需的单独列。.y。是rstatix::wilcox_test中的另一个输出变量,但我不确定它是否相关。
lm_abs <- lme4::lmer(log10(Avidity) ~ Timepoint+Range + (1|Sample), avw_abs) # make model
plot(resid(lm_abs)) # Checking model
qqnorm(resid(lm_abs))
comp_ls <- lsmeans::lsmeans(lm_abs, # Extract model comparisons
pairwise ~ c(Timepoint, Range), data = avw_abs,
adjust="BH")$contrast %>%
broom::tidy() %>%
separate(contrast, sep = " - ",into = c("group1","group2")) %>%
separate(group1, sep = " ", into = c("group1", "Range", "Range2")) %>%
unite("Range", c("Range", "Range2"), sep = " ",na.rm = T) %>%
separate(group2, sep = " ", into = c("group2", "Range3", "Range4")) %>%
unite("Range1", c("Range3", "Range4"), sep = " ",na.rm = T) %>%
mutate(.y. = "Avidity") %>%
filter(Range == Range1) %>% # Only interested in comparing the same range across timepoints
mutate(xmin = case_when(group1 == "V0" ~ 1,
group1 == "V1" ~ 2,
group1 == "V2" ~ 3,
group1 == "V3" ~ 4,
group1 == "V4" ~ 5,
group1 == "V5" ~ 6),
xmax = case_when(group2 == "V0" ~ 1,
group2 == "V1" ~ 2,
group2 == "V2" ~ 3,
group2 == "V3" ~ 4,
group2 == "V4" ~ 5,
group2 == "V5" ~ 6)) %>%
select(-Range1) %>%
group_by(Range) %>%
mutate(p.adj.signif = ifelse(adj.p.value > 0.05, "ns", ifelse(adj.p.value < 0.0001, "****", ifelse(adj.p.value < 0.001, "***", ifelse(adj.p.value < 0.01, "**", ifelse(adj.p.value < 0.05, "*", "ns")))))) %>% # Create mapping for p value symbols
mutate(Range = fct_relevel(Range, levels = c("Very low","Low", "Medium", "High", "Very high")))
comp_ls_bs <- full_join(comp, comp_ls, by = c("group1","group2","Range")) # Combine with rstatix output情节:
avw_abs_ %>%
ggplot(aes(as.factor(Timepoint), Avidity, fill = Range)) +
scale_fill_brewer(palette = "Blues") +
geom_col(data = ~avw_abs_ %>%
group_by(Range, Timepoint) %>%
summarize(Avidity = 10^mean(log10(Avidity))) %>%
ungroup(),
aes(group = Range),
colour = "black", position = position_dodge2()) +
ggnewscale::new_scale_fill() +
scale_fill_manual(values = RColorBrewer::brewer.pal(6, "Greys")) +
geom_jitter(pch = 21, size = 3, cex = 100, alpha = 0.7,
position = position_jitterdodge(0.9, jitter.width = 0.25, jitter.height = 0.00001), aes(fill = Range)) +
geom_errorbar(data = ~avw_abs_ %>%
group_by(Range, Timepoint) %>%
mutate(n = n(),
lowercv95 = 10^(mean(log10(Avidity))-qnorm(0.975)*sd(log10(Avidity))/sqrt(n)),
uppercv95 = 10^(mean(log10(Avidity))+qnorm(0.975)*sd(log10(Avidity))/sqrt(n))) %>%
mutate(lowercv95 = mean(lowercv95),uppercv95 = mean(uppercv95)),
aes(ymin = lowercv95, ymax = uppercv95),
position = position_dodge(0.9), size = 0.5, width = 0.5) +
scale_y_log10(
breaks = scales::trans_breaks("log10", function(x) 10^x,n = 9),
labels = scales::trans_format("log10", scales::math_format(10^.x))) +
facet_wrap(vars(Range), nrow = 1) +
coord_cartesian(ylim = c(10^2, 10^11), clip = "on") +
geom_hline(yintercept = 10^log10(317), linetype = "dashed", color = "darkred") +
ggprism::add_pvalue(comp_ls_bs, tip.length = 0, label.size = 2, step.group.by = "Range", step.increase = 0.035) + #### Here is where you add the p values
labs(y = "Avidity (U/mL)") +
guides(fill = guide_legend(title = c("Timepoint", "Avidity"),nrow = 1)) +
theme_classic() +
theme(axis.text= element_text(size = 8, color = "black"),
axis.title.x = element_blank(),
axis.title.y = element_text(size = 10, face = "bold"),
legend.text = element_text(size = 8),
legend.position = "none",
legend.key.size = unit(0.35, 'cm'),
legend.title = element_text(face = "bold"),
axis.ticks = element_line(colour = "black"),
strip.background = element_blank(),
plot.margin = unit(c(1.5,0.5,0.5,0.5), "cm"))
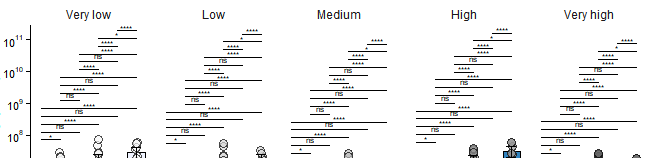
如果你有问题,我可以做一个更一般的例子。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/49961916
复制相关文章
相似问题

