在Neo4j中建立代谢途径
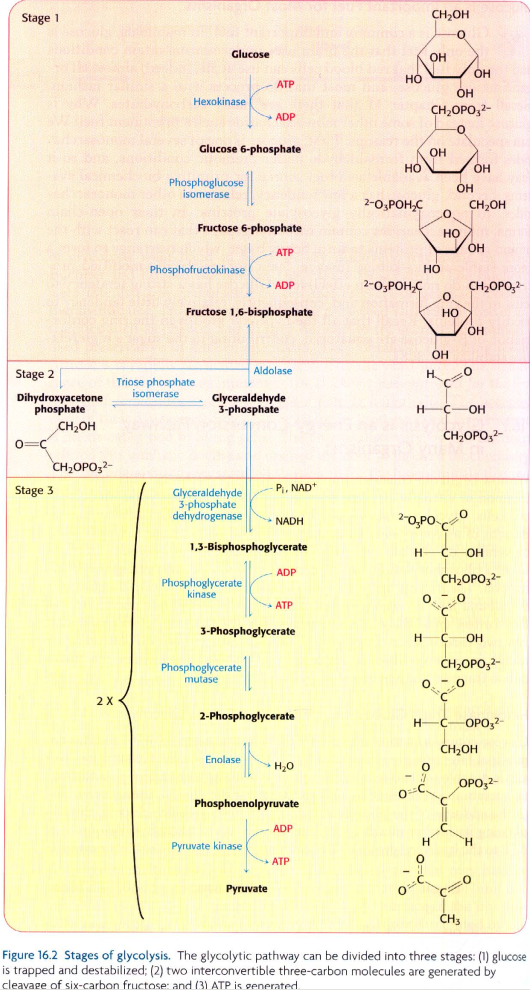
我试图在Neo4j中创建在这个问题底部的图像中显示的糖酵解途径,使用以下数据:
glycolysis_bioentities.csv
name
α-D-glucose
glucose 6-phosphate
fructose 6-phosphate
"fructose 1,6-bisphosphate"
dihydroxyacetone phosphate
D-glyceraldehyde 3-phosphate
"1,3-bisphosphoglycerate"
3-phosphoglycerate
2-phosphoglycerate
phosphoenolpyruvate
pyruvate
hexokinase
glucose-6-phosphatase
phosphoglucose isomerase
phosphofructokinase
"fructose-bisphosphate aldolase, class I"
triosephosphate isomerase (TIM)
glyceraldehyde-3-phosphate dehydrogenase
phosphoglycerate kinase
phosphoglycerate mutase
enolase
pyruvate kinaseglycolysis_relations.csv
source,relation,target
α-D-glucose,substrate_of,hexokinase
hexokinase,yields,glucose 6-phosphate
glucose 6-phosphate,substrate_of,glucose-6-phosphatase
glucose-6-phosphatase,yields,α-D-glucose
glucose 6-phosphate,substrate_of,phosphoglucose isomerase
phosphoglucose isomerase,yields,fructose 6-phosphate
fructose 6-phosphate,substrate_of,phosphofructokinase
phosphofructokinase,yields,"fructose 1,6-bisphosphate"
"fructose 1,6-bisphosphate",substrate_of,"fructose-bisphosphate aldolase, class I"
"fructose-bisphosphate aldolase, class I",yields,D-glyceraldehyde 3-phosphate
D-glyceraldehyde 3-phosphate,substrate_of,glyceraldehyde-3-phosphate dehydrogenase
D-glyceraldehyde 3-phosphate,substrate_of,triosephosphate isomerase (TIM)
triosephosphate isomerase (TIM),yields,dihydroxyacetone phosphate
glyceraldehyde-3-phosphate dehydrogenase,yields,"1,3-bisphosphoglycerate"
"1,3-bisphosphoglycerate",substrate_of,phosphoglycerate kinase
phosphoglycerate kinase,yields,3-phosphoglycerate
3-phosphoglycerate,substrate_of,phosphoglycerate mutase
phosphoglycerate mutase,yields,2-phosphoglycerate
2-phosphoglycerate,substrate_of,enolase
enolase,yields,phosphoenolpyruvate
phosphoenolpyruvate,substrate_of,pyruvate kinase
pyruvate kinase,yields,pyruvate到目前为止这就是我所拥有的,
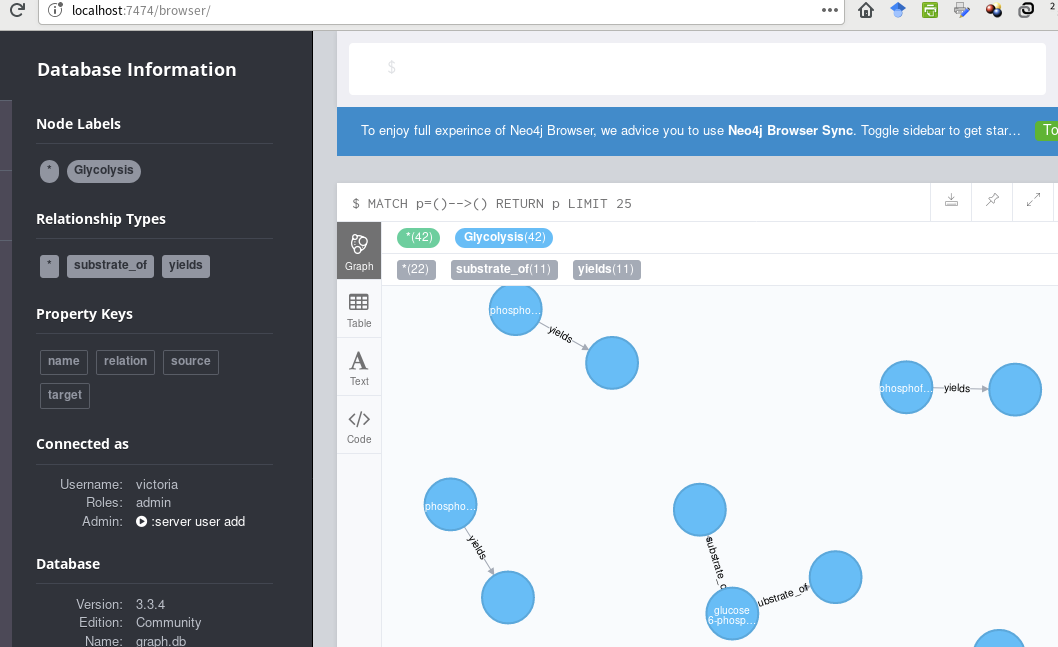
..。使用此密码码(传递给Cycli或cypher-shell):
LOAD CSV WITH HEADERS FROM "file:/glycolysis_relations.csv" AS row
MERGE (s:Glycolysis {source: row.source})
MERGE (r:Glycolysis {relation: row.relation})
MERGE (t:Glycolysis {target: row.target})
FOREACH (x in case row.relation when "substrate_of" then [1] else [] end |
MERGE (s)-[r:substrate_of]->(t)
)
FOREACH (x in case row.relation when "yields" then [1] else [] end |
MERGE (s)-[r:yields]->(t)
);我想要创建完全连接的路径,在所有节点上都有标题。有什么建议吗?
回答 3
Stack Overflow用户
发布于 2018-04-06 00:34:36
已更新
有多个问题和可能的改进:
- 应该删除第二个
MERGE,因为它创建了孤立的节点。关系类型不应该调优到Glycolysis节点,并且这些节点永远不会连接到任何其他节点。 - 第一条和第三条
MERGE子句必须对源节点和目标节点使用相同的属性名(例如,name),否则相同的化学物质可能会有两个节点(具有不同的属性键)。这就是为什么您最终得到的节点没有所有预期的连接。 - APOC过程apoc.cypher.doIt可以在某种程度上简化带有动态名称的关系的
MERGE。 - 这个用例不需要
glycolysis_bioentities.csv。
通过上述更改,您将得到这样的结果,这将生成一个与输入数据匹配的连通图:
LOAD CSV WITH HEADERS FROM "file:/glycolysis_relations.csv" AS row
MERGE (s:Glycolysis {name: row.source})
MERGE (t:Glycolysis {name: row.target})
WITH s, t, row
CALL apoc.cypher.doIt(
'MERGE (s)-[r:' + row.relation + ']->(t)',
{s:s, t:t}) YIELD value
RETURN 1;Stack Overflow用户
发布于 2018-04-06 02:44:58
@赛博赛姆的回答很好,提供了最优雅的解决方案(再次:谢谢!) --请投上接受的答案。
由于这个问题/答案/主题可能会引起其他人的兴趣,我想提一下我的代码(基于这个线程、如何在CSV中指定关系类型?和根据@赛博SO提供的提示进行修改)现在起作用了,并显示了结果:
解决方案1(我最初的文章,更新):
LOAD CSV WITH HEADERS FROM "file:/glycolysis_relations.csv" AS row
MERGE (s:Glycolysis {name:row.source})
MERGE (t:Glycolysis {name:row.target})
FOREACH (x in case row.relation when "substrate_of" then [1] else [] end |
MERGE (s)-[r:substrate_of]->(t)
)
FOREACH (x in case row.relation when "yields" then [1] else [] end |
MERGE (s)-[r:yields]->(t)
);解决方案2(赛博Solution,更新):
LOAD CSV WITH HEADERS FROM "file:/glycolysis_relations.csv" AS row
MERGE (s:Metabolism:Glycolysis {name: row.source})
MERGE (t:Metabolism:Glycolysis {name: row.target})
WITH s, t, row
// "Bug" -- additional duplicate relations with each iteration of this statement/script:
// CALL apoc.create.relationship(s, row.relation, {}, t) YIELD rel
// Solution:
// https://github.com/neo4j-contrib/neo4j-apoc-procedures/issues/271
// https://stackoverflow.com/questions/47808421/neo4j-load-csv-to-create-dynamic-relationship-types
CALL apoc.merge.relationship(s, row.relation, {}, {}, t) YIELD rel
RETURN COUNT(*);这两种解决方案都生成相同的图,如下所示。-D
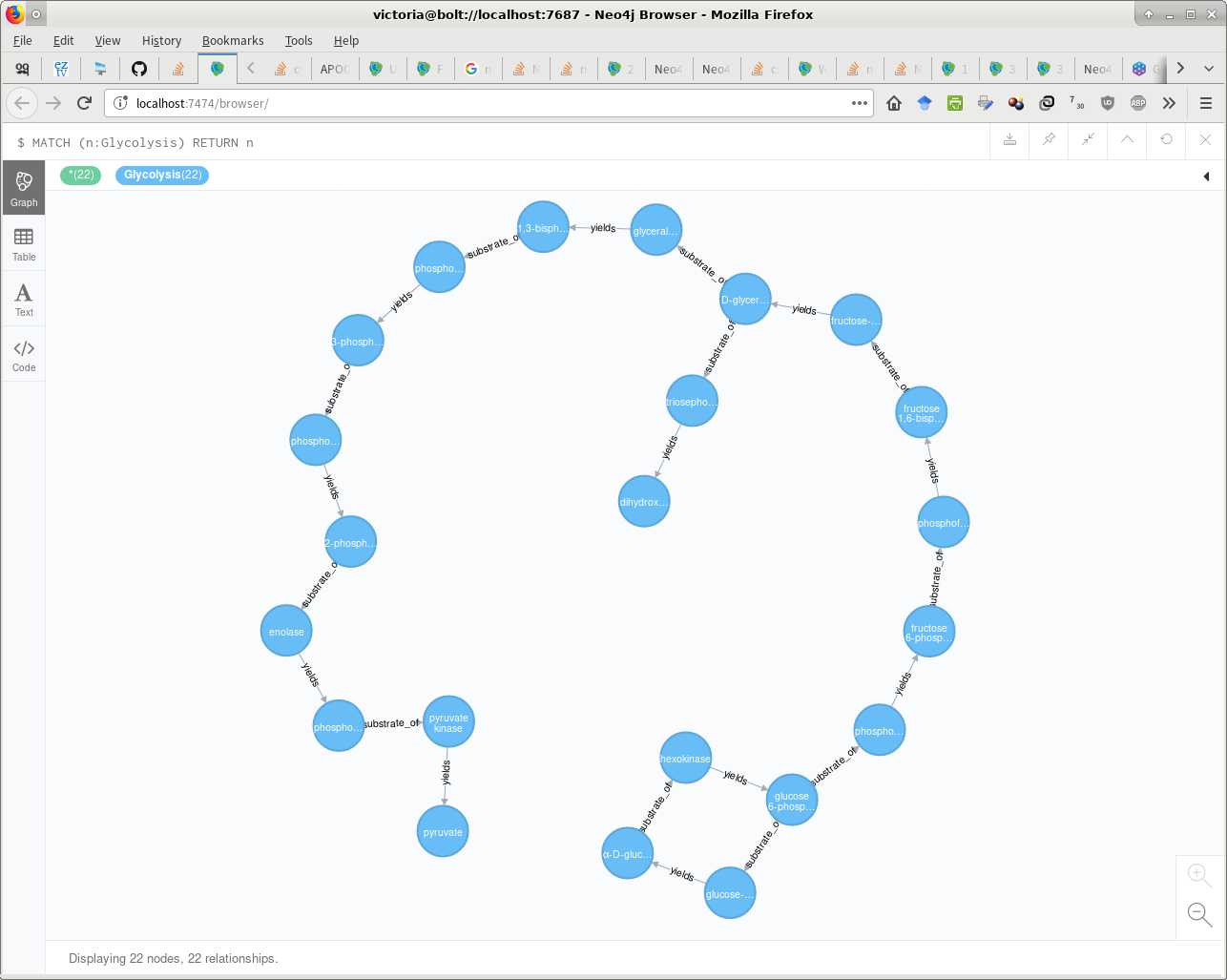
Stack Overflow用户
发布于 2018-04-12 20:04:58
如果允许的话,我还想发布一个后续的答案--我的理由是,目前很少有关于在Neo4j中重建代谢途径的信息,下面将在StackOverflow标题/主题下提供一个完整的摘要,“在Neo4j中创建一个代谢途径”。
像我的糖酵解途径,上面,我在Neo4j重建了TCA (柠檬酸循环?克雷伯循环)途径:
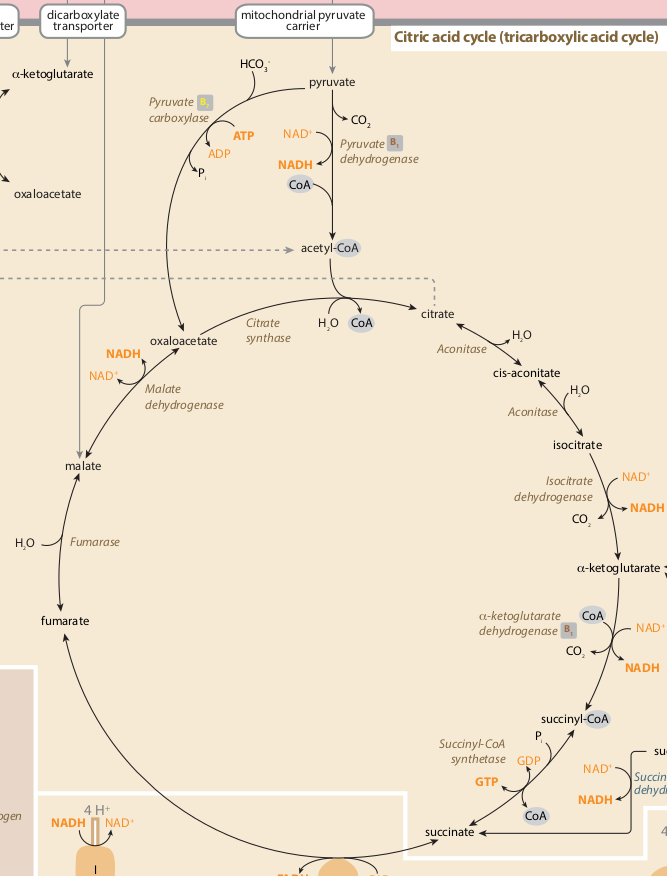
循环像源:[https://metabolicpathways.stanford.edu/] ]
在我的TCA路径图创建过程中出现的一个问题是,其中一个节点(酶,“乌头酶”)被使用了两次,所以在图形创建过程中,MERGE将公共节点aconitase合并为一个单一的实体,从而形成了这个布局,
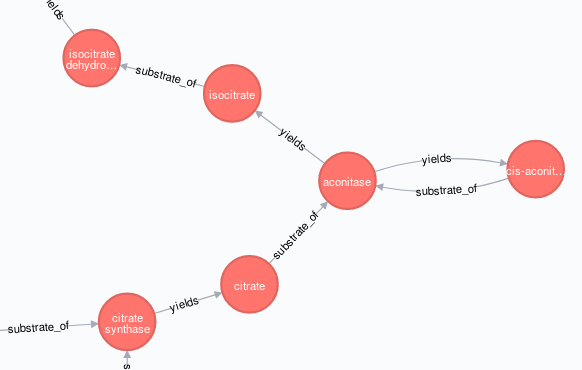
..。不是这个,如你所愿,
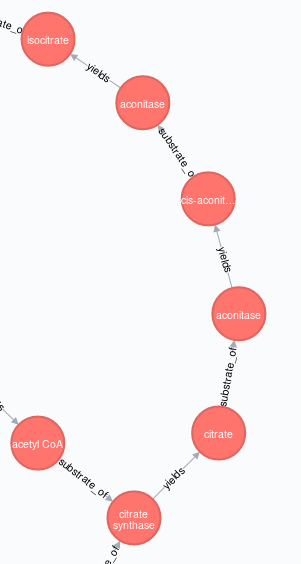
我对这个问题的解决方案是使用节点属性创建"TCA图“,临时区分-标记受影响的源节点和目标节点(稍后在正确创建图形之后删除这些标记)。
我还添加了一个:Metabolism标签,这样我就可以根据需要选择单独的路径(:Glycolysis \ :TCA)或完整的代谢网络(:Metabolism)。
最后,我需要通过它们的公共节点pyruvate连接这两条路径( :TCA),这是我能够通过APOC过程完成的(这里,附加到我的glycolysis.cql (Cypher)脚本的末尾)。
下面是我的CSV数据文件、*.cql Cypher脚本、脚本执行和结果图。
glycolysis.csv:
source,relation,target
α-D-glucose,substrate_of,hexokinase
hexokinase,yields,glucose 6-phosphate
glucose 6-phosphate,substrate_of,glucose-6-phosphatase
glucose-6-phosphatase,yields,α-D-glucose
glucose 6-phosphate,substrate_of,phosphoglucose isomerase
phosphoglucose isomerase,yields,fructose 6-phosphate
fructose 6-phosphate,substrate_of,phosphofructokinase
phosphofructokinase,yields,"fructose 1,6-bisphosphate"
"fructose 1,6-bisphosphate",substrate_of,"fructose-bisphosphate aldolase, class I"
"fructose-bisphosphate aldolase, class I",yields,D-glyceraldehyde 3-phosphate
D-glyceraldehyde 3-phosphate,substrate_of,glyceraldehyde-3-phosphate dehydrogenase
D-glyceraldehyde 3-phosphate,substrate_of,triosephosphate isomerase (TIM)
triosephosphate isomerase (TIM),yields,dihydroxyacetone phosphate
glyceraldehyde-3-phosphate dehydrogenase,yields,"1,3-bisphosphoglycerate"
"1,3-bisphosphoglycerate",substrate_of,phosphoglycerate kinase
phosphoglycerate kinase,yields,3-phosphoglycerate
3-phosphoglycerate,substrate_of,phosphoglycerate mutase
phosphoglycerate mutase,yields,2-phosphoglycerate
2-phosphoglycerate,substrate_of,enolase
enolase,yields,phosphoenolpyruvate
phosphoenolpyruvate,substrate_of,pyruvate kinase
pyruvate kinase,yields,pyruvatetca.csv:
source,relation,target,tag1,tag2
pyruvate,substrate_of,pyruvate dehydrogenase,,
pyruvate dehydrogenase,yields,acetyl CoA,,
acetyl CoA,substrate_of,citrate synthase,,
oxaloacetate,substrate_of,citrate synthase,,
citrate synthase,yields,citrate,,
citrate,substrate_of,aconitase,,1
aconitase,yields,cis-aconitate,1,
cis-aconitate,substrate_of,aconitase,,2
aconitase,yields,isocitrate,2,
isocitrate,substrate_of,isocitrate dehydrogenase,,
isocitrate dehydrogenase,yields,α-ketoglutarate,,
α-ketoglutarate,substrate_of,α-ketoglutarate dehydrogenase,,
α-ketoglutarate dehydrogenase,yields,succinyl-CoA,,
succinyl-CoA,substrate_of,succinyl-CoA synthetase,,
succinyl-CoA synthetase,yields,succinate,,
succinate,substrate_of,succinate dehydrogenase,,
succinate dehydrogenase,yields,fumarate,,
fumarate,substrate_of,fumarase,,
fumarase,yields,S-malate,,
S-malate,substrate_of,malate dehydrogenase,,
malate dehydrogenase,yields,oxaloacetate,,在通过"tag1“脚本创建源节点和目标节点时,"tsv.csv”中的"tca.cql“和”标记“2用于唯一地用于这些源节点和目标节点:
tca.cql:
// CREATE INDICES:
CREATE INDEX ON :Metabolism(name);
CREATE INDEX ON :TCA(name);
// CREATE GRAPH:
// USING PERIODIC COMMIT 5000
LOAD CSV WITH HEADERS FROM "file:/mnt/Vancouver/Programming/data/metabolism/tca.csv" AS row
MERGE (s:Metabolism:TCA {name: row.source, tag:COALESCE(row.tag1, '')})
MERGE (t:Metabolism:TCA {name: row.target, tag:COALESCE(row.tag2, '')})
WITH s, t, row
CALL apoc.merge.relationship(s, row.relation, {}, {}, t) YIELD rel
REMOVE s.tag, t.tag
RETURN COUNT(*);glycolysis.cql:
// CREATE INDICES:
CREATE INDEX ON :Metabolism(name);
CREATE INDEX ON :Glycolysis(name);
// CREATE GRAPH:
//USING PERIODIC COMMIT 5000
LOAD CSV WITH HEADERS FROM "file:/mnt/Vancouver/Programming/data/metabolism/glycolysis.csv" AS row
MERGE (s:Metabolism:Glycolysis {name: row.source})
MERGE (t:Metabolism:Glycolysis {name: row.target})
WITH s, t, row
CALL apoc.merge.relationship(s, row.relation, {}, {}, t) YIELD rel
RETURN COUNT(*);
// MERGE COMMON NODE (GLYCOLYSIS: PYRUVATE; TCA: PYRUVATE):
// As presented, run "tca.cql" first, then "glycolysis.cql"
MATCH (g:Glycolysis), (t:TCA) WHERE g.name = t.name
CALL apoc.refactor.mergeNodes([g,t]) YIELD node
RETURN node;脚本执行:
$ cat tca.cql | cypher-shell -u *** -p ***
COUNT(*)
21
$ cat glycolysis.cql | cypher-shell -u *** -p ***
COUNT(*)
22
node
(:Metabolism:TCA:Glycolysis {name: "pyruvate"})
$ Neo4j图(**:Metabolism** 视图):
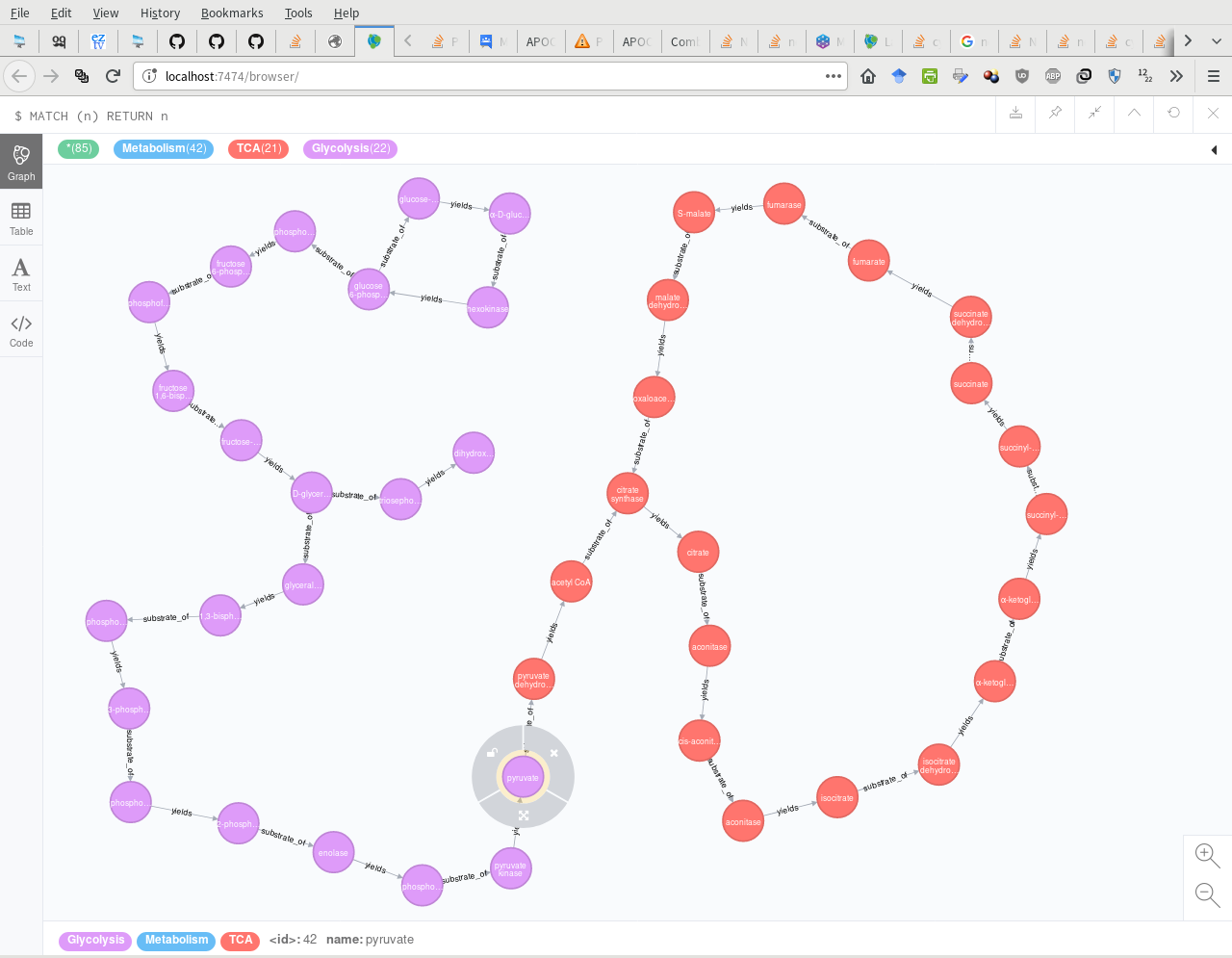
https://stackoverflow.com/questions/49682338
复制相似问题

