如何创建带有所有点的方框图,每个组的颜色可以手动分配。
如何创建带有所有点的方框图,每个组的颜色可以手动分配。
提问于 2017-09-25 17:15:02
我有一个数据框架:
> dput(df2)
structure(list(Genotype = c("miR-15/16 FL", "miR-15/16 FL", "miR-15/16 FL",
"miR-15/16 FL", "miR-15/16 FL", "miR-15/16 cKO", "miR-15/16 cKO",
"miR-15/16 cKO", "miR-15/16 cKO", "miR-15/16 cKO"),
`Cells/SC/Live/CD8—,, CD4+/Foxp3-,Median,<BV421-A>,CD127` = c(1191L, 1325L, 1089L, 1154L, 1147L, 1735L, 1441L, 1455L, 1560L, 1623L)),
.Names = c("Genotype", "Cells/SC/Live/CD8—,, CD4+/Foxp3-,Median,<BV421-A>,CD127"),
row.names = c(NA, -10L), class = c("tbl_df", "tbl", "data.frame"))
MFI=c(1191,1325,1089,1154,1147,1735,1441,1455,1560,1623))我想用每一组(miR-15/16 FL和miR-15/16 cKO)绘制一个盒形图,并将所有的
'miR-15/16 FL' 点是封闭的黑圈,所有的
'miR-15/16 cKO' 点开红色圆圈。我想要能够手动调整颜色和形状/大小的点数为每一组。
到目前为止,我已经尝试过:
library(ggplot2)
ggplot(data=df2, aes(x = df2$Genotype, y = df2[2])) +
geom_boxplot(outlier.shape = NA) +
geom_jitter(position = position_jitter(width = .2), shape=1, size=5) +
ylim(0,max(df2[2])+10)但是我还没有弄清楚如何独立地调整颜色/形状
'miR-15/16 FL'和
'miR-15/16 cKO'谢谢你在这方面的帮助!
回答 2
Stack Overflow用户
回答已采纳
发布于 2017-09-25 17:32:54
@巴尔特打败了我.唯一的区别是,我设置了全球图形调用之外的颜色和形状参数,用于未来的进度访问。
library(ggplot2)
df2 <- data.frame(Genotype = c('WT','WT','WT','WT','WT',
'cKO','cKO','cKO','cKO','cKO'),
MFI=c(1191,1325,1089,1154,1147,1735,1441,1455,1560,1623))
color.groups <- c(WT="black", cKO="red")
shape.groups <- c(WT=20, cKO=21)
ggplot(data=df2, aes(x = df2$Genotype, y = df2$MFI)) +
geom_boxplot(outlier.shape = NA) +
geom_point(position = position_jitter(width = .2), size=5,
aes(color=Genotype, shape = Genotype)) +
ylim(0,max(df2$MFI)+10) +
scale_color_manual(values=color.groups) +
scale_shape_manual(values=shape.groups)更新:
library(ggplot2)
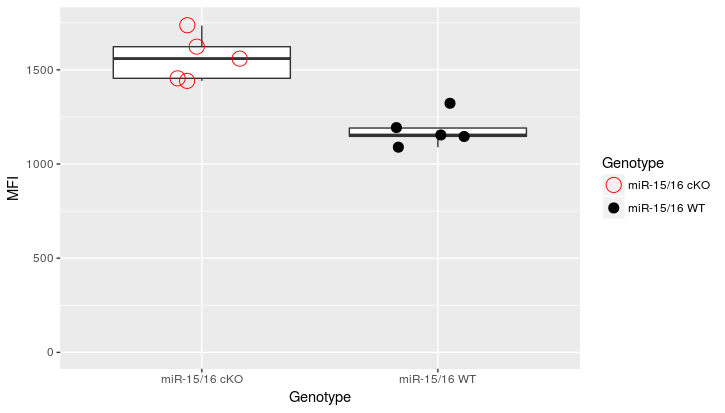
df2 <- data.frame(Genotype = c('miR-15/16 WT','miR-15/16 WT','miR-15/16 WT','miR-15/16 WT','miR-15/16 WT',
'miR-15/16 cKO','miR-15/16 cKO','miR-15/16 cKO','miR-15/16 cKO','miR-15/16 cKO'),
MFI=c(1191,1325,1089,1154,1147,1735,1441,1455,1560,1623))
color.groups <- c(`miR-15/16 WT`="black", `miR-15/16 cKO`="red")
shape.groups <- c(`miR-15/16 WT`=20, `miR-15/16 cKO`=21)
ggplot(data=df2, aes(x = Genotype, y = MFI)) +
geom_boxplot(outlier.shape = NA) +
geom_point(position = position_jitter(width = .2), size=5,
aes(color=Genotype, shape = Genotype)) +
ylim(0,max(df2$MFI)+10) +
scale_color_manual(values=color.groups) +
scale_shape_manual(values=shape.groups)
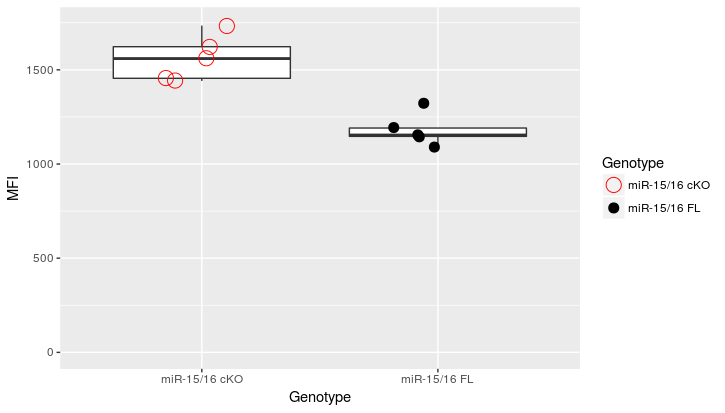
Update2:
df2 <- structure(list(Genotype = c("miR-15/16 FL", "miR-15/16 FL", "miR-15/16 FL",
"miR-15/16 FL", "miR-15/16 FL", "miR-15/16 cKO", "miR-15/16 cKO",
"miR-15/16 cKO", "miR-15/16 cKO", "miR-15/16 cKO"),
`Cells/SC/Live/CD8—,, CD4+/Foxp3-,Median,<BV421-A>,CD127` = c(1191L, 1325L, 1089L, 1154L, 1147L, 1735L, 1441L, 1455L, 1560L, 1623L)),
.Names = c("Genotype", "Cells/SC/Live/CD8—,, CD4+/Foxp3-,Median,<BV421-A>,CD127"),
row.names = c(NA, -10L), class = c("tbl_df", "tbl", "data.frame"))
colnames(df2) <- c("Genotype", "MFI")
color.groups <- c("black","red")
names(color.groups) <- unique(df2$Genotype)
shape.groups <- c(20, 21)
names(shape.groups) <- unique(df2$Genotype)
ggplot(data=df2, aes(x = Genotype, y = MFI)) +
geom_boxplot(outlier.shape = NA) +
geom_point(position = position_jitter(width = .2), size=5,
aes(color=Genotype, shape = Genotype)) +
ylim(0,max(df2$MFI)+10) +
scale_color_manual(values=color.groups) +
scale_shape_manual(values=shape.groups)
Stack Overflow用户
发布于 2017-09-25 17:27:24
这可能会让你开始:
ggplot(data=df2, aes(x = Genotype, y = MFI)) +
geom_boxplot(outlier.shape = NA) +
geom_jitter(aes(col = Genotype, shape = Genotype),position = position_jitter(width = .2), size=5) +
ylim(0,max(df2$MFI)+10)+
scale_shape_manual(values = c(1,16))+
scale_color_manual(values = c('red', 'black'))我发现这个网站非常有用:http://sape.inf.usi.ch/quick-reference/ggplot2/shape
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/46410688
复制相关文章
相似问题

