R闪亮图和d3heatmap兼容性问题
我试图添加一个交互式热图到我的闪亮的应用程序,但我也有互动的图形使用ggi相图。我目前正在使用d3heatmap包,但是热图不会在应用程序中呈现。我创建了一个玩具示例来说明这一点:
library(shiny)
library(ggiraph)
library(d3heatmap)
ui <- fluidPage(
d3heatmapOutput('d3'),
ggiraphOutput('gg')
)
server <- function(input, output, session) {
# Create heatmap
output$d3 <- renderD3heatmap({
d3heatmap(matrix(1:100, nrow = 100, ncol = 100))
})
# Create ggiraph
output$gg <- renderggiraph({
p <- ggplot(iris, aes(x = Sepal.Length, y = Petal.Width,
color = Species, tooltip = iris$Species) ) +
geom_point_interactive()
ggiraph(code = {print(p)})
})
}
shinyApp(ui = ui, server = server)加在一起,只有计时器呈现,而热图不呈现。但是,如果您注释掉了ggi相图代码,heatmap就会呈现。我试着改变加载包的顺序,但仍然没有效果。
我目前正在R3.2.2上运行(我必须使用这个版本,因为公司服务器只在这个版本上运行,我的经理和我都没有更新它的权限)。我尝试下载shinyheatmap、heatmaply和heatmap.2包,但是由于版本控制问题,安装失败了。
所以现在,我已经使用pheatmap创建了热图,但是它们不是交互的(也就是说,当我在单个单元格上悬停时无法得到值,并且不能放大)。有什么解决办法吗,还是有其他的交互式热图包可以工作?我希望避免将我的所有图形更改为平面图,因为我的代码中有很多这样的图形。
如果你需要其他信息,请告诉我。任何建议都将不胜感激!
回答 2
Stack Overflow用户
发布于 2017-06-22 22:54:11
(让你知道我是d3heatmap的作者)因为ggi相图使用的是d3.js版本4,d3heatmap使用的是D3.js版本3,所以两者之间存在冲突。我认为没有解决这一冲突的办法。
然而,使用ggplot2 2/ggi相图构建交互式热图并不是那么困难。见下文:
library(dplyr)
library(tidyr)
library(ggplot2)
library(ggiraph)
library(ggdendro)
# mydata <- cor(mtcars)
mydata <- matrix(runif(2500, min = -2, max = 2), ncol = 50)
row.names(mydata) <- paste0("row_", seq_len(nrow(mydata)))
colnames(mydata) <- paste0("col_", seq_len(ncol(mydata)))
# dendrogram for rows
hc <- hclust(dist(mydata), "ave")
dhr <- as.dendrogram(hc)
order_r <- rownames(mydata)[hc$order]
# dendrogram for columns
hc <- hclust(dist(t(mydata)), "ave")
dhc <- as.dendrogram(hc)
order_c <- colnames(mydata)[hc$order]
# the data
expr_set <- bind_cols(
data_frame(rowvar = rownames(mydata)),
as.data.frame(mydata)
)
expr_set <- gather(expr_set, colvar, measure, -rowvar)
expr_set$rowvar <- factor( expr_set$rowvar, levels = order_r )
expr_set$colvar <- factor( expr_set$colvar, levels = order_c )
expr_set <- arrange(expr_set, rowvar, colvar)
# get data for dendrograms - IMHO, ggdendro is the hero here...
data_c <- dendro_data(dhc, type = "rectangle")
data_c <- segment(data_c) %>% mutate(
y = y + length(order_r) + .5,
yend = yend + length(order_r) + .5
)
data_r <- dendro_data(dhr, type = "rectangle")
data_r <- segment(data_r)
data_r <- data_r %>%
mutate( x_ = y + length(order_c) + .5,
xend_ = yend + length(order_c) + .5,
y_ = x,
yend_ = xend )
expr_set <- expr_set %>%
mutate(
tooltip = sprintf("Row: %s<br/>Col: %s<br/>measure: %.02f",
rowvar, colvar, measure) ,
data_id = sprintf("%s_%s", rowvar, colvar)
)
# all data are tidy and can be now used with ggplot
p <- ggplot(data = expr_set, aes(x = colvar, y = rowvar) ) +
geom_tile_interactive(aes(fill = measure, tooltip = tooltip, data_id = data_id), colour = "white") +
scale_fill_gradient(low = "white", high = "#BC120A") +
geom_segment(
data = data_c,
mapping = aes(x = x, y = yend, xend = xend, yend = y),
colour = "gray20", size = .2) +
geom_segment(
data = data_r,
mapping = aes(x = x_, y = y_, xend = xend_, yend = yend_),
colour = "gray20", size = .2) +
coord_equal()
# cosmetics
p <- p + theme_minimal() +
theme(
legend.position = "right",
panel.grid.minor = element_line(color = "transparent"),
panel.grid.major = element_line(color = "transparent"),
axis.ticks.length = unit(2, units = "mm"),
plot.title = element_text(face = "bold", hjust = 0.5, size = 12),
axis.title = element_text(size = 9, colour = "gray30"),
axis.text.y = element_text(hjust = 1, size = 5, colour = "gray40"),
axis.text.x = element_text(angle = 90, hjust = 1, size = 5, colour = "gray40"),
legend.title=element_text(face = "bold", hjust = 0.5, size=8),
legend.text=element_text(size=6)
)
ggiraph(ggobj = p)
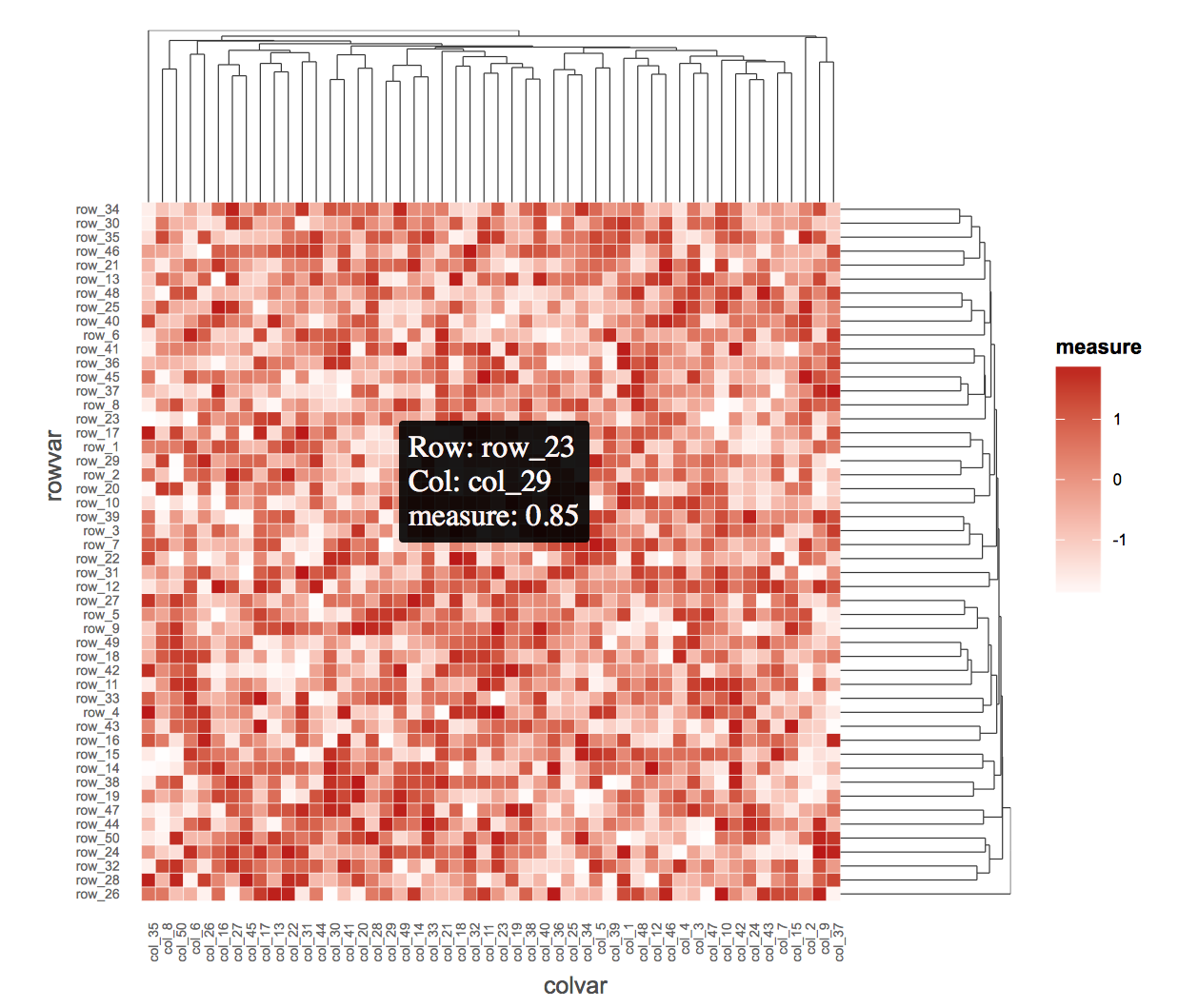
希望它能帮上忙
Stack Overflow用户
发布于 2019-10-04 14:48:41
我知道这个问题是在一段时间前回答的,但我遇到了同样的问题,我无法使用ggplot2,因为它只是缓慢地处理我的Shiny应用程序。heatmaply包的分配速度更快,实现起来更容易。我执行了一个迷你基准测试(n= 20)。ggplot2的平均时间为64秒。用heatmaply只花了2秒。这两种方法都使用了'ave'方法hclust.I,希望这是有帮助的。
下面是我使用的代码:
library(tidyr)
library(ggplot2)
library(ggiraph)
library(ggdendro)
library(heatmaply)
# mydata <- cor(mtcars)
create_data <- function(){
df <- matrix(runif(2500, min = -2, max = 2), ncol = 50)
row.names(df) <- paste0("row_", seq_len(nrow(df)))
colnames(df) <- paste0("col_", seq_len(ncol(df)))
return(df)
}
gg2heat <- function(mydata){
# dendrogram for rows
hc <- hclust(dist(mydata), "ave")
dhr <- as.dendrogram(hc)
order_r <- rownames(mydata)[hc$order]
# dendrogram for columns
hc <- hclust(dist(t(mydata)), "ave")
dhc <- as.dendrogram(hc)
order_c <- colnames(mydata)[hc$order]
# the data
expr_set <- bind_cols(
data_frame(rowvar = rownames(mydata)),
as.data.frame(mydata)
)
expr_set <- gather(expr_set, colvar, measure, -rowvar)
expr_set$rowvar <- factor( expr_set$rowvar, levels = order_r )
expr_set$colvar <- factor( expr_set$colvar, levels = order_c )
expr_set <- arrange(expr_set, rowvar, colvar)
# get data for dendrograms - IMHO, ggdendro is the hero here...
data_c <- dendro_data(dhc, type = "rectangle")
data_c <- segment(data_c) %>% mutate(
y = y + length(order_r) + .5,
yend = yend + length(order_r) + .5
)
data_r <- dendro_data(dhr, type = "rectangle")
data_r <- segment(data_r)
data_r <- data_r %>%
mutate( x_ = y + length(order_c) + .5,
xend_ = yend + length(order_c) + .5,
y_ = x,
yend_ = xend )
expr_set <- expr_set %>%
mutate(
tooltip = sprintf("Row: %s<br/>Col: %s<br/>measure: %.02f",
rowvar, colvar, measure) ,
data_id = sprintf("%s_%s", rowvar, colvar)
)
# all data are tidy and can be now used with ggplot
p <- ggplot(data = expr_set, aes(x = colvar, y = rowvar) ) +
geom_tile_interactive(aes(fill = measure, tooltip = tooltip, data_id = data_id), colour = "white") +
scale_fill_gradient(low = "white", high = "#BC120A") +
geom_segment(
data = data_c,
mapping = aes(x = x, y = yend, xend = xend, yend = y),
colour = "gray20", size = .2) +
geom_segment(
data = data_r,
mapping = aes(x = x_, y = y_, xend = xend_, yend = yend_),
colour = "gray20", size = .2) +
coord_equal()
# cosmetics
p <- p + theme_minimal() +
theme(
legend.position = "right",
panel.grid.minor = element_line(color = "transparent"),
panel.grid.major = element_line(color = "transparent"),
axis.ticks.length = unit(2, units = "mm"),
plot.title = element_text(face = "bold", hjust = 0.5, size = 12),
axis.title = element_text(size = 9, colour = "gray30"),
axis.text.y = element_text(hjust = 1, size = 5, colour = "gray40"),
axis.text.x = element_text(angle = 90, hjust = 1, size = 5, colour = "gray40"),
legend.title=element_text(face = "bold", hjust = 0.5, size=8),
legend.text=element_text(size=6)
)
ggiraph(ggobj = p)
}
htmp_gg <- c()
htmp_maply <-c()
for (i in 1:20){
df <- create_data()
time_gg <- (system.time(gg2heat(df)))[3]
htmp_gg<- append(htmp_gg, values = time_gg)
time_heatmaply <- (system.time(heatmaply::heatmaply(df, hclust_method = 'ave')))[3]
htmp_maply<- append(htmp_maply, values = time_heatmaply)
rm(df)
}
score <- data.frame(htmp_gg, htmp_maply)%>% gather(key = 'method', value = 'time')
p <- ggplot(score, aes(x = method, y = time, fill = method))+geom_violin()+ stat_summary(fun.y=median, geom="point", size=2, color="black")
print(p)https://stackoverflow.com/questions/44708029
复制相似问题

