将renderUI输入从一个闪亮的模块传递到另一个模块
将renderUI输入从一个闪亮的模块传递到另一个模块
提问于 2017-03-27 06:06:09
我试图模块化闪亮的代码,以上传CSV文件作为输入到scatterD3绘图。额外的UI控件将来自renderUI,以更改x变量和y变量.这只是来自How to organize large R Shiny apps?的Mikael回答的一个小小的修改,但我一直在挣扎,无法让这最后一点发挥作用。
对于这个数据集,我使用mtcar数据集469e9927/mtcars.csv。
## load libraries
library(shiny)
library(stringr)
library(scatterD3)
#source("/Users/echang/scratch/tmp/MSD_D3scatter/csvFile_Module.R")
csvFileInput <- function(id, label="CSV file") {
## Create namespace
ns<-NS(id)
tagList(
uiOutput(ns("controls"))
)
}
csvFileControl <- function(id){
ns <- NS(id)
tagList(
column(width=3, uiOutput(ns("ColName"))),
column(width=3, uiOutput(ns("ColEntry")))
)
}
csvFileUI <- function(id){
ns <- NS(id)
tagList(
uiOutput(ns("csvTable"))
)
}
## server module
csvFile <- function(input, output, session, stringsAsFactors) {
ns <- session$ns
## to reuse namespace, session must be first!!!
## User selected file
userFile <- reactive({
# If no file is selected, don't do anything
validate(need(input$file, message = FALSE))
input$file
})
dataframe <- reactive({
read.csv(
userFile()$datapath,
header = input$header,
sep=input$sep,
quote = input$quote,
stringsAsFactors = stringsAsFactors
)
})
# We can run observers in here if we want to
observe({
msg <- sprintf("File %s was uploaded", userFile()$name)
cat(msg, "\n")
})
output$controls <- renderUI({
## use taglist to keep everything together
tagList(
fileInput(ns('file'), 'Choose CSV file',
accept=c('txt/csv','text/comma-separated-values,text/plain','.csv')),
checkboxInput(ns('header'), 'Has heading', TRUE),
radioButtons(ns('sep'),'Separator', c(Comma=',',Semicolon=';',Tab='\t'), ','),
selectInput(ns('quote'),'Quote', c(None ='','Double Quote'='"','Single Quote'="'"),'"')
)
})
## use renderUI to display table
output$csvTable <- renderUI({
output$table <- renderDataTable(dataframe())
dataTableOutput(ns("table"))
})
## Column Name
output$ColName <- renderUI({
df <- dataframe()
if (is.null(df)) return(NULL)
items=names(df)
names(items)=items
tagList(
selectInput(ns("xvar"), "Column Names", items),
selectInput(ns("yvar"), "Column Names", items)
)
})
## Column Entry
output$ColEntry <- renderUI({
df <- dataframe()
if (is.null(input$col)) return(NULL)
tagList(
selectInput(ns("entry"), "Entry Names", df[,input$xvar])
)
})
# Return the reactive that yields the data frame
return(dataframe)
}## End of module
## scatterD3 module -------------------------------------------------------------
D3scatterUI <- function(id){
ns<-NS(id)
tagList(
scatterD3Output(ns("scatterplot1"))
)
}
D3scatter <- function(input,output,session,data,xvar,yvar){
ns <- session$ns
output$scatterplot1 <- renderScatterD3({
#scatterD3(data = data, x=mpg, y=carb,
scatterD3(data = data, x=xvar, y=yvar,
labels_size= 9, point_opacity = 1,
#col_var=cyl, symbol_var= data$Assay,
#lab= paste(mpg, carb, sep="|") , lasso=TRUE,
#xlab= "IFN-γ", ylab= "IL-10",
#click_callback = "function(id, index) {
# alert('scatterplot ID: ' + id + ' - Point index: ' + index)
# }",
transitions= T)
})
}
## Shiny ######################################################################
ui <- fluidPage(
titlePanel("Upload"),
tabsetPanel(type="tabs",
tabPanel("tab1",
sidebarLayout(
sidebarPanel(csvFileInput("basic")),
mainPanel(csvFileUI("basic"))
)
),
tabPanel("tab2",
tagList(
fluidRow(csvFileControl("basic")),
fluidRow(D3scatterUI("first"))
)
)
)
)
server <- function(input, output, session) {
## Option 1. CSV uploaded file
datafile <- callModule(csvFile, "basic", stringsAsFactors = FALSE)
## Option 2. mtcar data loaded at start
#datafile <- reactive({mtcars}) ## data loaded at runApp()
#callModule(csvFile, "basic")
xvar <- reactive(input$xvar)
yvar <- reactive(input$yvar)
callModule(D3scatter, "first", datafile(), xvar, yvar)
}
shinyApp(ui, server)我还参考了https://itsalocke.com/shiny-module-design-patterns-pass-module-input-to-other-modules/的闪亮模块设计。
我观看了网络研讨会,但无法正确理解我的逻辑。https://www.rstudio.com/resources/webinars/understanding-shiny-modules/任何帮助都将不胜感激!
回答 1
Stack Overflow用户
回答已采纳
发布于 2017-03-27 11:50:22
好吧,这确实有点困难,因为使用模块并不是很简单。你们很亲密..。您的主要问题不是将reactives的全部打包到一个列表中,并将它们传递到需要它们的地方。
我做了以下修改:
csvFile:在csvFile服务器模块中声明了附加的反应性函数xvar和yvar,类似于您已经为dataframe所做的工作。csvFile:将所有所需的反应性打包为列表,并将其作为返回值返回,如文章中的设计模式链接所述。(谢谢你,斯蒂芬·洛克)server:将该列表向下传递到callModule(D3scatter,... )中,同样如该链接中所述。D3scatter:通过调用scatterD3来使用从指定数据中提取的向量来重构一点。这是因为我无法让它作为列说明符处理字符串(但肯定有一种方法)。
下面是上面更改的代码部分:
csvFile服务器模块
csvFile <- function(input, output, session, stringsAsFactors) {
ns <- session$ns
## to reuse namespace, session must be first!!!
## User selected file
userFile <- reactive({
# If no file is selected, don't do anything
validate(need(input$file, message = FALSE))
input$file
})
dataframe <- reactive({
read.csv(
userFile()$datapath,
header = input$header,
sep=input$sep,
quote = input$quote,
stringsAsFactors = stringsAsFactors
)
})
# We can run observers in here if we want to
observe({
msg <- sprintf("File %s was uploaded", userFile()$name)
cat(msg, "\n")
})
xvar <- reactive({input[[ "xvar" ]] })
yvar <- reactive({input[[ "yvar" ]] })
output$controls <- renderUI({
## use taglist to keep everything together
tagList(
fileInput(ns('file'), 'Choose CSV file',
accept=c('txt/csv','text/comma-separated-values,text/plain','.csv')),
checkboxInput(ns('header'), 'Has heading', TRUE),
radioButtons(ns('sep'),'Separator', c(Comma=',',Semicolon=';',Tab='\t'), ','),
selectInput(ns('quote'),'Quote', c(None ='','Double Quote'='"','Single Quote'="'"),'"')
)
})
## use renderUI to display table
output$csvTable <- renderUI({
output$table <- renderDataTable(dataframe())
dataTableOutput(ns("table"))
})
## Column Name
output$ColName <- renderUI({
df <- dataframe()
if (is.null(df)) return(NULL)
items=names(df)
print(items)
names(items)=items
tagList(
selectInput(ns("xvar"), "Column Names", items),
selectInput(ns("yvar"), "Column Names", items)
)
})
## Column Entry
output$ColEntry <- renderUI({
df <- dataframe()
if (is.null(input$col)) return(NULL)
tagList(
selectInput(ns("entry"), "Entry Names", df[,input$xvar])
)
})
rlist <- list(dataframe=dataframe,xvar=xvar,yvar=yvar)
# Return the reactive that yields the data frame
return(rlist)
}## End of module服务器
server <- function(input, output, session) {
## Option 1. CSV uploaded file
rlist <- callModule(csvFile, "basic", stringsAsFactors = FALSE)
## Option 2. mtcar data loaded at start
#datafile <- reactive({mtcars}) ## data loaded at runApp()
#callModule(csvFile, "basic")
callModule(D3scatter, "first", rlist)
}D3scatter
D3scatter <- function(input,output,session,rlist){
ns <- session$ns
output$scatterplot1 <- renderScatterD3({
#scatterD3(data = data, x=mpg, y=carb,
mtdf <- rlist$dataframe()
x <- mtdf[[rlist$xvar()]]
y <- mtdf[[rlist$yvar()]]
scatterD3(x=x,y=y,
labels_size= 9, point_opacity = 1,
#col_var=cyl, symbol_var= data$Assay,
#lab= paste(mpg, carb, sep="|") , lasso=TRUE,
#xlab= "IFN-γ", ylab= "IL-10",
#click_callback = "function(id, index) {
# alert('scatterplot ID: ' + id + ' - Point index: ' + index)
# }",
transitions= T)
})
}然后它起了作用:
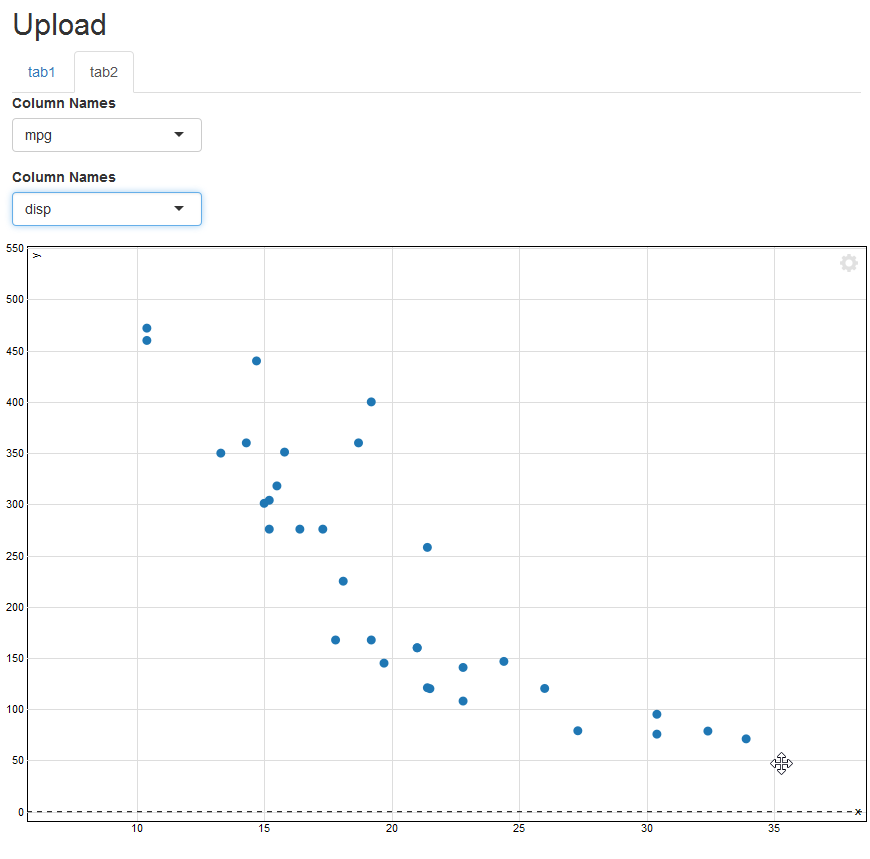
这是所有正在运行的代码,以防我忘了某个地方的更改,或者有人只想运行它。顺便说一下,散点图从一个点到另一个点变化的方式很酷.它不断地变形,具有动画般的效果。不同寻常。
一个文件中的整个应用程序
## load libraries
library(shiny)
library(stringr)
library(scatterD3)
#source("/Users/echang/scratch/tmp/MSD_D3scatter/csvFile_Module.R")
csvFileInput <- function(id, label="CSV file") {
## Create namespace
ns<-NS(id)
tagList(
uiOutput(ns("controls"))
)
}
csvFileControl <- function(id){
ns <- NS(id)
tagList(
column(width=3, uiOutput(ns("ColName"))),
column(width=3, uiOutput(ns("ColEntry")))
)
}
csvFileUI <- function(id){
ns <- NS(id)
tagList(
uiOutput(ns("csvTable"))
)
}
## server module
csvFile <- function(input, output, session, stringsAsFactors) {
ns <- session$ns
## to reuse namespace, session must be first!!!
## User selected file
userFile <- reactive({
# If no file is selected, don't do anything
validate(need(input$file, message = FALSE))
input$file
})
dataframe <- reactive({
read.csv(
userFile()$datapath,
header = input$header,
sep=input$sep,
quote = input$quote,
stringsAsFactors = stringsAsFactors
)
})
# We can run observers in here if we want to
observe({
msg <- sprintf("File %s was uploaded", userFile()$name)
cat(msg, "\n")
})
xvar <- reactive({input[[ "xvar" ]] })
yvar <- reactive({input[[ "yvar" ]] })
output$controls <- renderUI({
## use taglist to keep everything together
tagList(
fileInput(ns('file'), 'Choose CSV file',
accept=c('txt/csv','text/comma-separated-values,text/plain','.csv')),
checkboxInput(ns('header'), 'Has heading', TRUE),
radioButtons(ns('sep'),'Separator', c(Comma=',',Semicolon=';',Tab='\t'), ','),
selectInput(ns('quote'),'Quote', c(None ='','Double Quote'='"','Single Quote'="'"),'"')
)
})
## use renderUI to display table
output$csvTable <- renderUI({
output$table <- renderDataTable(dataframe())
dataTableOutput(ns("table"))
})
## Column Name
output$ColName <- renderUI({
df <- dataframe()
if (is.null(df)) return(NULL)
items=names(df)
print(items)
names(items)=items
tagList(
selectInput(ns("xvar"), "Column Names", items),
selectInput(ns("yvar"), "Column Names", items)
)
})
## Column Entry
output$ColEntry <- renderUI({
df <- dataframe()
if (is.null(input$col)) return(NULL)
tagList(
selectInput(ns("entry"), "Entry Names", df[,input$xvar])
)
})
rlist <- list(dataframe=dataframe,xvar=xvar,yvar=yvar)
# Return the reactive that yields the data frame
return(rlist)
}## End of module
## scatterD3 module -------------------------------------------------------------
D3scatterUI <- function(id){
ns<-NS(id)
tagList(
scatterD3Output(ns("scatterplot1"))
)
}
D3scatter <- function(input,output,session,rlist){
ns <- session$ns
output$scatterplot1 <- renderScatterD3({
#scatterD3(data = data, x=mpg, y=carb,
mtdf <- rlist$dataframe()
x <- mtdf[[rlist$xvar()]]
y <- mtdf[[rlist$yvar()]]
scatterD3(x=x,y=y,
labels_size= 9, point_opacity = 1,
#col_var=cyl, symbol_var= data$Assay,
#lab= paste(mpg, carb, sep="|") , lasso=TRUE,
#xlab= "IFN-γ", ylab= "IL-10",
#click_callback = "function(id, index) {
# alert('scatterplot ID: ' + id + ' - Point index: ' + index)
# }",
transitions= T)
})
}
## Shiny ######################################################################
ui <- fluidPage(
titlePanel("Upload"),
tabsetPanel(type="tabs",
tabPanel("tab1",
sidebarLayout(
sidebarPanel(csvFileInput("basic")),
mainPanel(csvFileUI("basic"))
)
),
tabPanel("tab2",
tagList(
fluidRow(csvFileControl("basic")),
fluidRow(D3scatterUI("first"))
)
)
)
)
server <- function(input, output, session) {
## Option 1. CSV uploaded file
rlist <- callModule(csvFile, "basic", stringsAsFactors = FALSE)
## Option 2. mtcar data loaded at start
#datafile <- reactive({mtcars}) ## data loaded at runApp()
#callModule(csvFile, "basic")
callModule(D3scatter, "first", rlist)
}
shinyApp(ui, server)页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/43038967
复制相关文章
相似问题

