GenomicRanges添加覆盖范围
我正在研究RNA seq数据,并试图按基因型绘制平均覆盖范围,类似于这里所做的工作。
每个基因型的RNA seq覆盖率(资料来源: pickrell等人,自然,2010年)
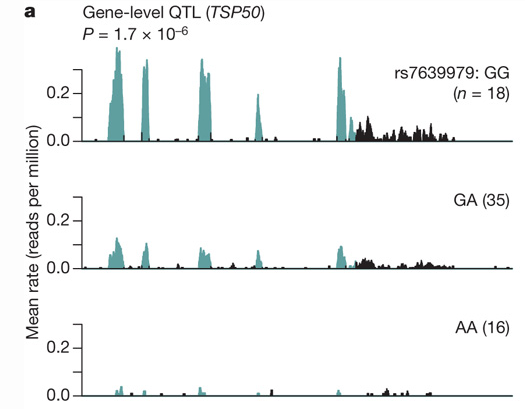
为此,我有来自100个个人的大型文件,其中包含来自RNA-seq数据的覆盖信息(在特定区域),并且我在R中以GenomicRanges对象的形式读取这些信息。
这给了我一些GRanges对象,比如在下面的玩具示例中获得的对象:
gr1=GRanges(seqname=1,range=IRanges(start=c(1,5,10,15,30,55),end=c(4,9,14,29,39,60)) gr1$cov=c(3,1,8,6,2,10) gr2=GRanges(seqname=1,range=IRanges(start=c(3,20,24),end=c(7,23,26) gr2$cov=c(3,5,3) Start=unique(排序(c(range(Gr1)@start,range(Gr2)@start) gr1
GRanges object with 6 ranges and 1 metadata column:
seqnames ranges strand | cov
<Rle> <IRanges> <Rle> | <numeric>
1 [ 1, 4] * | 3
1 [ 5, 9] * | 1
1 [10, 14] * | 8
1 [15, 29] * | 6
1 [30, 39] * | 2
1 [55, 60] * | 10
-------
seqinfo: 1 sequence from an unspecified genome; no seqlengthsgr2
GRanges object with 3 ranges and 1 metadata column:
seqnames ranges strand | cov
<Rle> <IRanges> <Rle> | <numeric>
1 [ 3, 7] * | 3
1 [20, 23] * | 5
1 [24, 26] * | 3
-------
seqinfo: 1 sequence from an unspecified genome; no seqlengths问题是,我每个人都有这些(gr1和gr2是两个不同的个体),我想将它们组合起来,创建一个基因组范围对象,给出每个位置的总体覆盖范围,包括1和2,如下所示:
gr3
GRanges object with 6 ranges and 1 metadata column:
seqnames ranges strand | cov
<Rle> <IRanges> <Rle> | <numeric>
1 [ 1, 2] * | 3
1 [ 3, 4] * | 6 (=3+3)
1 [ 5, 7] * | 4 (=1+3)
1 [ 8, 9] * | 1
1 [10, 14] * | 8
1 [15, 19] * | 6
1 [20, 23] * | 11 (=6+5)
1 [24, 26] * | 9 (=6+3)
1 [27, 29] * | 6
1 [30, 39] * | 2
1 [55, 60] * | 10 有人知道一个简单的方法吗?还是我注定了?
谢谢你的回答。
PS:我的数据不是搁浅的,但如果你有它作为搁浅数据,那就更好了。
PPS:理想情况下,我也希望能够对乘法进行处理,或者应用带有两个参数x和y的任何函数,而不是简单地添加覆盖范围。
回答 1
Stack Overflow用户
发布于 2018-01-09 21:49:08
已经快一年了,但这是我的答案,供将来参考。
每当我找不到函数直接执行像这个任务这样的任务时,我只需将GRanges对象展开为单bp解析。这允许我对元数据列执行任何必需的操作,将它们视为简单的data.frame列,因为IRanges现在在两个Granges对象之间匹配。
在这个问题的具体案例中,有以下几点。
### Sort seqlevels
# (not necessary here, but in real world examples,
# with multiple sequences, you will want to do this)
gr1 <- sort(GenomeInfoDb::sortSeqlevels(gr1))
gr2 <- sort(GenomeInfoDb::sortSeqlevels(gr2))
### Add seqlengths
# (this corresponds to the actual sequence lengths;
# here we use the highest position between the two objects: 60)
seqlengths(gr1) <- 60
### Make 1-bp tiles covering the genome
# (using either one of gr1 and gr2 as a reference)
bins <- GenomicRanges::tileGenome(GenomeInfoDb::seqlengths(gr1),
tilewidth=1,
cut.last.tile.in.chrom=TRUE)
### Get coverage signal as Rle object
gr1_cov <- coverage(gr1, weight="cov")
gr2_cov <- coverage(gr2, weight="cov")
### Get average coverage in each bin
# (since the bins are 1-bp wide, this just keeps the original coverage value)
gr1_bins <- GenomicRanges::binnedAverage(bins, gr1_cov, "binned_cov")
gr2_bins <- GenomicRanges::binnedAverage(bins, gr2_cov, "binned_cov")
### Make final object:
# We can now sum the values in the metadata columns
# Addressing the PPS, you could do any other operation or apply a function
gr3 <- gr1_bins
gr3$binned_cov <- gr1_bins$binned_cov + gr2_bins$binned_cov这产生了最终的GRanges对象的单bp分辨率.
> gr3
GRanges object with 60 ranges and 1 metadata column:
seqnames ranges strand | binned_cov
<Rle> <IRanges> <Rle> | <numeric>
[1] 1 [1, 1] * | 3
[2] 1 [2, 2] * | 3
[3] 1 [3, 3] * | 6
[4] 1 [4, 4] * | 6
[5] 1 [5, 5] * | 4
... ... ... ... . ...
[56] 1 [56, 56] * | 10
[57] 1 [57, 57] * | 10
[58] 1 [58, 58] * | 10
[59] 1 [59, 59] * | 10
[60] 1 [60, 60] * | 10
-------
seqinfo: 1 sequence from an unspecified genome为了压缩它并得到问题中的确切gr3,我们可以执行以下操作。
### Compress back to variable-width IRanges (by cov)
gr3_Rle <- coverage(gr3, weight='binned_cov')
gr3 <- as(gr3_Rle, "GRanges")
### Drop 0-score rows
gr3 <- gr3[gr3$score > 0]
### Rename metadata column
names(mcols(gr3)) <- 'cov'
> gr3
GRanges object with 11 ranges and 1 metadata column:
seqnames ranges strand | cov
<Rle> <IRanges> <Rle> | <numeric>
[1] 1 [ 1, 2] * | 3
[2] 1 [ 3, 4] * | 6
[3] 1 [ 5, 7] * | 4
[4] 1 [ 8, 9] * | 1
[5] 1 [10, 14] * | 8
[6] 1 [15, 19] * | 6
[7] 1 [20, 23] * | 11
[8] 1 [24, 26] * | 9
[9] 1 [27, 29] * | 6
[10] 1 [30, 39] * | 2
[11] 1 [55, 60] * | 10
-------
seqinfo: 1 sequence from an unspecified genomehttps://stackoverflow.com/questions/42400257
复制相似问题

