在R中同步和绘制两个timeseries数据集
我有两组数据来自一个人在不同环境温度下的实验。P1代表患者的生理反应数据,P1IAQ代表实验期间的环境监测数据。P1每秒记录32次数据,但P1IAQ每10秒记录一次数据。
head(P1IAQ)
Time RH Temp CO2
1 12:04:07 44.2 19.89 664
2 12:04:17 44.2 19.89 664
3 12:04:27 44.2 19.89 665
4 12:04:37 44.2 19.89 665
5 12:04:47 44.2 19.89 666
6 12:04:57 44.2 19.89 668
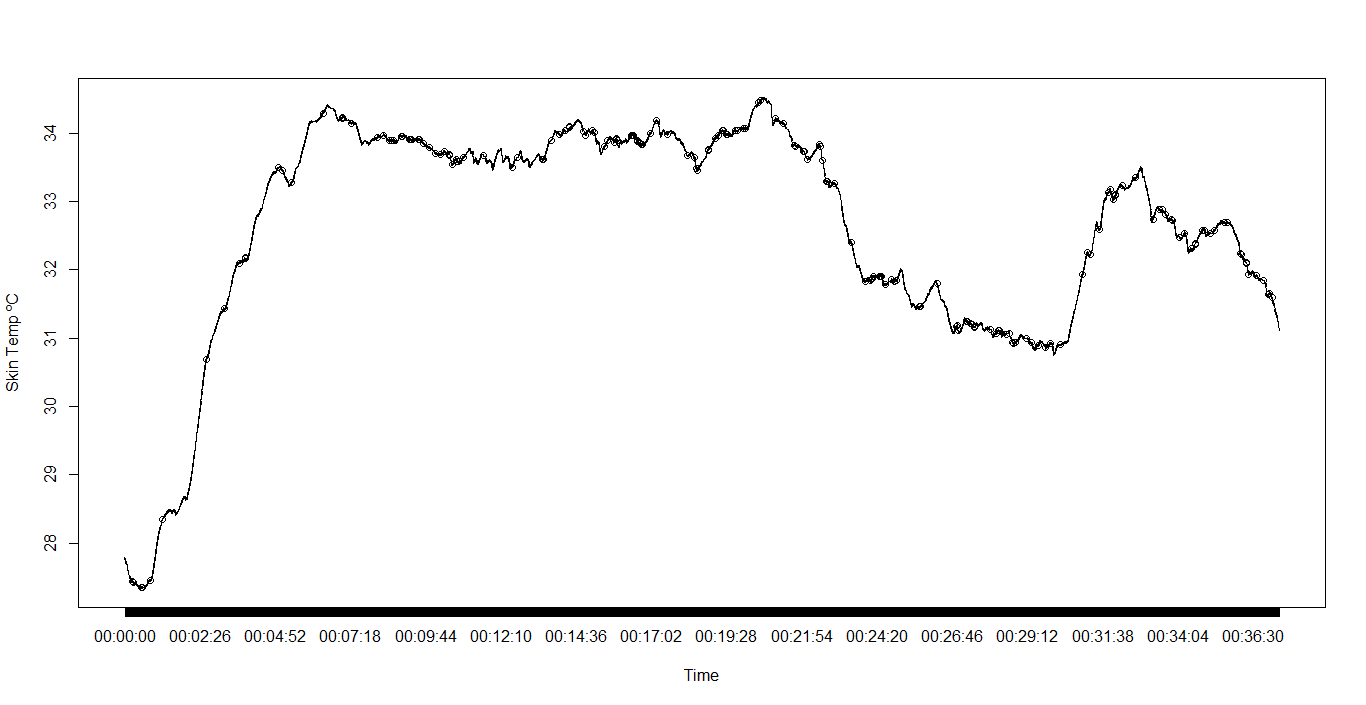
head(P1)
Time SkinTemp HeartRate RespirationRate
1 00:00:00 27.781 70 10
2 00:00:00 27.780 70 10
3 00:00:00 27.779 70 10
4 00:00:00 27.779 70 10
5 00:00:00 27.778 70 10
6 00:00:00 27.777 70 10我的问题是,P1上的时间戳是错误的。我如何将它们放在同一个图表上,以查看在环境温度降低之后,SkinTemp是否有时间延迟?
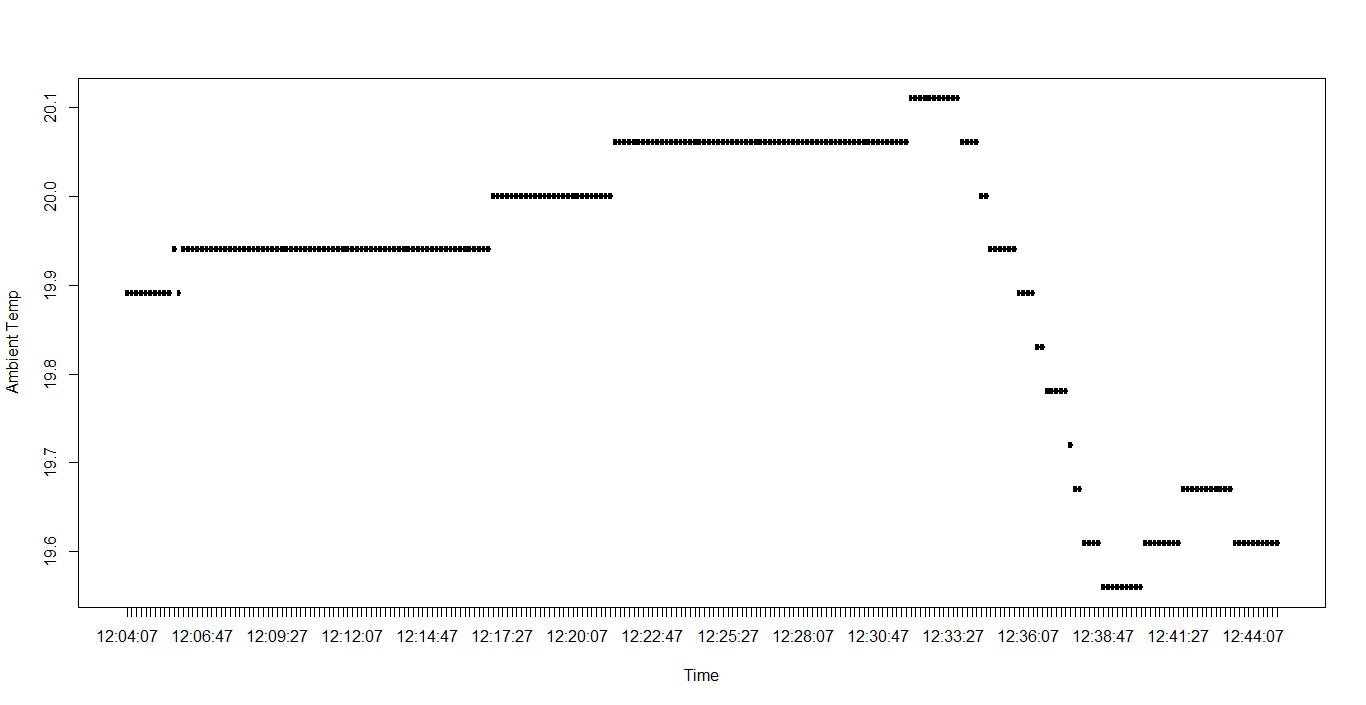
编辑: dput for P1IAQ我为环境数据添加了前20个值。我认为最好是从所有的值中减去12:04:07,使开始时间为00:00:00。我试过看润滑油了。
library(lubridate)
P1IAQ$Time<-hms(P1IAQ$Time)这个datetime帖子看起来很有趣,但它是用来绘制日期的,而不是实际更改日期的。Plotting data against time in R
dput(P1IAQ)
structure(list(Time = structure(1:19, .Label = c("12:04:07",
"12:04:17", "12:04:27", "12:04:37", "12:04:47", "12:04:57", "12:05:07",
"12:05:17", "12:05:27", "12:05:37", "12:05:47", "12:05:57", "12:06:07",
"12:06:17", "12:06:27", "12:06:37", "12:06:47", "12:06:57", "12:07:07"
), class = "factor"), RH = c(44.2, 44.2, 44.2, 44.2, 44.2, 44.2,
44.2, 44.2, 44.1, 44.1, 44.2, 44.2, 44.2, 44.3, 44.2, 44.2, 44.2,
44.3, 44.3), Temp = c(19.89, 19.89, 19.89, 19.89, 19.89, 19.89,
19.89, 19.89, 19.89, 19.89, 19.94, 19.89, 19.94, 19.94, 19.94,
19.94, 19.94, 19.94, 19.94), CO2 = c(664L, 664L, 665L, 665L,
666L, 668L, 668L, 669L, 667L, 670L, 670L, 672L, 675L, 677L, 682L,
684L, 685L, 686L, 687L)), .Names = c("Time", "RH", "Temp", "CO2"
), class = "data.frame", row.names = c(NA, -19L))编辑:我使用lubridate:同步了时间
P1IAQ$Time<-period_to_seconds(hms(as.character(P1IAQ$Time))-hms("12:04:07"))
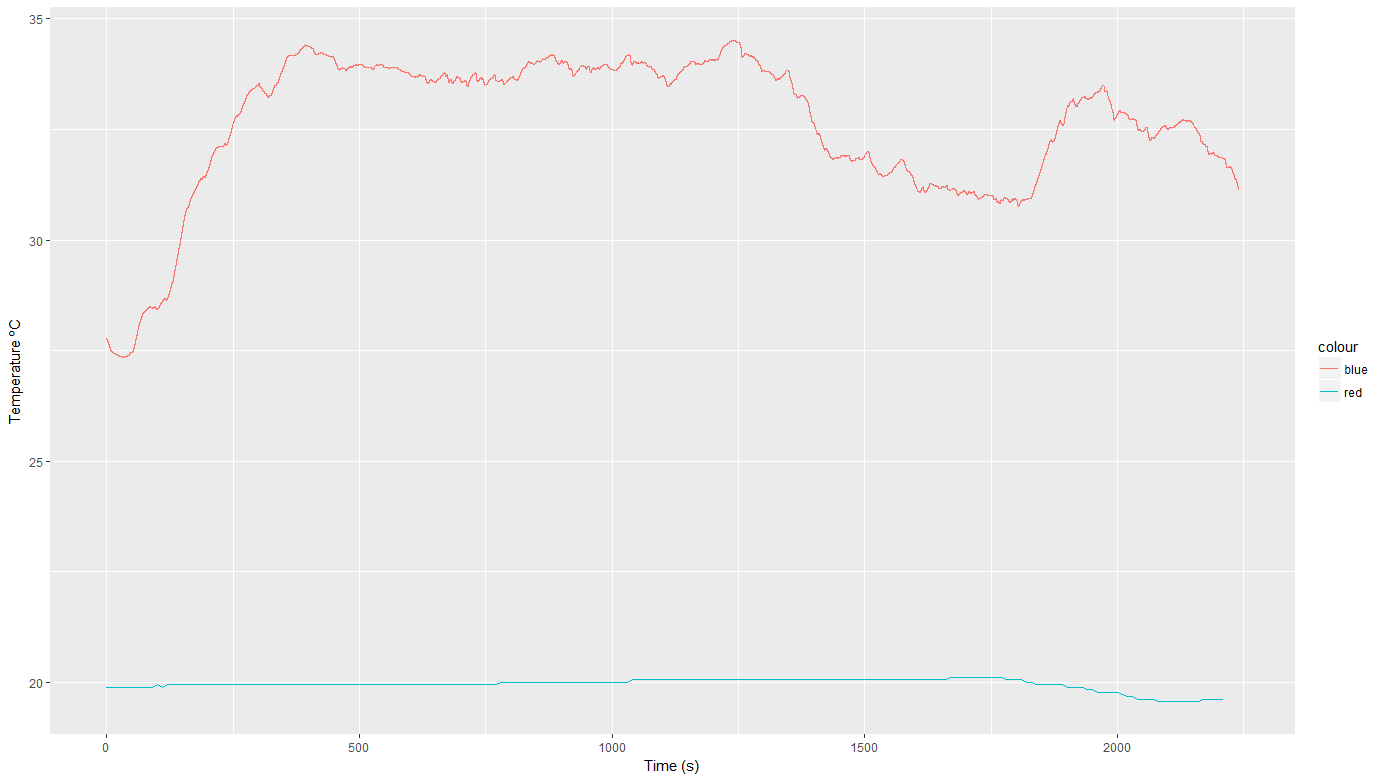
P1$Time<-period_to_seconds(hms(as.character(P1$Time)))但现在把它们勾勒在一起是很棘手的。我试过ggplot2,但我不能得到两个垂直轴。任何想法
ggplot() +
geom_line(data = P1IAQ, aes(x = Time, y = Temp, color = "red")) +
geom_line(data = P1, aes(x = Time, y = Temp, color = "blue")) +
xlab('Time (s)') +
ylab('Temperature ºC')
回答 2
Stack Overflow用户
发布于 2016-12-28 18:19:49
如果您只需要每个数据帧中的时间在一个通用的比例上,您可以将它们转换为自实验开始以来经过的数字秒,而不必担心日期或时间类。然后,您可以根据共同的时间尺度加入这两个数据帧。
我使用了您的P1IAQ数据示例,并创建了假P1数据以配合它。我的P1中的P1可能与实际数据的格式不同。如果您发布了一个P1示例,我可以调整下面的示例以适应您的实际数据。
library(dplyr)
library(reshape2)
library(hms)
library(zoo)
library(ggplot2)
theme_set(theme_light())
# Fake P1 data frame
set.seed(10)
n=32*60*3 + 1
P1 = data.frame(Time=as.POSIXct(seq(0,180,length.out=n), origin=as.Date("2016-05-01"), tz="GMT"),
SkinTemp = round(cumsum(rnorm(n, 0, 0.01)) + 27.78, 2),
RespirationRate=round(rnorm(n, 10, 0.5)))将P1$Time和P1IAQ$Time转换为等于实验开始以来经过的秒数的数值。(请注意,您发布的数据中的P1IAQ$Time是一个factor,因此我在进一步处理之前转换为字符):
P1$nTime = as.numeric(as.hms(P1$Time))
P1IAQ$nTime = as.numeric(as.hms(as.character(P1IAQ$Time)))
P1IAQ$nTime = P1IAQ$nTime - min(P1IAQ$nTime)通过P1和P1IAQ加入nTime
P1j = full_join(P1, P1IAQ, by="nTime", suffix=c("_P1","_P1IAQ")) %>%
# Make sure joined data frame is sorted by nTime
arrange(nTime) %>%
# Fill missing values with Last One Carried Forward
mutate_at(vars(Time_P1IAQ, RH, Temp, CO2), na.locf)将数据从宽格式转换为长格式后绘制:
ggplot(P1j %>% select(Time_P1IAQ, nTime, Skin=SkinTemp, Ambient=Temp) %>%
# Convert from wide to long format for plotting
melt(id.var=c("Time_P1IAQ", "nTime")),
aes(nTime, value, group=Time_P1IAQ)) +
geom_line() +
facet_grid(variable ~ ., scales="free_y") +
scale_y_continuous(expand=c(0.5,0)) +
labs(x="Elapsed Time (sec)", y=expression(Temperature~"("*degree*C*")"))
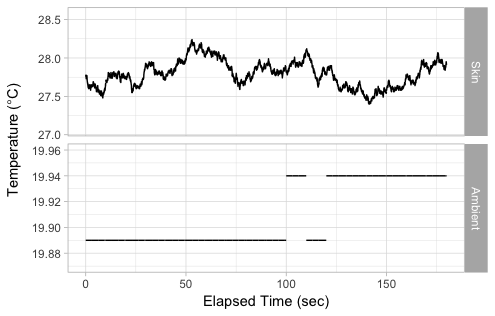
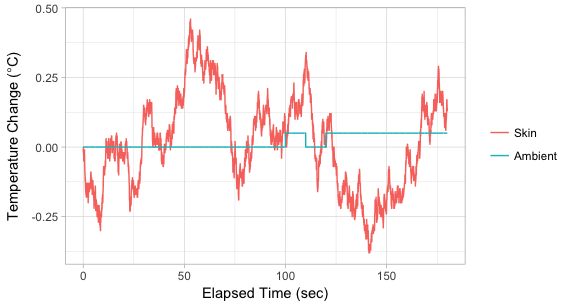
另一种选择是绘制实验开始时的温度变化图。这样,您就可以在同一面板上拥有两条线,而不必处理它们位于不同的位置:
ggplot(P1j %>% select(Time_P1IAQ, nTime, Skin=SkinTemp, Ambient=Temp) %>%
# Convert from wide to long format for plotting
melt(id.var=c("Time_P1IAQ", "nTime")) %>%
# Convert temperatures to difference from starting values
group_by(variable) %>%
mutate(value = value - value[nTime==min(nTime)]),
aes(nTime, value, colour=variable)) +
geom_line() +
labs(x="Elapsed Time (sec)", y=expression(Temperature~Change~"("*degree*C*")"),
colour="")
Stack Overflow用户
发布于 2016-12-28 16:32:39
我没有你的数据,但我会准备一些类似的.在这种情况下,P1的长度与P1IAQ不同:
library(ggplot2)
#I create a sample of your data
P1<-data.frame(1:10,51:60)
P1IAQ<-data.frame(1:8,1:8)
colnames(P1)<-c("Time","Temp")
colnames(P1IAQ)<-c("Time","Temp")
# I cathegory your data for plot
df = data.frame(Time=c(P1$Time,P1IAQ$Time), values=c(P1$Temp,P1IAQ$Temp),type=c(rep("P1",length(P1$Time)),rep("P1IAQ",length(P1IAQ$Time))))
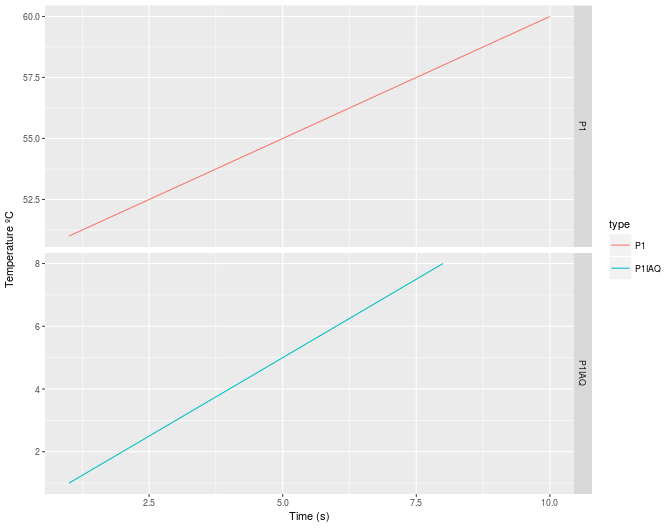
ggplot(data=df, aes(x=Time, y=values, color=type)) +
geom_line() +
facet_grid(type ~ ., scales="free") +
xlab('Time (s)') +
ylab('Temperature ºC')
https://stackoverflow.com/questions/41352042
复制相似问题

