R interaction.plot
R interaction.plot
提问于 2016-11-08 19:04:09
我想为我的数据绘制一个interaction.plot。
with(data, interaction.plot(biofeedback, diet, as.numeric(drug),
xlab="Biofeedback", ylab="Drug", ylim = c(0, 4), col = 2:3))结果:
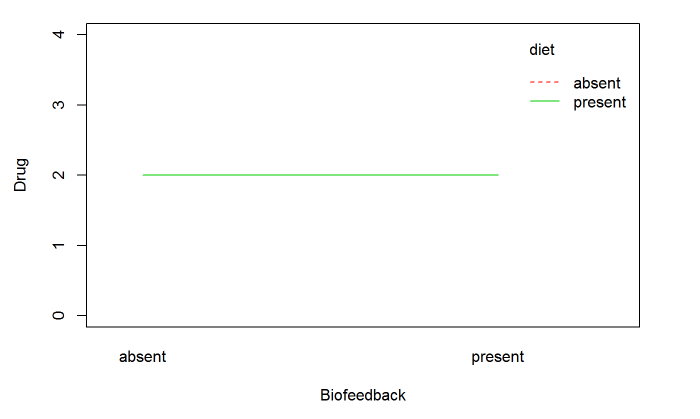
但显然这是不对的,因为我的数据是:
## biofeedback diet drug pressure
## 1 present absent 1 170
## 2 present absent 1 175
## 3 present absent 1 165
...
## 7 present present 1 161
## 8 present present 1 173
## 9 present present 1 157
...
## 19 present present 2 164
## 20 present present 2 166
## 21 present present 2 159
...这个数据对于缺席有不同的drugs,对于现在的diet有不同的Biofeedback。对于缺席的drugs,它有一些diet。这一切都没有在情节上显示出来。为什么?
dput(data)
## structure(list(biofeedback = structure(c(2L, 2L, 2L, 2L, 2L,
## 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
## 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L,
## 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
## 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
## 1L, 1L, 1L), .Label = c("absent", "present"), class = "factor"),
## diet = structure(c(1L, 1L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L,
## 2L, 2L, 1L, 1L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 1L,
## 1L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 1L, 1L, 1L,
## 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 1L, 1L, 1L, 1L, 1L, 2L,
## 2L, 2L, 2L, 2L, 2L, 1L, 1L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L,
## 2L, 2L), .Label = c("absent", "present"), class = "factor"),
## drug = structure(c(1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
## 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 3L,
## 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 1L, 1L, 1L, 1L,
## 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
## 2L, 2L, 2L, 2L, 2L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L,
## 3L, 3L), .Label = c("1", "2", "3"), class = "factor"), pressure = c(170L,
## 175L, 165L, 180L, 160L, 158L, 161L, 173L, 157L, 152L, 181L,
## 190L, 186L, 194L, 201L, 215L, 219L, 209L, 164L, 166L, 159L,
## 182L, 187L, 174L, 180L, 187L, 199L, 170L, 204L, 194L, 162L,
## 184L, 183L, 156L, 180L, 173L, 173L, 194L, 197L, 190L, 176L,
## 198L, 164L, 190L, 169L, 164L, 176L, 175L, 189L, 194L, 217L,
## 206L, 199L, 195L, 171L, 173L, 196L, 199L, 180L, 203L, 202L,
## 228L, 190L, 206L, 224L, 204L, 205L, 199L, 170L, 160L, 179L,
## 179L)), .Names = c("biofeedback", "diet", "drug", "pressure"
## ), row.names = c(NA, -72L), class = "data.frame")回答 1
Stack Overflow用户
回答已采纳
发布于 2016-11-08 19:47:28
看上去没有什么错误。每个组的平均值是2.0,这就是情节所显示的。
> df$drug <- as.numeric(levels(df$drug))[df$drug]
> library(plyr)
> ddply(df, .(biofeedback, diet), summarise, val = mean(drug))
biofeedback diet val
1 absent absent 2
2 absent present 2
3 present absent 2
4 present present 2页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/40494567
复制相关文章
相似问题

