在多个文件中使用ggplot绘制数据
在多个文件中使用ggplot绘制数据
提问于 2016-07-31 10:46:49
我有一个时间序列数据文件,有4种代谢产物A,B,AE和E的浓度随时间的变化。我有许多这种类型的数据文件(大约100个)。我想绘制一个图表中所有文件中所有四个代谢物的时间序列。每种代谢物都被指定为特定的颜色。
我编译了下面的代码,但是它只在一个文件(最后一个)中绘制数据。我认为这是因为当我调用ggplot()时,它会创建一个新的绘图。我试着在四个循环之外创建一个情节,但没有成功。
p = NULL
for(i in 1:length(filesToProcess)){
fileName = filesToProcess[i]
fileContent = read.csv(fileName)
#fileContent$Time <- NULL
p <- ggplot()+
geom_line(data = fileContent, aes(x = Time, y = A, color = "A"), size =0.8) +
geom_line(data = fileContent, aes(x = Time, y = B, color = "B"), size =0.8) +
geom_line(data = fileContent, aes(x = Time, y = AE, color = "AE"), size =0.8) +
geom_line(data = fileContent, aes(x = Time, y = E, color = "E"), size =0.8) +
xlab('Time') +
ylab('Metabolite Concentration')+
ggtitle('Step Scan') +
labs(color="Metabolites")
}
plot(p)下面是图
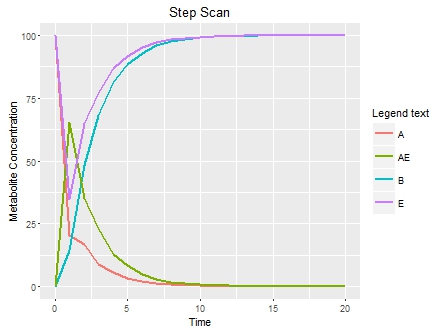
示例文件可以找到 here
回答 2
Stack Overflow用户
发布于 2016-07-31 12:01:15
我通常采用以下方法(未经测试,因为缺乏可重复的示例)
read_one <- function(f, ...){
w <- read.csv(f, ...)
m <- reshape2::melt(w, id = c("Time"))
m$source <- tools::file_path_sans_ext(f) # keep track of filename
m
}
plot_one <- function(d){
ggplot(d, aes(x=Time, y=value)) +
geom_line(aes(colour=variable), size = 0.8) +
ggtitle('Step Scan') +
labs(x = 'Time', y = 'Metabolite Concentration', color="Metabolites")
}
## strategy 1 (multiple independent plots)
ml <- lapply(filesToProcess, read_one)
pl <- lapply(ml, plot_one)
gridExtra::grid.arrange(grobs = pl)
## strategy 2: facetting
m <- plyr::ldply(filesToProcess, read_one)
ggplot(m, aes(x=Time, y=value)) +
facet_wrap(~source) +
geom_line(aes(colour=variable), size = 0.8) +
ggtitle('Step Scan') +
labs(x = 'Time', y = 'Metabolite Concentration', color="Metabolites")Stack Overflow用户
发布于 2016-08-01 11:05:31
我想我解决了这个问题。我承认答案有点粗糙。但是,如果我能够在for循环之外初始化"p“变量,它就解决了这个问题。
filesToProcess = readLines("FilesToProcess.txt")
#initializing the variable with ggplot() object
p <- ggplot()
for(i in 1:length(filesToProcess)){
fileName = filesToProcess[i]
fileContent = read.csv(fileName)
p <- p +
geom_line(data = fileContent, aes(x = Time, y = A, color = "A"), size =0.8) +
geom_line(data = fileContent, aes(x = Time, y = B, color = "B"), size =0.8) +
geom_line(data = fileContent, aes(x = Time, y = AE, color = "AE"), size =0.8) +
geom_line(data = fileContent, aes(x = Time, y = E, color = "E"), size =0.8)
}
p <- p + theme_bw() + scale_x_continuous(breaks=1:20) +
xlab('Time') +
ylab('Metabolite Concentration')+
ggtitle('Step Scan') +
labs(color="Legend text")
plot(p)
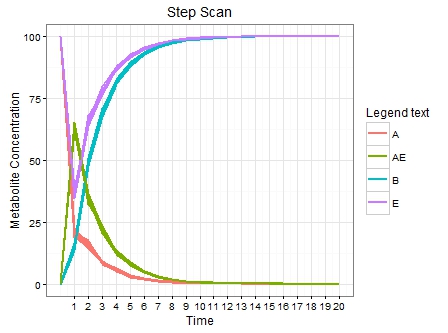
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/38683167
复制相关文章
相似问题

