捕获组可以嵌套在非捕获组中吗?
捕获组可以嵌套在非捕获组中吗?
提问于 2016-03-02 14:44:42
我试图在R中使用类PERL正则表达式分割一个FASTA头。
输入字符串的一些示例:
>P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B
>ENSEMBL:ENSBTAP00000024146 (Bos taurus) similar to alpha-2-macroglobulin isoform 1
>ENSEMBL:ENSBTAP00000024462 (Bos taurus) 47 kDa protein;>ENSEMBL:ENSBTAP00000024466 (Bos taurus) 44 kDa protein
>Q3SX28 TREMBL:Q3SX28;Q5KR48 (Bos taurus) Tropomyosin 2
>P00761 SWISS-PROT:P00761|TRYP_PIG Trypsin - Sus scrofa (Pig).我想要得到信息
- 标识符(P04259、ENSBTAP00000024146、ENSBTAP00000024462、Q3SX28、P00761)
- 如果有类似于案例1 (KRT6B)的基因符号
- 蛋白质名称(角蛋白,Ⅱ型细胞骨架6B,(牛金牛)类似于α-2-巨球蛋白异构体1,(牛金牛) 47 kDa蛋白,(牛金牛)肌球蛋白2,胰蛋白酶- scrofa (猪)
我建议的正则表达式是R格式的:
^(?:>.*?(?:ENSEMBL:|SWISS-PROT:|TREMBL:))([A-Z0-9.-]+)(?:.*?Gene_Symbol=(\\S+)|\\s+|;\\S+ |\\|\\S+)(.*?)(?:;>.*?|\\n)并以pcre格式:
^(?:>.*?(?:ENSEMBL:|SWISS-PROT:|TREMBL:))([A-Z0-9.-]+)(?:.*?Gene_Symbol=(\S+)|\s|;\S+ |\|\S+)(.*?)(?:;>.*?|\n)我在regex101.com上测试了正则表达式,结果正是我想要的。但是如果我在R中执行它,每个捕获组都包含完整的字符串。我认为并非每个条目都有基因符号的特殊情况会产生一些内部问题。也许这也是我将捕获组嵌套在非捕获组中的一个问题。
> gsub(pattern = regex, replacement = "\\1", x = ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B", perl = TRUE)
[1] ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B"
> gsub(pattern = regex, replacement = "\\2", x = ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B", perl = TRUE)
[1] ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B"
> gsub(pattern = regex, replacement = "\\3", x = ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B", perl = TRUE)
[1] ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B"regex101.com测试结果
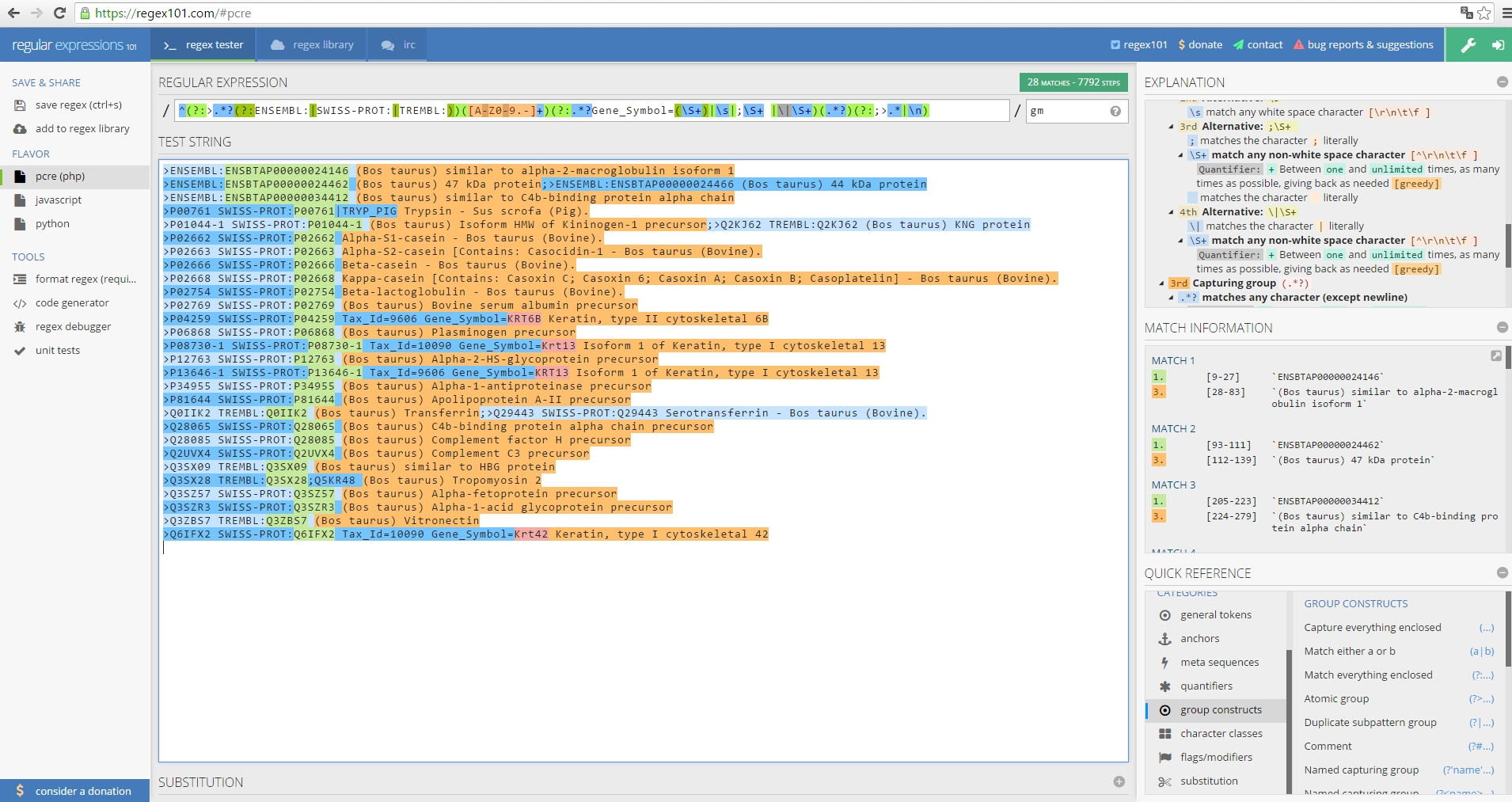
最后,我想要绿色块,红色块(如果可用的话)和橙色块作为结果。希望你能帮我。
回答 1
Stack Overflow用户
回答已采纳
发布于 2016-03-02 15:35:36
在在线regex测试器中测试字符串时,使用了多行字符串。regex中的最后一个\n匹配一个换行符,因此,您有一个>P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B匹配。
您只需要用一个字符串尾锚替换\n,就可以使它与gsub一起工作。
x = ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B"
gsub("^(?:>.*?(?:ENSEMBL:|SWISS-PROT:|TREMBL:))([A-Z0-9.-]+)(?:.*?Gene_Symbol=(\\S+)|\\s|;\\S+ |\\|\\S+)(.*?)(?:;>.*|$)", "\\1", x, perl = TRUE)
gsub("^(?:>.*?(?:ENSEMBL:|SWISS-PROT:|TREMBL:))([A-Z0-9.-]+)(?:.*?Gene_Symbol=(\\S+)|\\s|;\\S+ |\\|\\S+)(.*?)(?:;>.*|$)", "\\2", x, perl = TRUE)
gsub("^(?:>.*?(?:ENSEMBL:|SWISS-PROT:|TREMBL:))([A-Z0-9.-]+)(?:.*?Gene_Symbol=(\\S+)|\\s|;\\S+ |\\|\\S+)(.*?)(?:;>.*|$)", "\\3", x, perl = TRUE)但是,由于您想提取文本,我建议使用正则表达式从str_match包中提取文本:
> library(stringr)
> x = ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B"
> str_match(x, "^(?:>.*?(?:ENSEMBL:|SWISS-PROT:|TREMBL:))([A-Z0-9.-]+)(?:.*?Gene_Symbol=(\\S+)|\\s|;\\S+ |\\|\\S+)(.*?)(?:;>.*|$)")
[,1] [,2] [,3]
[1,] ">P04259 SWISS-PROT:P04259 Tax_Id=9606 Gene_Symbol=KRT6B Keratin, type II cytoskeletal 6B" "P04259" "KRT6B"
[,4]
[1,] " Keratin, type II cytoskeletal 6B"页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/35750411
复制相关文章
相似问题

