ggplot2:在facet中创建facet
ggplot2:在facet中创建facet
提问于 2015-11-05 08:34:19
我不知道我是否正确地表达了自己的标题,但我希望这是容易理解,当你看到以下数字。首先,这是我的数据:
hydrocarbons average SD type group
N 6,21 4,632774217 PAHs Naphtalenes
N1 4,71 2,43670665 PAHs Naphtalenes
N2 7,6 3,266286228 PAHs Naphtalenes
N3 16,18 11,00643289 PAHs Naphtalenes
N4 18,8 4,59631824 PAHs Naphtalenes
F 16,87 7,022165062 PAHs Fluorenes
F1 16,64 5,721267073 PAHs Fluorenes
F2 18,67 8,467132345 PAHs Fluorenes
F3 22,79 0,988021105 PAHs Fluorenes
P 7,97 0,211647391 PAHs Phenanthrenes
P1 26,66 16,64819987 PAHs Phenanthrenes
P2 21,72 4,416811664 PAHs Phenanthrenes
P3 18,99 4,635405486 PAHs Phenanthrenes
P4 66,28 7,706085861 PAHs Phenanthrenes
D 8,33 0,89862145 PAHs Dibenzothiophenes
D1 8,63 PAHs Dibenzothiophenes
D2 9,57 PAHs Dibenzothiophenes
D3 20,69 3,453922632 PAHs Dibenzothiophenes
D4 32,5 8,191613185 PAHs Dibenzothiophenes
FL 10,37 PAHs Fluoranthenes
PY 10,53 PAHs Fluoranthenes
FL1 24,42 8,886055918 PAHs Fluoranthenes
FL2 42,52 9,466539232 PAHs Fluoranthenes
FL3 51,99 15,77786373 PAHs Fluoranthenes
C 74,28 9,560499532 PAHs Chrysenes
C1 46,56 15,86163409 PAHs Chrysenes
C2 82,85 4,854714782 PAHs Chrysenes
C3 114,42 41,70884318 PAHs Chrysenes
nC-10 2,24 alkanes
nC-11 2,24 alkanes
nC-12 4,85 1,414267191 alkanes
nC-13 5,54 0,089306765 alkanes
nC-14 6,81 0,241222891 alkanes
nC-15 5,56 alkanes
nC-16 5,95 alkanes
nC-17 5,82 alkanes
nC-18 5,7 alkanes
nC-19 6,41 alkanes
nC-20 7,36 alkanes
nC-21 6,24 alkanes
nC-22 6,07 alkanes
nC-23 6,35 alkanes
nC-24 7,32 alkanes
nC-25 6,6 2,215395794 alkanes
nC-26 5,97 1,839829721 alkanes
nC-27 6,51 1,972060107 alkanes
nC-28 7,57 1,797509743 alkanes
nC-29 8,37 3,004883333 alkanes
nC-30 9,05 3,503601406 alkanes
nC-31 10,27 4,242811665 alkanes
nC-32 11,5 5,087821955 alkanes
nC-33 14,31 8,085948386 alkanes
nC-34 16,96 10,10105484 alkanes
nC-35 20,52 14,1878649 alkanes
nC-36 21,88 13,40071226 alkanes
n-C5 (Pentane) 10,63 1,715015757 VOCs
n-C6 (Hexane) 1,74 0,859880844 VOCs
n-C7 (Heptane) 9,62 4,316473516 VOCs
n-C9 (Nonane) 2,34 0,044641 VOCs
Benzene 23,51 0,631882255 VOCs
Toluene 18,48 2,369137637 VOCs
Ethylbenzene 7,55 7,171631537 VOCs
m-Xylene 12,53 7,250491275 VOCs
p-Xylene 15,21 1,800247445 VOCs
o-Xylene 21,96 2,184177383 VOCs
Propylbenzene 12,8 15,31136895 VOCs
n-Butylbenzene 9,33 5,486543125 VOCs
n-Pentylbenzene 6,77 0,420247353 VOCs我想绘制我的碳氢化合物的半衰期(“平均”),并按“类型”对其进行分类。然而,只有类型"PAHs“,我想有额外的方面,根据”组“。我能编写的最好的代码和图是下面的图,但这并不完全是我想要的:
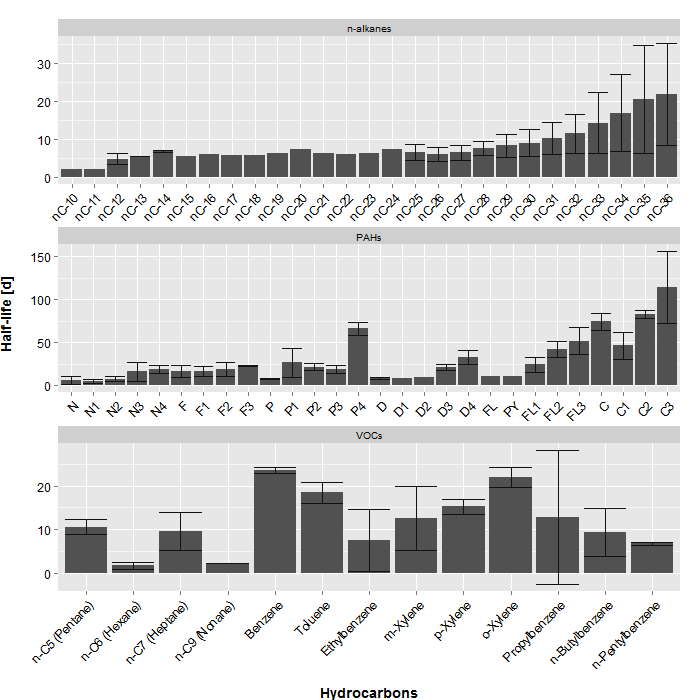
这是我使用的代码:
all <- read.delim2("E#6-results_chemistry.txt", header=TRUE)
library(ggplot2)
all$hydrocarbons <- factor(all$hydrocarbons, levels = all$hydrocarbons) #keeps the order of x-axis same as in table
levels(all$type)[levels(all$type)=="alkanes"] <- "n-alkanes" #if needed to change specific labels in a column
ggplot(all, aes(x=hydrocarbons, y=average)) +
geom_bar(position = position_dodge(), stat="identity", fill="gray32") +
geom_errorbar(aes(ymin=average-SD, ymax=average+SD), color="gray8") +
facet_wrap(~type, scales="free", ncol=1) +
ylab("Half-life [d]") + xlab("Hydrocarbons") +
theme(axis.text.x=element_text(size=12, color="black", angle=45, hjust=1),
axis.title.x = element_text(size=14, face="bold", vjust=-0.7),
axis.title.y = element_text(size=14, face="bold", vjust=2),
axis.text.y = element_text(size=12, colour="black"))现在,问题是,是否有可能制作一个图形,我可以在现有的一个,只为"PAHs“类型,根据”组“额外的小面条?图形看起来应该是这样的:
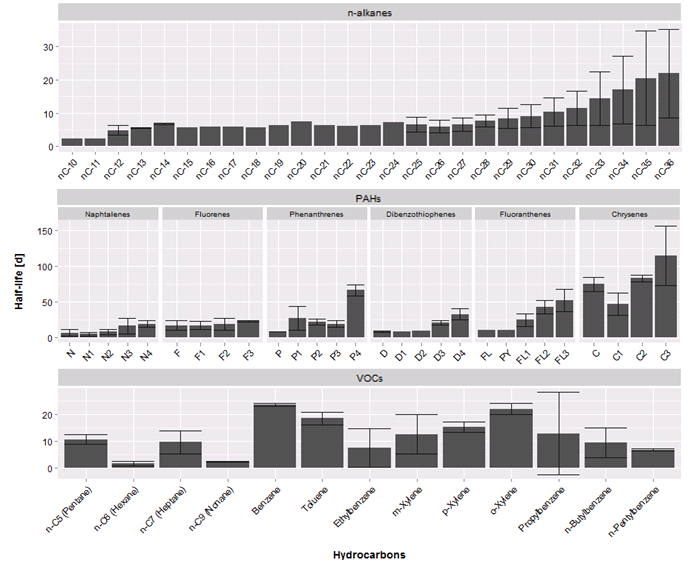
如果可能的话,它也适用于我,为"PAHs“做第二个x轴,然后按照”群组“排列它。
谢谢你,丹尼斯
回答 1
Stack Overflow用户
回答已采纳
发布于 2015-11-09 02:16:24
我想你需要深入研究ggplot的结构。这种方法采用了原来的情节,但去掉了中间面板(但保留了空间)。我将中间面板作为一个单独的图来构造,包括“PAHs”级别的各个方面。我从这个图中提取相关的材料(地块、条形、x轴和y轴),将其插入到原图中的空空间中。然后,构造一个包含变量名称“PAHs”的新条。该方法确保三个面板的高度相同,unit(1, "null")。(请参阅下面我使用的all数据框架的图表。)
次要编辑:更新到ggplot2 2.2.1
library(ggplot2)
library(gtable)
library(grid)
all$hydrocarbons <- factor(all$hydrocarbons, levels = all$hydrocarbons) #keeps the order of x-axis same as in table
levels(all$type)[levels(all$type)=="alkanes"] <- "n-alkanes" #if needed to change specific labels in a column
# Original plot
pAll <- ggplot(all, aes(x = hydrocarbons, y = average)) +
geom_bar(position = position_dodge(), stat="identity", fill = "gray32") +
geom_errorbar(aes(ymin = average-SD, ymax = average+SD), color = "gray8") +
facet_wrap( ~ type, scales = "free", ncol = 1) +
ylab("Half-life [d]") + xlab("Hydrocarbons") +
theme(axis.text.x = element_text(size = 12, color = "black", angle = 45, hjust = 1),
axis.title.x = element_text(size = 14, face = "bold", vjust = -0.7),
axis.title.y = element_text(size = 14, face = "bold", vjust = 2),
axis.text.y = element_text(size = 12, colour = "black"))
# Middle plot, but with facets for 'PAHs'
pMid <- ggplot(subset(all, type == "PAHs"), aes(x = hydrocarbons, y = average)) +
geom_bar(position = position_dodge(), stat = "identity", fill = "gray32") +
geom_errorbar(aes(ymin = average - SD, ymax = average + SD), color = "gray8") +
facet_grid(. ~ group, scales = "free", space = "free") +
ylab("Half-life [d]") +
theme(axis.text.x = element_text(size = 12, color = "black", angle = 45, hjust = 1),
axis.title.x = element_text(size = 14, face = "bold", vjust = -0.7),
axis.title.y = element_text(size = 14, face = "bold", vjust = 2),
axis.text.y = element_text(size = 12, colour = "black"))
# Get the ggplot grobs
gAll <- ggplotGrob(pAll)
gMid <- ggplotGrob(pMid)
# In gMid, get the positions of the panels in the layout: t = top, l = left, ...
pos1 <- c(subset(gMid$layout, grepl("panel", gMid$layout$name), select = t:r))
# Extract 'panels' (along with x-axis and strips)
panels <- gMid[(unique(pos1$t) - 1) : (unique(pos1$t) + 1), min(pos1$l) : max(pos1$l)]
# Extract 'axisL' - y-axis
axisL <- gMid[unique(pos1$t), min(pos1$l) - 1]
# In gAll, get the positions of the panel of the middle plot in the layout
pos2 <- c(subset(gAll$layout, grepl("panel", gAll$layout$name), select = t:r))
pos2 <- lapply(pos2, "[", 2)
# Drop original panel material in the middle plot
drop <- gAll$layout$name %in% c("panel-1-2", "strip-t-1-2", "axis-b-1-2", "axis-l-2-1")
gAll$grobs[drop] <- NULL
gAll$layout <- gAll$layout[!drop,]
# Add new 'panels' to gAll
gAll <- gtable_add_grob(gAll, panels, t = pos2$t-1, l = pos2$l, b = pos2$t+1, name = "panels")
# Add new 'axisL' to gAll
gAll <- gtable_add_grob(gAll, axisL, t = pos2$t, l = pos2$l-1, name = "strips")
# Add row above the current strips
gAll <- gtable_add_rows(gAll, gAll$heights[pos2$t-1], pos2$t-2)
# Add grob, a new strip containing variable name 'PAHs', into the new row
gAll <- gtable_add_grob(gAll,
list(rectGrob(gp = gpar(col = NA, fill = "gray85")),
textGrob("PAHs", gp = gpar(fontsize = 9.6))),
t = pos2$t-1, l = pos2$l, name = c("background", "text"))
# Add a small gap between the strips
gAll <- gtable_add_rows(gAll, unit(1/10, "line"), pos2$t-1)
# Probably not needed, but if axisB font size in pMid is different from pAll
gAll$heights[pos2$t + 3] <- gMid$heights[unique(pos1$t) + 1]
# Draw it
grid.newpage()
grid.draw(gAll)
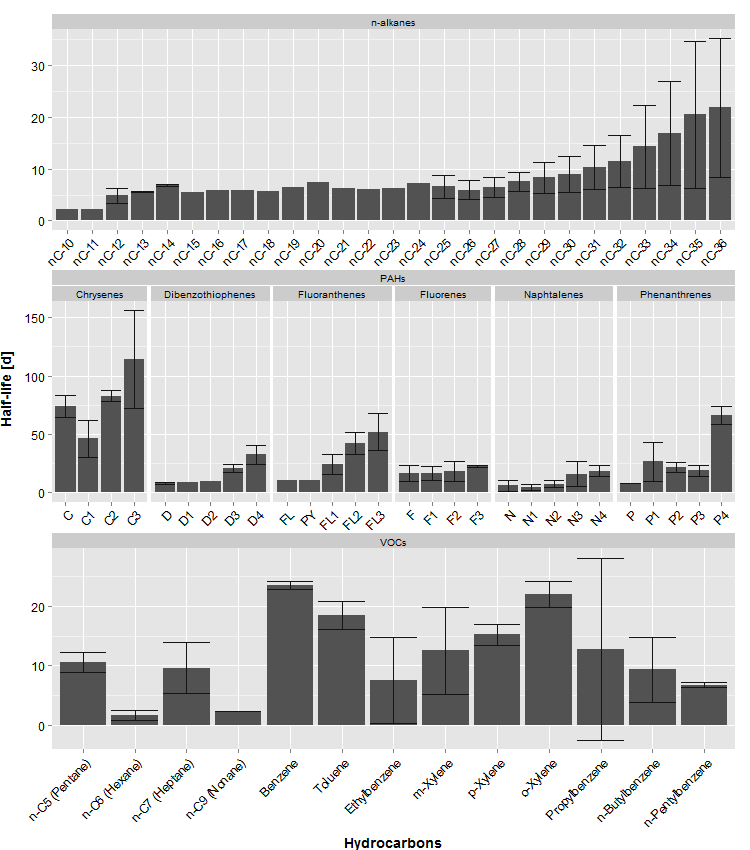
all = structure(list(hydrocarbons = structure(c(21L, 28L, 29L, 30L,
31L, 12L, 13L, 14L, 15L, 60L, 62L, 63L, 64L, 65L, 6L, 7L, 8L,
9L, 10L, 16L, 67L, 17L, 18L, 19L, 2L, 3L, 4L, 5L, 32L, 33L, 34L,
35L, 36L, 37L, 38L, 39L, 40L, 41L, 42L, 43L, 44L, 45L, 46L, 47L,
48L, 49L, 50L, 51L, 52L, 53L, 54L, 55L, 56L, 57L, 58L, 23L, 24L,
25L, 26L, 1L, 68L, 11L, 20L, 61L, 59L, 66L, 22L, 27L), .Label = c("Benzene",
"C", "C1", "C2", "C3", "D", "D1", "D2", "D3", "D4", "Ethylbenzene",
"F", "F1", "F2", "F3", "FL", "FL1", "FL2", "FL3", "m-Xylene",
"N", "n-Butylbenzene", "n-C5 (Pentane)", "n-C6 (Hexane)", "n-C7 (Heptane)",
"n-C9 (Nonane)", "n-Pentylbenzene", "N1", "N2", "N3", "N4", "nC-10",
"nC-11", "nC-12", "nC-13", "nC-14", "nC-15", "nC-16", "nC-17",
"nC-18", "nC-19", "nC-20", "nC-21", "nC-22", "nC-23", "nC-24",
"nC-25", "nC-26", "nC-27", "nC-28", "nC-29", "nC-30", "nC-31",
"nC-32", "nC-33", "nC-34", "nC-35", "nC-36", "o-Xylene", "P",
"p-Xylene", "P1", "P2", "P3", "P4", "Propylbenzene", "PY", "Toluene"
), class = "factor"), average = c(6.21, 4.71, 7.6, 16.18, 18.8,
16.87, 16.64, 18.67, 22.79, 7.97, 26.66, 21.72, 18.99, 66.28,
8.33, 8.63, 9.57, 20.69, 32.5, 10.37, 10.53, 24.42, 42.52, 51.99,
74.28, 46.56, 82.85, 114.42, 2.24, 2.24, 4.85, 5.54, 6.81, 5.56,
5.95, 5.82, 5.7, 6.41, 7.36, 6.24, 6.07, 6.35, 7.32, 6.6, 5.97,
6.51, 7.57, 8.37, 9.05, 10.27, 11.5, 14.31, 16.96, 20.52, 21.88,
10.63, 1.74, 9.62, 2.34, 23.51, 18.48, 7.55, 12.53, 15.21, 21.96,
12.8, 9.33, 6.77), SD = c(4.632774217, 2.43670665, 3.266286228,
11.00643289, 4.59631824, 7.022165062, 5.721267073, 8.467132345,
0.988021105, 0.211647391, 16.64819987, 4.416811664, 4.635405486,
7.706085861, 0.89862145, NA, NA, 3.453922632, 8.191613185, NA,
NA, 8.886055918, 9.466539232, 15.77786373, 9.560499532, 15.86163409,
4.854714782, 41.70884318, NA, NA, 1.414267191, 0.089306765, 0.241222891,
NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, 2.215395794, 1.839829721,
1.972060107, 1.797509743, 3.004883333, 3.503601406, 4.242811665,
5.087821955, 8.085948386, 10.10105484, 14.1878649, 13.40071226,
1.715015757, 0.859880844, 4.316473516, 0.044641, 0.631882255,
2.369137637, 7.171631537, 7.250491275, 1.800247445, 2.184177383,
15.31136895, 5.486543125, 0.420247353), type = structure(c(2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L,
3L, 3L, 3L), .Label = c("alkanes", "PAHs", "VOCs"), class = "factor"),
group = structure(c(6L, 6L, 6L, 6L, 6L, 5L, 5L, 5L, 5L, 7L,
7L, 7L, 7L, 7L, 3L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L, 4L, 2L,
2L, 2L, 2L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L), .Label = c("",
"Chrysenes", "Dibenzothiophenes", "Fluoranthenes", "Fluorenes",
"Naphtalenes", "Phenanthrenes"), class = "factor")), .Names = c("hydrocarbons",
"average", "SD", "type", "group"), class = "data.frame", row.names = c(NA,
-68L))页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/33539824
复制相关文章
相似问题

