图分析:识别循环路径
图分析:识别循环路径
提问于 2015-06-24 18:55:46
我有一个无向图,如:
import igraph as ig
G = ig.Graph()
G.add_vertices(9)
G.add_edges([(0,1), (1,2),(2,3),(3,0),(0,4),(4,5),(5,6),(6,7),(6,8)])从节点0到1,2,3返回到0有一个“循环路径”(抱歉,图中的节点标记以1而不是0开头)。
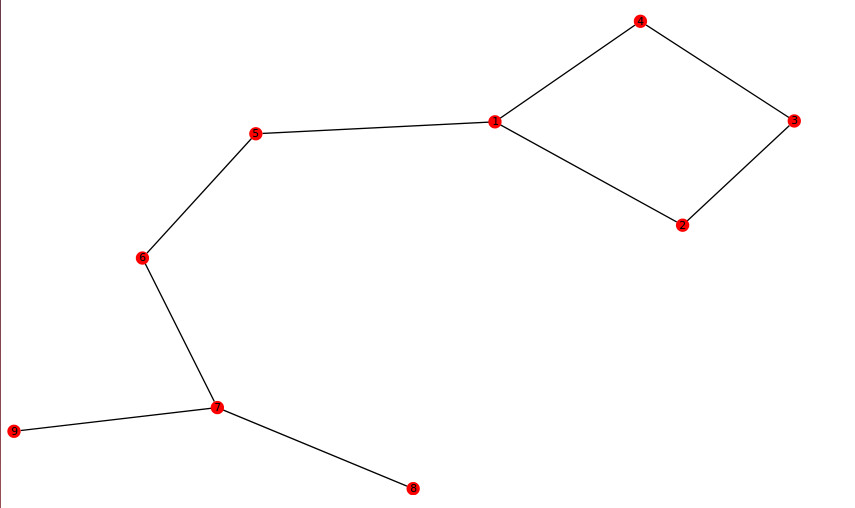
对于赋值,我需要标识“循环路径”连接到图的其余部分的节点,即0,最重要的是,“循环路径”本身,即[0,1,2,3,0]和/或[0,3,2,1,0]。
我正在努力,但真的毫无头绪。如果我像这里一样使用来自find_all_paths(G,0,0)的函数,我当然只能得到[0]
回答 4
Stack Overflow用户
回答已采纳
发布于 2015-06-26 09:18:31
好的,这是我自己问题的答案之一:
多亏了马克斯李和深烯‘的帮助,我有了在python_igraph中重写基函数以便工作的想法:
import igraph as ig
G = ig.Graph()
G.add_vertices(9)
G.add_edges([(0,1), (1,2),(2,3),(3,0),(0,4),(4,5),(5,6),(6,7),(6,8)])
def cycle_basis_ig(G,root=None):
gnodes=set(n.index for n in G.vs())
cycles=[]
while gnodes: # loop over connected components
if root is None:
root=gnodes.pop()
stack=[root]
pred={root:root}
used={root:set()}
while stack: # walk the spanning tree finding cycles
z=stack.pop() # use last-in so cycles easier to find
zused=used[z]
for nbr in G.neighbors(z,mode='ALL'):
if nbr not in used: # new node
pred[nbr]=z
stack.append(nbr)
used[nbr]=set([z])
elif nbr is z: # self loops
cycles.append([z])
elif nbr not in zused:# found a cycle
pn=used[nbr]
cycle=[nbr,z]
p=pred[z]
while p not in pn:
cycle.append(p)
p=pred[p]
cycle.append(p)
cycles.append(cycle)
used[nbr].add(z)
gnodes-=set(pred)
root=None
return cycles
cb = cycle_basis_ig(G)
print 'cycle_basis_ig: ', cbStack Overflow用户
发布于 2015-06-25 21:19:35
因为这个问题也是用networkx标记的,所以我用它来举例说明代码。
在图论中,“循环路径”通常称为循环。
我看到的最简单(可能不是最快的)想法是找到循环和交点集(或切割顶点,即增加连通部件数量的点),然后它们的交集就是解决方案。
在同样的基础上开始:
import networkx as nx
G.add_nodes_from([9])
G.add_edges_from([(0,1), (1,2),(2,3),(3,0),(0,4),(4,5),(5,6),(6,7),(6,8)])现在问题的解决办法是:
cycles = nx.cycle_basis(G) # list of cycles
cuts = list(nx.articulation_points(G)) # list of cut verteces
nodes_needed = set() # the set of nodes we are looking for
for cycle in cycles:
for node in cycle:
if node in cuts:
nodes_needed.add(node)Stack Overflow用户
发布于 2015-06-25 22:36:47
下面是一个用宽度优先搜索寻找循环的例子。我想知道是否有更有效的方法。即使是中等大的图形,或者更长的最大循环长度,这也可能会持续很长时间。深度优先搜索也可以这样做。首先,我相信您使用R发布了这个问题,所以下面也可以找到R版本。由于同样的原因,python版本并不完全是pythonic的,因为我很快就从R翻译过来了。有关解释,请参见代码中的注释。
import igraph
# creating a toy graph
g = igraph.Graph.Erdos_Renyi(n = 100, p = 0.04)
# breadth first search of paths and unique loops
def get_loops(adj, paths, maxlen):
# tracking the actual path length:
maxlen -= 1
nxt_paths = []
# iterating over all paths:
for path in paths['paths']:
# iterating neighbors of the last vertex in the path:
for nxt in adj[path[-1]]:
# attaching the next vertex to the path:
nxt_path = path + [nxt]
if path[0] == nxt and min(path) == nxt:
# the next vertex is the starting vertex, we found a loop
# we keep the loop only if the starting vertex has the
# lowest vertex id, to avoid having the same loops
# more than once
paths['loops'].append(nxt_path)
# if you don't need the starting vertex
# included at the end:
# paths$loops <- c(paths$loops, list(path))
elif nxt not in path:
# keep the path only if we don't create
# an internal loop in the path
nxt_paths.append(nxt_path)
# paths grown by one step:
paths['paths'] = nxt_paths
if maxlen == 0:
# the final return when maximum search length reached
return paths
else:
# recursive return, to grow paths further
return get_loops(adj, paths, maxlen)
adj = []
loops = []
# the maximum length to limit computation time on large graphs
# maximum could be vcount(graph), but that might take for ages
maxlen = 4
# creating an adjacency list
# for directed graphs use the 'mode' argument of neighbors()
# according to your needs ('in', 'out' or 'all')
adj = [[n.index for n in v.neighbors()] for v in g.vs]
# recursive search of loops
# for each vertex as candidate starting point
for start in xrange(g.vcount()):
loops += get_loops(adj,
{'paths': [[start]], 'loops': []}, maxlen)['loops']在R中相同
require(igraph)
# creating a toy graph
g <- erdos.renyi.game(n = 100, p.or.m = 0.04)
# breadth first search of paths and unique loops
get_loops <- function(adj, paths, maxlen){
# tracking the actual path length:
maxlen <- maxlen - 1
nxt_paths <- list()
# iterating over all paths:
for(path in paths$paths){
# iterating neighbors of the last vertex in the path:
for(nxt in adj[[path[length(path)]]]){
# attaching the next vertex to the path:
nxt_path <- c(path, nxt)
if(path[1] == nxt & min(path) == nxt){
# the next vertex is the starting vertex, we found a loop
# we keep the loop only if the starting vertex has the
# lowest vertex id, to avoid having the same loops
# more than once
paths$loops <- c(paths$loops, list(nxt_path))
# if you don't need the starting vertex included
# at the end:
# paths$loops <- c(paths$loops, list(path))
}else if(!(nxt %in% path)){
# keep the path only if we don't create
# an internal loop in the path
nxt_paths <- c(nxt_paths, list(nxt_path))
}
}
}
# paths grown by one step:
paths$paths <- nxt_paths
if(maxlen == 0){
# the final return when maximum search length reached
return(paths)
}else{
# recursive return, to grow paths further
return(get_loops(adj, paths, maxlen))
}
}
adj <- list()
loops <- list()
# the maximum length to limit computation time on large graphs
# maximum could be vcount(graph), but that might take for ages
maxlen <- 4
# creating an adjacency list
for(v in V(g)){
# for directed graphs use the 'mode' argument of neighbors()
# according to your needs ('in', 'out' or 'all')
adj[[as.numeric(v)]] <- neighbors(g, v)
}
# recursive search of loops
# for each vertex as candidate starting point
for(start in seq(length(adj))){
loops <- c(loops, get_loops(adj, list(paths = list(c(start)),
loops = list()), maxlen)$loops)
}页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/31034730
复制相关文章
相似问题

