R-将特定列绘制为x,并以两行作为行。
R-将特定列绘制为x,并以两行作为行。
提问于 2015-06-02 17:50:17
下面是我的数据的一个小样本:
gene_name ctrl_lsm1_ratio_t0 ctrl_lsm1_ratio_t1 ctrl_lsm1_ratio_t2
22 ABP140 -0.262682 -0.303352 -0.223626
246 ARI1 -0.163952 -0.374765 -0.321876
454 BPH1 -0.517519 -0.524553 -0.747609
513 BUR6 0.645573 0.217433 0.390403
588 CDC20 -0.264072 -0.665268 -0.594191
ctrl_lsm1_ratio_t3 ctrl_lsm1_stat_t0 ctrl_lsm1_stat_t1 ctrl_lsm1_stat_t2
22 -0.421704 no no no
246 -0.692391 no no no
454 -0.793595 no no yes
513 0.200799 yes no no
588 -0.523884 no yes yes
ctrl_lsm1_stat_t3 systematic_name
22 yes YOR239W
246 yes YGL157W
454 yes YCR032W
513 no YER159C
588 yes YGL116W我想在x轴(如时间点0、1、2和3)上绘制2:5的列,y轴与比率列相匹配。
如果有一种方法在特定的时间点将点涂成“是”或“否”的颜色,我也希望能够这样做。(例如,ctrl_lsm1_ratio_t0列中的点将根据ctrl_lsm1_stat_t0列中的值着色)。
我还希望一次只绘制两行,这两行都是行(例如,第22行和第513行)。希望这有意义!我对R不熟悉,不知道该怎么做。我愿意下载任何必要的软件包。
回答 1
Stack Overflow用户
回答已采纳
发布于 2015-06-02 19:51:53
data.csv:
gene_name,ctrl_lsm1_ratio_t0,ctrl_lsm1_ratio_t1,ctrl_lsm1_ratio_t2,ctrl_lsm1_ratio_t3,ctrl_lsm1_stat_t0,ctrl_lsm1_stat_t1,ctrl_lsm1_stat_t2,ctrl_lsm1_stat_t3,systematic_name
ABP140,-0.262682,-0.303352,-0.22362,-0.421704,no,no,no,yes,YOR239W6
ARI1,-0.163952,-0.374765,-0.32187,-0.692391,no,no,no,yes,YGL157W6
BPH1,-0.517519,-0.524553,-0.74760,-0.793595,no,no,yes,yes,YCR032W9
BUR6,0.645573,0.217433,0.39040,0.200799,yes,no,no,no,YER159C3
CDC20,-0.264072,-0.665268,-0.59419,-0.523884,no,yes,yes,yes,YGL116W1代码:
d<-read.csv("data.csv", header=T, stringsAsFactors=F)
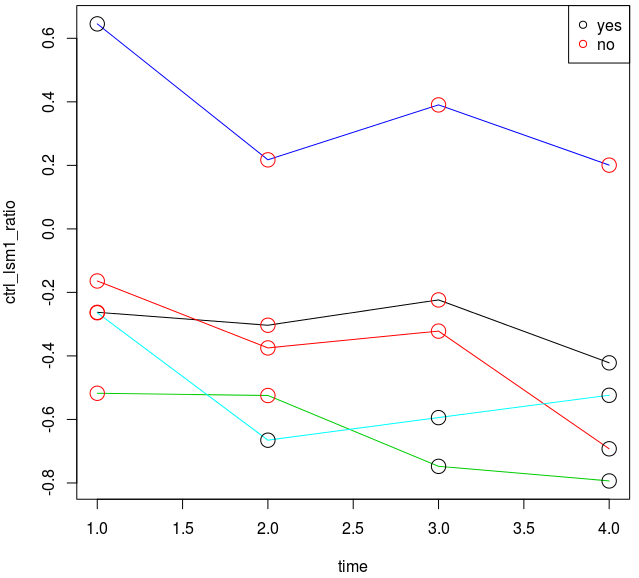
matplot(t(d[,2:5]), type="l", pch=20, lty=1, xlab="time", ylab="ctrl_lsm1_ratio")
d2<-reshape(d[,6:9],varying=list(names(d[,6:9])),direction="long",v.name="ctrl_lsm1_stat", ids=d$gene_name)
points(d2$time, unlist(d[,2:5]), col=ifelse(d2$ctrl_lsm1_stat=="yes",1,2),cex=2.0)
legend("topright",legend=c("yes","no"), col=c(1,2), pch=21)
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/30603114
复制相关文章
相似问题

