使用` `party`‘包运行` `ctree`’,列作为因子,而不是字符
使用` `party`‘包运行` `ctree`’,列作为因子,而不是字符
提问于 2014-03-01 21:17:28
我已经提到了convert data.frame column format from character to factor,Converting multiple data.table columns to factors in R和Convert column classes in data.table
不幸的是,它并没有解决我的问题。我正在处理体脂数据集,我的数据被称为> bf。我增加了一个名为agegrp的专栏,将不同年龄的人归类为年轻人、中年人或老年人:
bf$agegrp<-ifelse(bf$age<=40, "young", ifelse(bf$age>40 & bf$age<55,"middle", "old"))这是ctree分析:
> set.seed(1234)
> modelsample<-sample(2, nrow(bf), replace=TRUE, prob=c(0.7, 0.3))
> traindata<-bf[modelsample==1, ]
> testdata<-bf[modelsample==2, ]
> predictor<-agegrp~DEXfat+waistcirc+hipcirc+kneebreadth` and ran, `bf_ctree<-ctree(predictor, data=traindata)
> bf_ctree<-ctree(predictor, data=traindata)我得到了以下错误:
Error in trafo(data = data, numeric_trafo = numeric_trafo, factor_trafo = factor_trafo, :
data class character is not supported
In addition: Warning message:
In storage.mode(RET@predict_trafo) <- "double" : NAs introduced by coercion既然bf$agegrp是我跑的“角色”,
> bf$agegrp<-as.factor(bf$agegrp)现在强制使用agegrp列进行因子分析。
> Class (bf$agegrp)给了[1] "Factor"。
我再次尝试运行ctree,但是它抛出了相同的错误。有人知道问题的根源是什么吗?
回答 1
Stack Overflow用户
回答已采纳
发布于 2014-03-01 22:27:51
这对我来说很管用:
library(mboot)
library(party)
bf <- bodyfat
bf$agegrp <- cut(bf$age,c(0,40,55,100),labels=c("young","middle","old"))
predictor <- agegrp~DEXfat+waistcirc+hipcirc+kneebreadth
set.seed(1234)
modelsample <-sample(2, nrow(bf), replace=TRUE, prob=c(0.7, 0.3))
traindata <-bf[modelsample==1, ]
testdata <-bf[modelsample==2, ]
bf_ctree <-ctree(predictor, data=traindata)
plot(bf_ctree)
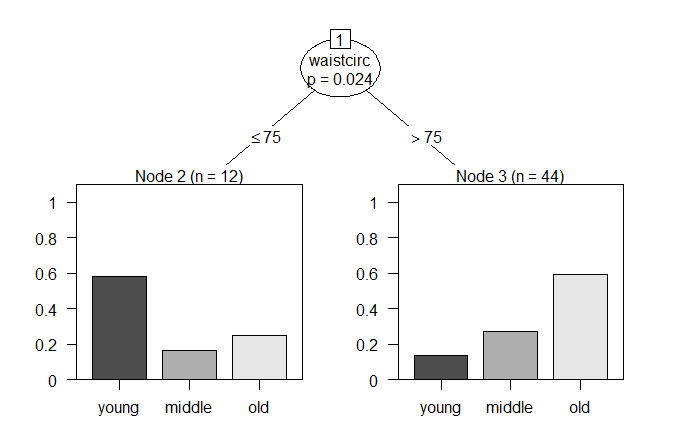
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/22120498
复制相关文章
相似问题

