在分面ggplot2图中独立地定位两个传说
我有一个由ggplot2生成的情节,其中包含两个传说。传说的位置并不理想,所以我想调整它们。我一直试图模仿the answer to "How do I position two legends independently in ggplot"中显示的方法。答案中所示的例子起作用了。然而,我无法得到所描述的方法来适应我的情况。
我使用R2.15.3 (2013-03-01),ggplot2_0.9.3.1,点阵0.20-13,gtable_0.1.2,gridExtra_0.9.1对Debian挤压.
考虑一下minimal.R生成的图。这和我的实际情节很相似。
########################
minimal.R
########################
get_stat <- function()
{
n = 20
q1 = qnorm(seq(3, 17)/20, 14, 5)
q2 = qnorm(seq(1, 19)/20, 65, 10)
Stat = data.frame(value = c(q1, q2),
pvalue = c(dnorm(q1, 14, 5)/max(dnorm(q1, 14, 5)), d = dnorm(q2, 65, 10)/max(dnorm(q2, 65, 10))),
variable = c(rep('true', length(q1)), rep('data', length(q2))))
return(Stat)
}
stat_all<- function()
{
library(ggplot2)
library(gridExtra)
stathuman = get_stat()
stathuman$dataset = "human"
statmouse = get_stat()
statmouse$dataset = "mouse"
stat = merge(stathuman, statmouse, all=TRUE)
return(stat)
}
simplot <- function()
{
Stat = stat_all()
Pvalue = subset(Stat, variable=="true")
pdf(file = "CDF.pdf", width = 5.5, height = 2.7)
stat = ggplot() + stat_ecdf(data=Stat, n=1000, aes(x=value, colour = variable)) +
theme(legend.key = element_blank(), legend.background = element_blank(), legend.position=c(.9, .25), legend.title = element_text(face = "bold")) +
scale_x_continuous("Negative log likelihood") +
scale_y_continuous("Proportion $<$ x") +
facet_grid(~ dataset, scales='free') +
scale_colour_manual(values = c("blue", "red"), name="Data type",
labels=c("Gene segments", "Model"), guide=guide_legend(override.aes = list(size = 2))) +
geom_area(data=Pvalue, aes(x=value, y=pvalue, fill=variable), position="identity", alpha=0.5) +
scale_fill_manual(values = c("gray"), name="Pvalue", labels=c(""))
print(stat)
dev.off()
}
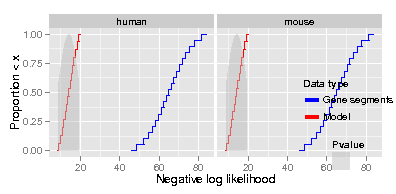
simplot()这将导致下面的情节。可以看出,Data type和Pvalue传说的位置并不好。我将此代码修改为minimal2.R。
使用版本1,它应该将图例放在顶部,代码运行时没有错误,但没有显示图例。
编辑:有两个框显示,一个在另一个之上。上面那个是空白的。如果我不像@巴普蒂斯特所建议的那样在grid.arrange()中设定高度,那么传说和情节都会放在底部的盒子里。如果我设置的高度如图所示,那我就看不到传说了。
EDIT2:看起来额外的空白框是由grid.newpage调用的,我是从前面的问题中复制的。我不知道它为什么会在那里。如果我不使用那一行,那么我只得到一个框/页。
在版本2中,我得到了这个错误。
Error in UseMethod("grid.draw") :
no applicable method for 'grid.draw' applied to an object of class "c('gg', 'ggplot')"
Calls: simplot -> grid.draw编辑:如果我按照@baptiste的建议使用print(plotNew),那么我将得到以下错误
Error in if (empty(data)) { : missing value where TRUE/FALSE needed
Calls: simplot ... facet_map_layout -> facet_map_layout.grid -> locate_grid.我试图弄清楚这是怎么回事,但我找不到多少相关的信息。
备注:
- 我不知道为什么我要得到经验民防的楼梯效应。我肯定有一个很明显的解释。如果你知道的话请告诉我。
- 如果有人能提出替代方案,例如matplotlib,我从未认真尝试过,我愿意考虑使用这种代码的替代品,甚至ggplot2来生成这个图。
- 添加
打印(ggplot_gtable(ggplot_build(Stat2)
minimal2.R给我的 TableGrob (7x7)“布局”:12个grobs z单元格名为grob 1 0 (1-7,1-7)背景矩形.背景.背景.矩形.186 2 1 (3-3,4-4)条-顶层绝对grobs.绝对值135 3 (3-3,6-6)条.顶绝对grobs.AbteGrob.141 4 5 (4-4,3-3)轴-l绝对GrobGRID.AbteGrob.129 53 (4-4,4-4)面板gTreeGRID.gTree.155 6 4 (4-4,6-6)面板gTreeGRID.gTree.169 7 6 (5-5,4-4)轴-b绝对Grob.117 8 (5-5,6-6)轴-b-绝对GrobID.AbteGrob.123 9 8(6-6-6) xlab出租车。Title.x.text.171 10 9(4-4,2-2) Ylabtextaxis.Title.y.text.173 11 10 (4-4,4-6)指南-方框gtableguide-方框12 11 (2-2,4-6)标题textplot.title.text.184 我不明白这是什么故障。有人能解释吗?guide-box是否与传说相对应,人们如何知道这一点?
下面是我的代码minimal2.R的修改版本。
########################
minimal2.R
########################
get_stat <- function()
{
n = 20
q1 = qnorm(seq(3, 17)/20, 14, 5)
q2 = qnorm(seq(1, 19)/20, 65, 10)
Stat = data.frame(value = c(q1, q2),
pvalue = c(dnorm(q1, 14, 5)/max(dnorm(q1, 14, 5)), d = dnorm(q2, 65, 10)/max(dnorm(q2, 65, 10))),
variable = c(rep('true', length(q1)), rep('data', length(q2))))
return(Stat)
}
stat_all<- function()
{
library(ggplot2)
library(gridExtra)
library(gtable)
stathuman = get_stat()
stathuman$dataset = "human"
statmouse = get_stat()
statmouse$dataset = "mouse"
stat = merge(stathuman, statmouse, all=TRUE)
return(stat)
}
simplot <- function()
{
Stat = stat_all()
Pvalue = subset(Stat, variable=="true")
pdf(file = "CDF.pdf", width = 5.5, height = 2.7)
## only include data type legend
stat1 = ggplot() + stat_ecdf(data=Stat, n=1000, aes(x=value, colour = variable)) +
theme(legend.key = element_blank(), legend.background = element_blank(), legend.position=c(.9, .25), legend.title = element_text(face = "bold")) +
scale_x_continuous("Negative log likelihood") +
scale_y_continuous("Proportion $<$ x") +
facet_grid(~ dataset, scales='free') +
scale_colour_manual(values = c("blue", "red"), name="Data type", labels=c("Gene segments", "Model"), guide=guide_legend(override.aes = list(size = 2))) +
geom_area(data=Pvalue, aes(x=value, y=pvalue, fill=variable), position="identity", alpha=0.5) +
scale_fill_manual(values = c("gray"), name="Pvalue", labels=c(""), guide=FALSE)
## Extract data type legend
dataleg <- gtable_filter(ggplot_gtable(ggplot_build(stat1)), "guide-box")
## only include pvalue legend
stat2 = ggplot() + stat_ecdf(data=Stat, n=1000, aes(x=value, colour = variable)) +
theme(legend.key = element_blank(), legend.background = element_blank(), legend.position=c(.9, .25), legend.title = element_text(face = "bold")) +
scale_x_continuous("Negative log likelihood") +
scale_y_continuous("Proportion $<$ x") +
facet_grid(~ dataset, scales='free') +
scale_colour_manual(values = c("blue", "red"), name="Data type", labels=c("Gene segments", "Model"), guide=FALSE) +
geom_area(data=Pvalue, aes(x=value, y=pvalue, fill=variable), position="identity", alpha=0.5) +
scale_fill_manual(values = c("gray"), name="Pvalue", labels=c(""))
## Extract pvalue legend
pvalleg <- gtable_filter(ggplot_gtable(ggplot_build(stat2)), "guide-box")
## no legends
stat = ggplot() + stat_ecdf(data=Stat, n=1000, aes(x=value, colour = variable)) +
theme(legend.key = element_blank(), legend.background = element_blank(), legend.position=c(.9, .25), legend.title = element_text(face = "bold")) +
scale_x_continuous("Negative log likelihood") +
scale_y_continuous("Proportion $<$ x") +
facet_grid(~ dataset, scales='free') +
scale_colour_manual(values = c("blue", "red"), name="Data type", labels=c("Gene segments", "Model"), guide=FALSE) +
geom_area(data=Pvalue, aes(x=value, y=pvalue, fill=variable), position="identity", alpha=0.5) +
scale_fill_manual(values = c("gray"), name="Pvalue", labels=c(""), guide=FALSE)
## Add data type legend: version 1 (data type legend should be on top)
## plotNew <- arrangeGrob(dataleg, stat, heights = unit.c(dataleg$height, unit(1, "npc") - dataleg$height), ncol = 1)
## Add data type legend: version 2 (data type legend should be somewhere in the interior)
## plotNew <- stat + annotation_custom(grob = dataleg, xmin = 7, xmax = 10, ymin = 0, ymax = 4)
grid.newpage()
grid.draw(plotNew)
dev.off()
}
simplot()回答 1
Stack Overflow用户
发布于 2013-07-04 12:27:46
这可以用grid.arrange和arrangeGrob来完成,但是正确地调整高度和宽度是很痛苦的。
grid.arrange(arrangeGrob(dataleg, pvalleg, nrow=1, ncol=2, widths=c(unit(1, "npc"), unit(5, "cm"))), stat, nrow=2, heights=c(unit(.2, "npc"), unit(.8, "npc")))我通常更喜欢用一个合适的图例来制作一个新的情节,并使用这个新的传说:
h <- ggplot(data.frame(a=rnorm(10), b=rnorm(10), c=factor(rbinom(10, 1,.5), labels=c("Gene segments", "Model")), d=factor("")),
aes(x=a, y=b)) +
geom_line(aes(color=c), size=1.3) + geom_polygon(aes(fill=d)) +
scale_color_manual(values=c("blue", "red"), name="Data type") +
scale_fill_manual(values="gray", name="P-value")
g_legend<-function(a.gplot){
tmp <- ggplot_gtable(ggplot_build(a.gplot))
leg <- which(sapply(tmp$grobs, function(x) x$name) == "guide-box")
legend <- tmp$grobs[[leg]]
return(legend)
}
legend <- g_legend(h)
grid.arrange(stat, legend, nrow=1, ncol=2, widths=c(unit(.8, "npc"), unit(.2, "npc")))
grid.arrange(legend, stat, nrow=2, ncol=1, heights=c(unit(.2, "npc"), unit(.8, "npc")))https://stackoverflow.com/questions/16501999
复制相似问题

