Emgu CV -各向异性扩散
Emgu CV -各向异性扩散
提问于 2013-02-05 19:54:41
有谁能给我介绍一些现有的各向异性扩散的实现,最好是perona-malik扩散?
回答 2
Stack Overflow用户
回答已采纳
发布于 2013-02-05 21:28:44
翻译以下MATLAB代码:
% pm2.m - Anisotropic Diffusion routines
function ZN = pm2(ZN,K,iterate);
[m,n] = size(ZN);
% lambda = 0.250;
lambda = .025;
%K=16;
rowC = [1:m]; rowN = [1 1:m-1]; rowS = [2:m m];
colC = [1:n]; colE = [2:n n]; colW = [1 1:n-1];
result_save=0;
for i = 1:iterate,
%i;
% result=PSNR(Z,ZN);
% if result>result_save
% result_save=result;
% else
% break;
% end
deltaN = ZN(rowN,colC) - ZN(rowC,colC);
deltaS = ZN(rowS,colC) - ZN(rowC,colC);
deltaE = ZN(rowC,colE) - ZN(rowC,colC);
deltaW = ZN(rowC,colW) - ZN(rowC,colC);
% deltaN = deltaN .*abs(deltaN<K);
% deltaS = deltaS .*abs(deltaS<K);
% deltaE = deltaE .*abs(deltaE<K);
% deltaW = deltaW .*abs(deltaW<K);
fluxN = deltaN .* exp(-((abs(deltaN) ./ K).^2) );
fluxS = deltaS .* exp(-((abs(deltaS) ./ K).^2) );
fluxE = deltaE .* exp(-((abs(deltaE) ./ K).^2) );
fluxW = deltaW .* exp(-((abs(deltaW) ./ K).^2) );
ZN = ZN + lambda*(fluxN +fluxS + fluxE + fluxW);
%ZN=max(0,ZN);ZN=min(255,ZN);
end代码不是我的,而是从:http://www.csee.wvu.edu/~xinl/code/pm2.m中提取的。
Stack Overflow用户
发布于 2020-08-15 01:40:39
OpenCV Implementation (它需要3通道映像):
from cv2.ximgproc import anisotropicDiffusion
ultrasound_ad_cv2 = anisotropicDiffusion(im,0.075 ,80, 100)并列比较
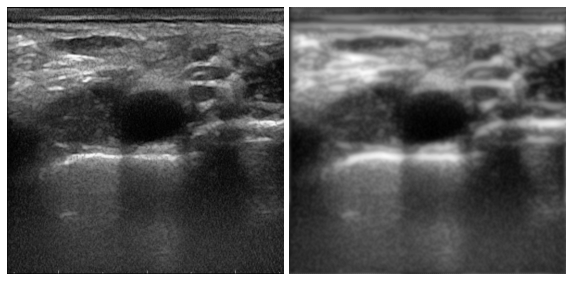
:(仅适用于灰度图像)
import scipy.ndimage.filters as flt
import numpy as np
import warnings
def anisodiff(img,niter=1,kappa=50,gamma=0.1,step=(1.,1.),sigma=0, option=1,ploton=False):
"""
Anisotropic diffusion.
Usage:
imgout = anisodiff(im, niter, kappa, gamma, option)
Arguments:
img - input image
niter - number of iterations
kappa - conduction coefficient 20-100 ?
gamma - max value of .25 for stability
step - tuple, the distance between adjacent pixels in (y,x)
option - 1 Perona Malik diffusion equation No 1
2 Perona Malik diffusion equation No 2
ploton - if True, the image will be plotted on every iteration
Returns:
imgout - diffused image.
kappa controls conduction as a function of gradient. If kappa is low
small intensity gradients are able to block conduction and hence diffusion
across step edges. A large value reduces the influence of intensity
gradients on conduction.
gamma controls speed of diffusion (you usually want it at a maximum of
0.25)
step is used to scale the gradients in case the spacing between adjacent
pixels differs in the x and y axes
Diffusion equation 1 favours high contrast edges over low contrast ones.
Diffusion equation 2 favours wide regions over smaller ones.
"""
# ...you could always diffuse each color channel independently if you
# really want
if img.ndim == 3:
warnings.warn("Only grayscale images allowed, converting to 2D matrix")
img = img.mean(2)
# initialize output array
img = img.astype('float32')
imgout = img.copy()
# initialize some internal variables
deltaS = np.zeros_like(imgout)
deltaE = deltaS.copy()
NS = deltaS.copy()
EW = deltaS.copy()
gS = np.ones_like(imgout)
gE = gS.copy()
# create the plot figure, if requested
if ploton:
import pylab as pl
from time import sleep
fig = pl.figure(figsize=(20,5.5),num="Anisotropic diffusion")
ax1,ax2 = fig.add_subplot(1,2,1),fig.add_subplot(1,2,2)
ax1.imshow(img,interpolation='nearest')
ih = ax2.imshow(imgout,interpolation='nearest',animated=True)
ax1.set_title("Original image")
ax2.set_title("Iteration 0")
fig.canvas.draw()
for ii in np.arange(1,niter):
# calculate the diffs
deltaS[:-1,: ] = np.diff(imgout,axis=0)
deltaE[: ,:-1] = np.diff(imgout,axis=1)
if 0<sigma:
deltaSf=flt.gaussian_filter(deltaS,sigma);
deltaEf=flt.gaussian_filter(deltaE,sigma);
else:
deltaSf=deltaS;
deltaEf=deltaE;
# conduction gradients (only need to compute one per dim!)
if option == 1:
gS = np.exp(-(deltaSf/kappa)**2.)/step[0]
gE = np.exp(-(deltaEf/kappa)**2.)/step[1]
elif option == 2:
gS = 1./(1.+(deltaSf/kappa)**2.)/step[0]
gE = 1./(1.+(deltaEf/kappa)**2.)/step[1]
# update matrices
E = gE*deltaE
S = gS*deltaS
# subtract a copy that has been shifted 'North/West' by one
# pixel. don't as questions. just do it. trust me.
NS[:] = S
EW[:] = E
NS[1:,:] -= S[:-1,:]
EW[:,1:] -= E[:,:-1]
# update the image
imgout += gamma*(NS+EW)
if ploton:
iterstring = "Iteration %i" %(ii+1)
ih.set_data(imgout)
ax2.set_title(iterstring)
fig.canvas.draw()
# sleep(0.01)
return imgout- 用法
#茴香( niter=1,kappa=50,gamma=0.1,step=(1,1.),sigma=0,option=1,ploton=False)
us_im_ad = anisodiff(ultrasound,100,80,0.075,(1,1),2.5,1)并列比较
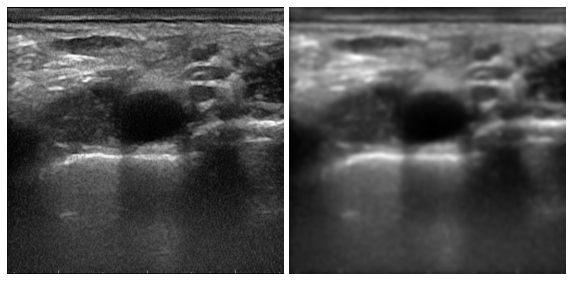
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/14715817
复制相关文章
相似问题

