试图复制数据时发生意外错误
试图复制数据时发生意外错误
提问于 2021-11-26 11:51:37
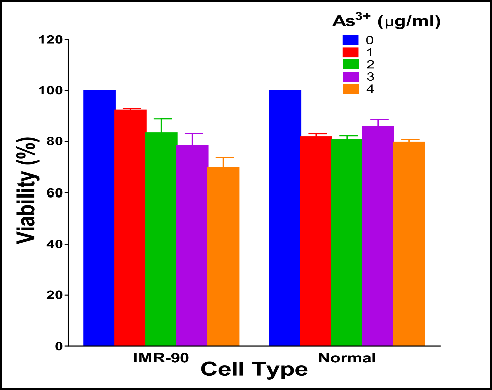
我是新的R,并试图建立一个分组条形图,显示两组为两种不同的情况。我是一个初学者,所以我试图复制我在网上看到的例子。
#create dataframe
Gene<-c("IMR-90","Normal","IMR-90","Normal","IMR-90","Normal","IMR-90","Normal","IMR-90","Normal")
count1<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,69.75771,79.64783)
count2<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,69.75771,79.64783)
count3<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,69.75771,79.64783)
count4<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,69.75771,79.64783)
Species<-c("0","0","1","1","2","2","3","3","4","4")
cols = c(2,3,4,5,6,7,8,9,10,11)
df<-data.frame(Gene,count1,count2,count3,count4,Species)
df1 = transform(df, mean=rowMeans(df[cols]), sd=apply(df[cols],1, sd))
ggplot(df1, aes(x=as.factor(Gene), y=mean, fill=Species)) +
geom_bar(position=position_dodge(), stat="identity", colour='black') +
geom_errorbar(aes(ymin=mean-sd, ymax=mean+sd), width=.2,position=position_dodge(.9))我的R代码在上面和下面是我想要实现的。
回答 2
Stack Overflow用户
回答已采纳
发布于 2021-11-26 12:41:54
似乎您的问题是在给我带来错误的转换中发现的。
要向数据添加均值和标准差,可以使用dplyr包和函数突变。要计算每一行的平均值和sd,只需向管道中添加行级()
Gene<-c("IMR-90","Normal","IMR-90","Normal","IMR-90","Normal","IMR-90","Normal","IMR-90","Normal")
count1<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,69.75771,79.64783)
count2<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,69.75771,79.64783)
count3<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,69.75771,79.64783)
count4<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,69.75771,79.64783)
Species<-c("0","0","1","1","2","2","3","3","4","4")
cols = c(2,3,4,5,6,7,8,9,10,11)
df<-data.frame(Gene,count1,count2,count3,count4,Species)
df1 <- df %>% rowwise() %>% mutate(mean=(count1+count2+count3+count4)/3, stdev = sd(c(count1, count2, count3, count4)))
ggplot(df1, aes(x=as.factor(Gene), y=mean, fill=Species)) +
geom_bar(position=position_dodge(), stat="identity", colour='black') +
geom_errorbar(aes(ymin=mean-stdev, ymax=mean+stdev), width=.2,position=position_dodge(.9))+
labs(title="title", x= "X axis name", y = "Y axis name")Stack Overflow用户
发布于 2021-11-26 12:39:22
最好先检查错误发生在哪里,因为您的图表没有任何问题(好的,也许没有全部完成,但没有抛出错误)。
你的数据准备出了问题。这也是明智的,如果你想要处理的标准差,你实际上给我们的数据,给我们一个SD值,而不是0 ;)
Gene<-c("IMR-90","Normal","IMR-90","Normal","IMR-90","Normal","IMR-90","Normal","IMR-90","Normal")
count1<-c(100,100,92.2453,81.76718,83.326,83.61812,78.34494,85.92033,62.75771,79.64783)
count2<-c(110,100,92.2453,87.76718,83.326,80.61812,78.34494,85.92033,65.75771,79.64783)
count3<-c(100,110,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,55.75771,70.64783)
count4<-c(100,100,92.2453,81.76718,83.326,80.61812,78.34494,85.92033,68.75771,79.64783)
Species<-c("0","0","1","1","2","2","3","3","4","4")
library(data.table)
df<- data.table(Gene, count1, count2, count3, count4, Species)
cols <- c("count1", "count2", "count3", "count4")
df1 <- df[, .(mean = rowMeans(.SD), sd = sd(.SD)), by = .(Gene, Species), .SDcols = cols]
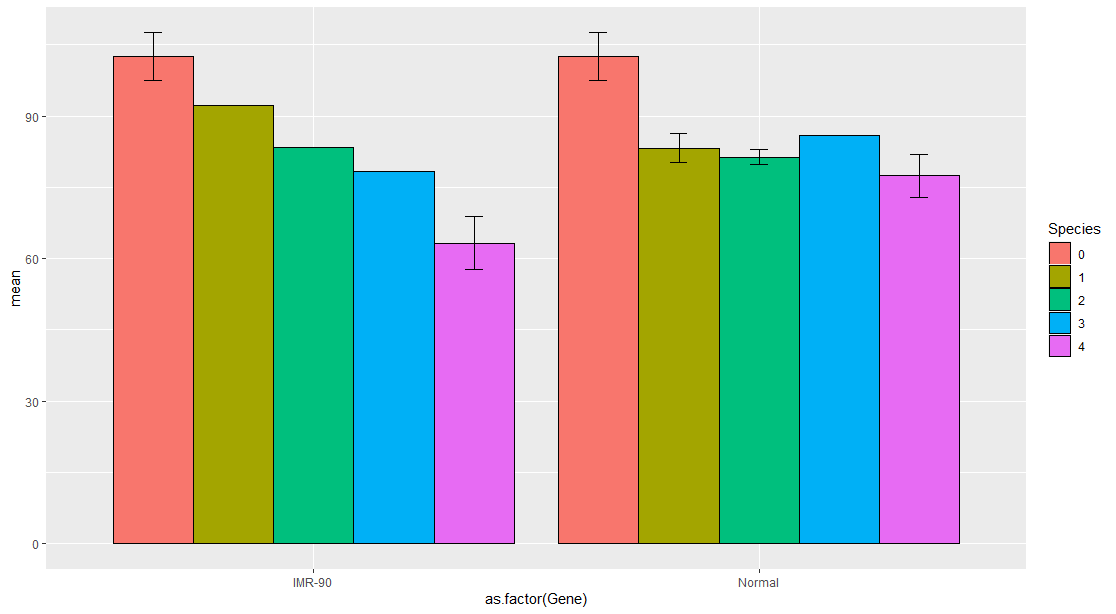
ggplot(df1, aes(x=as.factor(Gene), y=mean, fill=Species)) +
geom_bar(position=position_dodge(), stat="identity", colour='black') +
geom_errorbar(aes(ymin=mean-sd, ymax=mean+sd), width=.2,position=position_dodge(.9))
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/70123975
复制相关文章
相似问题

