使用stat_fit_tb()在ggpmisc & ggplot中编辑*行*表格显示的名称
使用stat_fit_tb()在ggpmisc & ggplot中编辑*行*表格显示的名称
提问于 2020-11-03 15:01:50
虽然stat_poly_eq()允许使用eq.with.lhs和eq.x.rhs更改变量名,但根据我对ggpmisc文档的阅读,类似的功能似乎在stat_fit_tb()中不可用。
下面的示例中是否有一种修改plt对象的方法,以强制表显示显示参数名称,这些参数名称更容易查看,并且更符合等式和轴标签?
## dummy data
set.seed(1)
df <- data.frame(month = c(1:60))
df$observed <- 2.5 + 0.05*df$month + rnorm(60, sd = 1)## min plot example
my.formula <- y ~ poly(x,2,raw=TRUE) ## formula with generic variable names
plt <- ggplot(df, aes(x=month, y=observed)) +
geom_point() +
## show fit and CI
geom_smooth(method = "lm", se=TRUE, level=0.95, formula = my.formula) +
## display equation with useful variable names (i.e. not x and y)
stat_poly_eq(eq.with.lhs = "italic(Obs)~`=`~",
eq.x.rhs = ".month",
aes(label = paste(..eq.label.., ..rr.label.., sep = "~~~")),
parse = TRUE,
formula = my.formula, label.y = 0.9) +
## show table of each coefficient's p-value
stat_fit_tb(method.args = list(formula = my.formula),
tb.vars = c(parameter = "term", ## can change column headings
coeff = "estimate",
"p-val" = "p.value"),
label.y = 0.8, label.x = "left")
plt
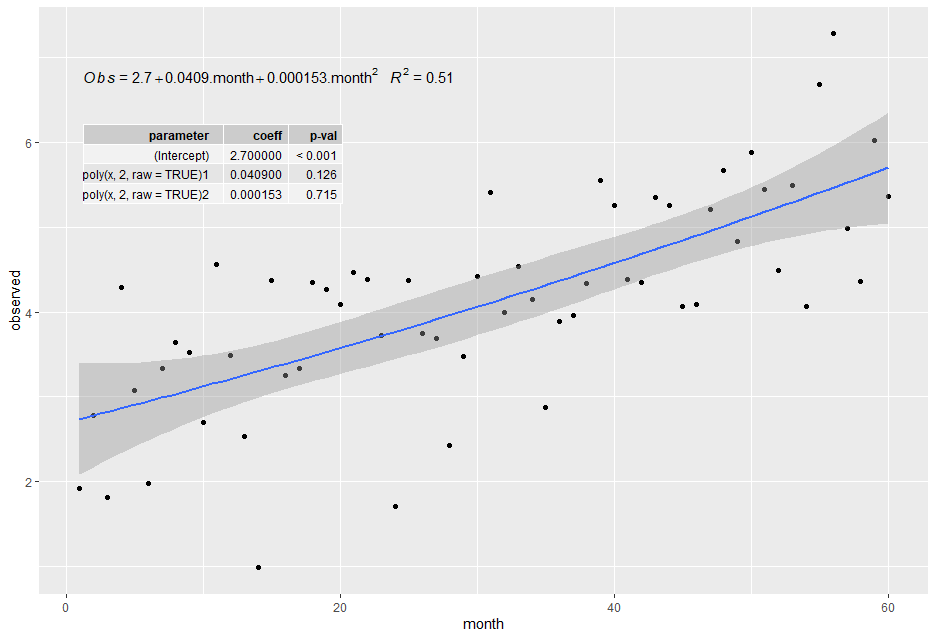
回答 3
Stack Overflow用户
回答已采纳
发布于 2020-11-09 21:30:11
更新的'ggpmisc‘(>= 0.3.7)使这个答案成为可能,在我看来应该是首选的答案。
## ggpmisc (>= 0.3.7)
library(ggpmisc)
## dummy data
set.seed(1)
df <- data.frame(month = c(1:60))
df$observed <- 2.5 + 0.05*df$month + rnorm(60, sd = 1)
## min plot example
my.formula <- y ~ poly(x,2,raw=TRUE) ## formula with generic variable names
plt <- ggplot(df, aes(x=month, y=observed)) +
geom_point() +
## show fit and CI
geom_smooth(method = "lm", se=TRUE, level=0.95, formula = my.formula) +
## display equation with useful variable names (i.e. not x and y)
stat_poly_eq(eq.with.lhs = "italic(Obs)~`=`~",
eq.x.rhs = '" month"',
aes(label = paste(..eq.label.., ..rr.label.., sep = "~~~")),
parse = TRUE,
formula = my.formula, label.y = 0.9) +
## show table of each coefficient's p-value
stat_fit_tb(method.args = list(formula = my.formula),
tb.vars = c(parameter = "term", ## can change column headings
coeff = "estimate",
"p-val" = "p.value"),
tb.params = c(1, month = 2, "month^2" = 3), ##
label.y = 0.8, label.x = "left",
parse = TRUE)
plt给出下面的情节。(我也把论点改成了eq.x.rhs,虽然不是问题的直接部分。P-值更好的格式是在新版本的“ggpmisc”包中实现的。)
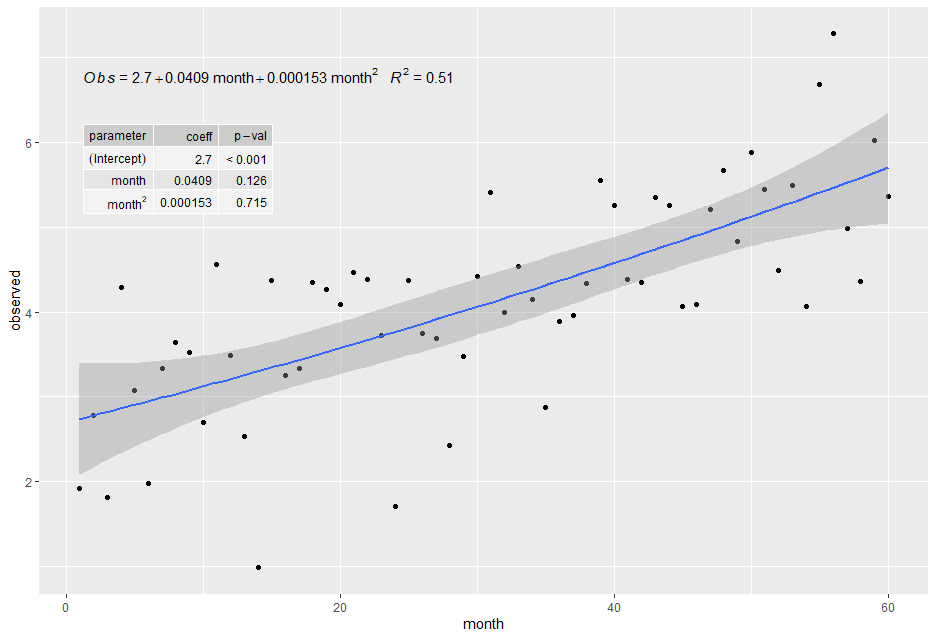
Stack Overflow用户
发布于 2020-11-04 09:23:41
在将plt转换成grob对象之后,这可能会被黑客攻击,但是现在我喜欢解决一次问题&用它来解决问题,所以我用底层的ggproto对象进行了黑客攻击。
运行以下代码(注释中显示了原始代码中的更改):
library(ggpmisc)
StatFitTb2 <- ggproto(
"StatFitTb2",
StatFitTb,
compute_panel = function (data, scales, method, method.args, tb.type, tb.vars,
tb.row.names, digits, npc.used = TRUE, label.x, label.y) {
force(data)
if (length(unique(data$x)) < 2) {
return(data.frame())
}
panel.idx <- as.integer(as.character(data$PANEL[1]))
if (length(label.x) >= panel.idx) {
label.x <- label.x[panel.idx]
}
else if (length(label.x) > 0) {
label.x <- label.x[1]
}
if (length(label.y) >= panel.idx) {
label.y <- label.y[panel.idx]
}
else if (length(label.y) > 0) {
label.y <- label.y[1]
}
method.args <- c(method.args, list(data = quote(data)))
if (is.character(method))
method <- match.fun(method)
mf <- do.call(method, method.args)
if (tolower(tb.type) %in% c("fit.anova", "anova")) {
mf_tb <- broom::tidy(stats::anova(mf))
}
else if (tolower(tb.type) %in% c("fit.summary", "summary")) {
mf_tb <- broom::tidy(mf)
}
else if (tolower(tb.type) %in% c("fit.coefs", "coefs")) {
mf_tb <- broom::tidy(mf)[c("term", "estimate")]
}
num.cols <- sapply(mf_tb, is.numeric)
mf_tb[num.cols] <- signif(mf_tb[num.cols], digits = digits)
if (!is.null(tb.vars)) {
mf_tb <- dplyr::select(mf_tb, !!tb.vars)
}
# new condition for modifying row names, if they are specified
if(!is.null(tb.row.names)) {
mf_tb[, 1] <- tb.row.names
}
z <- tibble::tibble(mf_tb = list(mf_tb))
if (npc.used) {
margin.npc <- 0.05
}
else {
margin.npc <- 0
}
if (is.character(label.x)) {
label.x <- switch(label.x, right = (1 - margin.npc),
center = 0.5, centre = 0.5,
middle = 0.5, left = (0 + margin.npc))
if (!npc.used) {
x.delta <- abs(diff(range(data$x)))
x.min <- min(data$x)
label.x <- label.x * x.delta + x.min
}
}
if (is.character(label.y)) {
label.y <- switch(label.y, top = (1 - margin.npc), center = 0.5,
centre = 0.5, middle = 0.5, bottom = (0 + margin.npc))
if (!npc.used) {
y.delta <- abs(diff(range(data$y)))
y.min <- min(data$y)
label.y <- label.y * y.delta + y.min
}
}
if (npc.used) {
z$npcx <- label.x
z$x <- NA_real_
z$npcy <- label.y
z$y <- NA_real_
}
else {
z$x <- label.x
z$npcx <- NA_real_
z$y <- label.y
z$npcy <- NA_real_
}
z
})
stat_fit_tb2 <- function(mapping = NULL, data = NULL, geom = "table_npc",
method = "lm", method.args = list(formula = y ~ x),
tb.type = "fit.summary", tb.vars = NULL, digits = 3,
tb.row.names = NULL, # new parameter for row names (defaults to NULL)
label.x = "center", label.y = "top", label.x.npc = NULL,
label.y.npc = NULL, position = "identity", table.theme = NULL,
table.rownames = FALSE, table.colnames = TRUE, table.hjust = 1,
parse = FALSE, na.rm = FALSE, show.legend = FALSE, inherit.aes = TRUE,
...) {
if (!is.null(label.x.npc)) {
stopifnot(grepl("_npc", geom))
label.x <- label.x.npc
}
if (!is.null(label.y.npc)) {
stopifnot(grepl("_npc", geom))
label.y <- label.y.npc
}
ggplot2::layer(stat = StatFitTb2, # reference modified StatFitTb2 instead of the original
data = data, mapping = mapping,
geom = geom, position = position, show.legend = show.legend,
inherit.aes = inherit.aes,
params = list(method = method, method.args = method.args,
tb.type = tb.type, tb.vars = tb.vars,
tb.row.names = tb.row.names, # new parameter here
digits = digits, label.x = label.x, label.y = label.y,
npc.used = grepl("_npc", geom), table.theme = table.theme,
table.rownames = table.rownames, table.colnames = table.colnames,
table.hjust = table.hjust, parse = parse, na.rm = na.rm,
...))
}用法:
ggplot(df, aes(x=month, y=observed)) +
geom_point() +
## show fit and CI
geom_smooth(method = "lm", se=TRUE, level=0.95, formula = my.formula) +
## display equation with useful variable names (i.e. not x and y)
stat_poly_eq(eq.with.lhs = "italic(Obs)~`=`~",
eq.x.rhs = ".month",
aes(label = paste(..eq.label.., ..rr.label.., sep = "~~~")),
parse = TRUE,
formula = my.formula, label.y = 0.9) +
## show table of each coefficient's p-value
stat_fit_tb2(method.args = list(formula = my.formula),
tb.vars = c(parameter = "term", ## can change column headings
coeff = "estimate",
"p-val" = "p.value"),
tb.row.names = c("(Intercept)", "month", "month^2"),
label.y = 0.8, label.x = "left", parse = TRUE)注意:parse = TRUE使month^2行名看起来更好看,但它也会影响表中的所有其他值(例如p值的破折号变成减号,数字被四舍五入到不同的数字数,等等)。
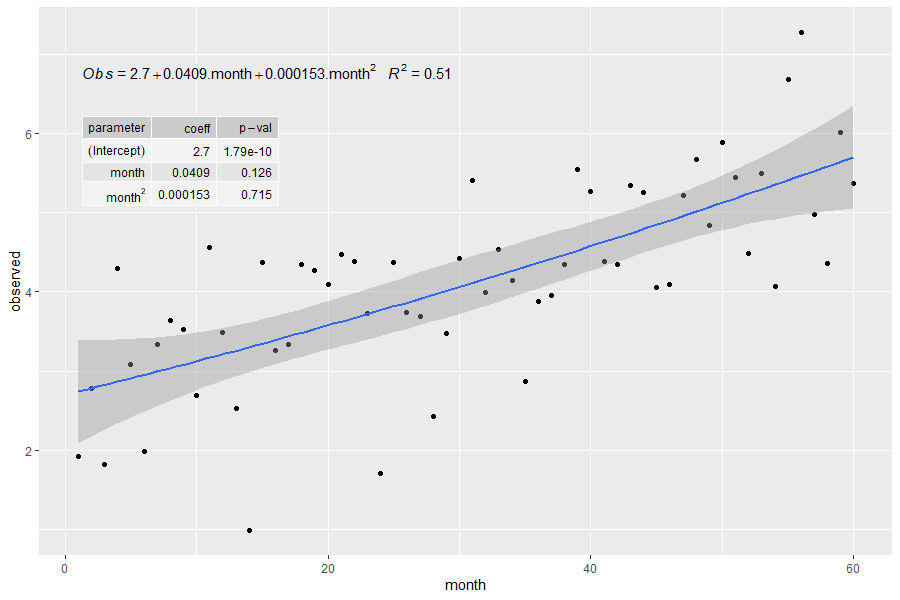
Stack Overflow用户
发布于 2020-11-04 23:44:03
注意:如果您仍然使用'ggpmisc‘(<= 0.3.6),这个答案可能是有用的。否则,请参见使用“ggpmisc”(>= 0.3.7)的单独答案。
在将其内置到包中之前,一种相当简单的方法是在aes()中动态编辑tibble。为了避免代码混乱,我首先定义了一个函数。
library(ggpmisc)
## dummy data
set.seed(1)
df <- data.frame(month = c(1:60))
df$observed <- 2.5 + 0.05*df$month + rnorm(60, sd = 1)
## min plot example
my.formula <- y ~ poly(x,2,raw=TRUE) ## formula with generic variable names
## define function for renaming parameters in tibble(s) returned by the stat
## walk through the list an operate on all the tibbles found so that
## grouping and facets are also supported.
set_param_names <- function(x, names) {
for (i in seq_along(x)) {
x[[i]][[1]] <- names
}
x
}
plt <- ggplot(df, aes(x=month, y=observed)) +
geom_point() +
## show fit and CI
geom_smooth(method = "lm", se=TRUE, level=0.95, formula = my.formula) +
## display equation with useful variable names (i.e. not x and y)
stat_poly_eq(eq.with.lhs = "italic(Obs)~`=`~",
eq.x.rhs = ".month",
aes(label = paste(..eq.label.., ..rr.label.., sep = "~~~")),
parse = TRUE,
formula = my.formula, label.y = 0.9) +
## show table of each coefficient's p-value
stat_fit_tb(method.args = list(formula = my.formula),
tb.vars = c(parameter = "term", ## can change column headings
coeff = "estimate",
"p-val" = "p.value"),
label.y = 0.8, label.x = "left",
aes(label = set_param_names(stat(mf_tb),
c("intercept", "month", "month^2"))),
parse = TRUE)
plt这意味着:
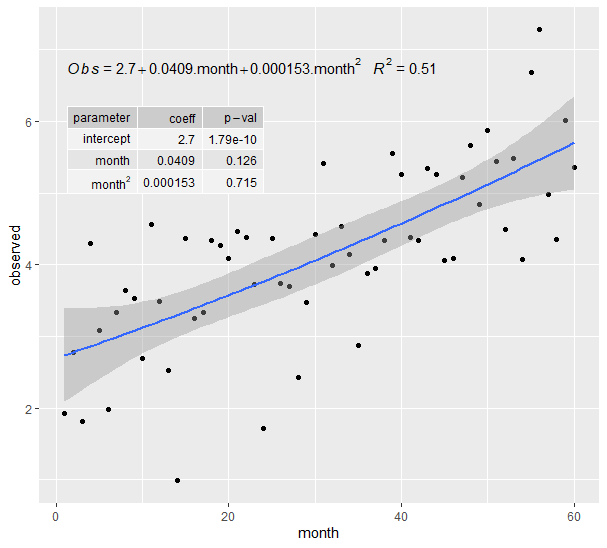
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/64665345
复制相关文章
相似问题

