为什么ggplot不显示一个方框的错误条?
为什么ggplot不显示一个方框的错误条?
提问于 2020-09-12 12:04:17
我在R中做了一些带有ggplot的盒子图,我想知道为什么它不显示一个框图的错误栏呢?
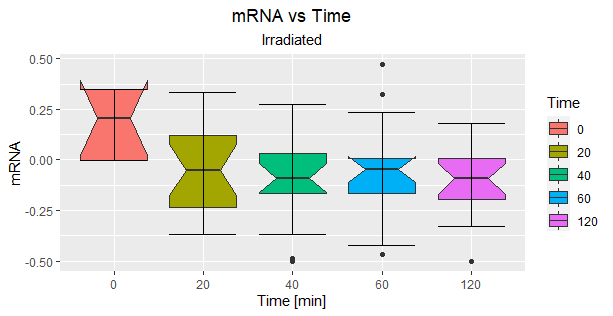
代码就是这样的:
ID1.4.5.6.7[,"Time"] <- as.factor(ID1.4.5.6.7[,"Time"])
ggplot(data=ID1.4.5.6.7,aes(x=Time, y=mRNA, fill=Time)) +
geom_boxplot(notch = TRUE) +
stat_boxplot(geom="errorbar")+
labs(title="mRNA vs Time", subtitle="Irradiated",x = "Time [min]",y = "mRNA")+
theme(plot.title = element_text(hjust = 0.5),plot.subtitle = element_text(hjust = 0.5))我不知道它是与代码相关的问题,还是与数据相关的问题,而不是问题。
structure(list(Gene = c("ID-1", "ID-1", "ID-1", "ID-1", "ID-1",
"ID-1", "ID-1", "ID-1", "ID-1", "ID-1", "ID-1", "ID-1", "ID-1",
"ID-1", "ID-1", "ID-4", "ID-4", "ID-4", "ID-4", "ID-4", "ID-4",
"ID-4", "ID-4", "ID-4", "ID-4", "ID-4", "ID-4", "ID-4", "ID-4",
"ID-4", "ID-4", "ID-5", "ID-5", "ID-5", "ID-5", "ID-5", "ID-5",
"ID-5", "ID-5", "ID-5", "ID-5", "ID-5", "ID-5", "ID-5", "ID-5",
"ID-5", "ID-5", "ID-5", "ID-5", "ID-6", "ID-6", "ID-6", "ID-6",
"ID-6", "ID-6", "ID-6", "ID-6", "ID-6", "ID-6", "ID-6", "ID-6",
"ID-6", "ID-6", "ID-6", "ID-6", "ID-6", "ID-6", "ID-7", "ID-7",
"ID-7", "ID-7", "ID-7", "ID-7", "ID-7", "ID-7", "ID-7", "ID-7",
"ID-7", "ID-7", "ID-7", "ID-7", "ID-7", "ID-7", "ID-7", "ID-7"
), mRNA = c(-0.181385669, -0.059647494, 0.104476117, -0.052190978,
-0.040484945, 0.194226742, -0.501601326, 0.102342605, -0.127143845,
-0.008523742, -0.102946211, -0.042894028, 0.002922923, -0.134394347,
-0.214204393, -0.138122686, 0.203242361, 0.097935502, 0.147068146,
-0.089430917, 0.331565412, -0.034572422, -0.129896329, 0.324191,
0.470108479, -0.027268223, 0.232304713, 0.090348708, 0.070848402,
0.181540708, -0.502255367, -0.267631441, -0.368647839, -0.040910404,
-0.003983171, -0.003983171, -0.003983171, -0.14980589, -0.119449612,
-0.309154214, -0.487589361, 0.272803506, -0.421733575, -0.467108567,
0.024868338, -0.156025729, -0.044680175, -0.206716896, -0.272014193,
-0.230499883, -0.238597397, -0.118130949, 0.349957464, 0.349957464,
0.349957464, 0.172048587, -0.186226994, 0.16113822, -0.293029136,
-0.111636253, -0.044189887, 0.081555274, -0.048106079, -0.05853566,
0.010407814, -0.066981809, -0.09828484, -0.315190986, -0.005102456,
0.221556197, 0.206584568, 0.206584568, 0.206584568, 0.102649006,
-0.011777384, -0.36963487, -0.054853074, -0.230240699, -0.210508323,
-0.208889919, -0.050763372, 0.023073782, -0.095118984, -0.091076071,
-0.330257395), Time = structure(c(2L, 2L, 2L, 3L, 3L, 2L, 3L,
3L, 4L, 4L, 4L, 4L, 5L, 5L, 5L, 2L, 2L, 2L, 3L, 3L, 2L, 3L, 3L,
4L, 4L, 4L, 4L, 5L, 5L, 5L, 5L, 2L, 2L, 2L, 1L, 1L, 1L, 3L, 3L,
2L, 3L, 3L, 4L, 4L, 4L, 5L, 5L, 5L, 5L, 2L, 2L, 2L, 1L, 1L, 1L,
3L, 3L, 2L, 3L, 3L, 4L, 4L, 4L, 4L, 5L, 5L, 5L, 2L, 2L, 2L, 1L,
1L, 1L, 3L, 2L, 3L, 3L, 4L, 4L, 4L, 4L, 5L, 5L, 5L, 5L), .Label = c("0",
"20", "40", "60", "120"), class = "factor"), predicted_mRNA = c(-0.00551000342030954,
-0.00551000342030954, -0.00551000342030954, -0.0302695238715682,
-0.0302695238715682, -0.00551000342030954, -0.0302695238715682,
-0.0302695238715682, -0.0550290443228268, -0.0550290443228268,
-0.0550290443228268, -0.0550290443228268, -0.129307605676603,
-0.129307605676603, -0.129307605676603, -0.00551000342030954,
-0.00551000342030954, -0.00551000342030954, -0.0302695238715682,
-0.0302695238715682, -0.00551000342030954, -0.0302695238715682,
-0.0302695238715682, -0.0550290443228268, -0.0550290443228268,
-0.0550290443228268, -0.0550290443228268, -0.129307605676603,
-0.129307605676603, -0.129307605676603, -0.129307605676603, -0.00551000342030954,
-0.00551000342030954, -0.00551000342030954, 0.0192495170309491,
0.0192495170309491, 0.0192495170309491, -0.0302695238715682,
-0.0302695238715682, -0.00551000342030954, -0.0302695238715682,
-0.0302695238715682, -0.0550290443228268, -0.0550290443228268,
-0.0550290443228268, -0.129307605676603, -0.129307605676603,
-0.129307605676603, -0.129307605676603, -0.00551000342030954,
-0.00551000342030954, -0.00551000342030954, 0.0192495170309491,
0.0192495170309491, 0.0192495170309491, -0.0302695238715682,
-0.0302695238715682, -0.00551000342030954, -0.0302695238715682,
-0.0302695238715682, -0.0550290443228268, -0.0550290443228268,
-0.0550290443228268, -0.0550290443228268, -0.129307605676603,
-0.129307605676603, -0.129307605676603, -0.00551000342030954,
-0.00551000342030954, -0.00551000342030954, 0.0192495170309491,
0.0192495170309491, 0.0192495170309491, -0.0302695238715682,
-0.00551000342030954, -0.0302695238715682, -0.0302695238715682,
-0.0550290443228268, -0.0550290443228268, -0.0550290443228268,
-0.0550290443228268, -0.129307605676603, -0.129307605676603,
-0.129307605676603, -0.129307605676603)), row.names = c(NA, -85L
), class = "data.frame")这里是dput(ID1.4.5.6.7),所以是dataframe。
回答 2
Stack Overflow用户
回答已采纳
发布于 2020-09-12 13:01:19
我建议采用这种方法,在这里您可以启用varwidth以查看错误栏。在这里,代码:
#Plot
ggplot(data=ID1.4.5.6.7,aes(x=Time, y=mRNA, fill=Time)) +
geom_boxplot(varwidth = TRUE,notch=TRUE) +
stat_boxplot(geom="errorbar")+
labs(title="mRNA vs Time", subtitle="Irradiated",x = "Time [min]",y = "mRNA")+
theme(plot.title = element_text(hjust = 0.5),plot.subtitle = element_text(hjust = 0.5))输出:
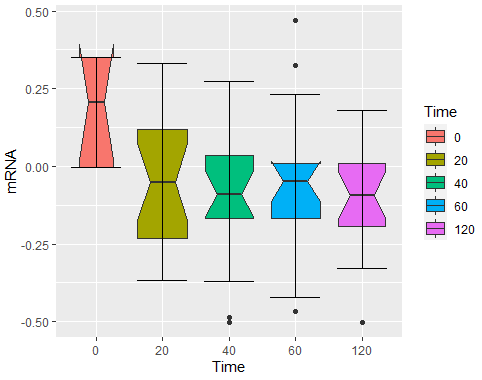
Stack Overflow用户
发布于 2020-09-12 12:48:00
因为方框没有错误条。盒子图只是五个数字的图形表示:最小,Q1 (第一个四分位数),中位数,Q3 (第三个四分位数)和最大值。晶须(向上和向下的“条形”)只是以数据中的最小值(下一条)和最大值(上一条)结尾的行。“框”的底部边缘是Q1,上边是Q3。
可以安排一组数据,使最小值与Q1相同,最大值与Q3相同。或多或少,这似乎是在没有胡须的盒子里发生的事情。ggplot在盒子图中添加了一些额外的细节(“腰部”被拉了进来,算法上的调整导致了在0组顶部出现倒排的可能性),但这似乎或多或少就是正在发生的事情。
编辑:这似乎是一个关于代码的问题,但实际上是关于统计的。交叉验证可能更好(虽然我认为它现在可能已经得到了充分的回答)。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/63860118
复制相关文章
相似问题

