筛选集合事件,如果同一日期发生多个集合事件,则只保留第一个事件。
筛选集合事件,如果同一日期发生多个集合事件,则只保留第一个事件。
提问于 2020-05-05 17:10:12
我已经包含了我拥有的数据的一小部分。
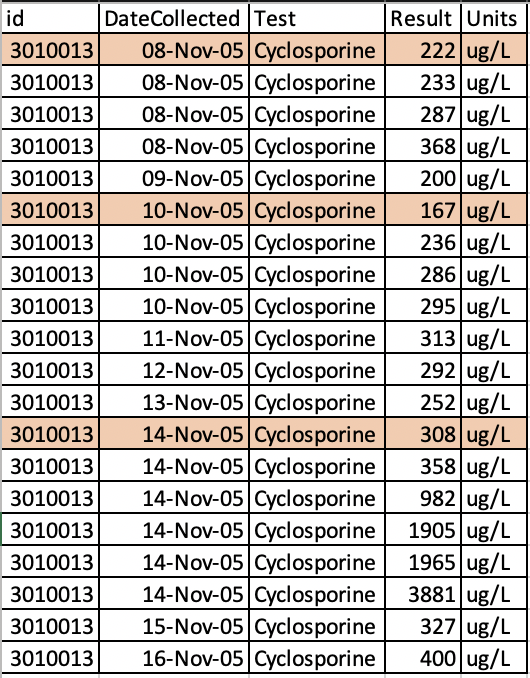
它包含治疗药物监测水平的日期。我只需要包括第一个事件的病人,有多个在同一天,并删除其他。
我在下面的图片中突出了一些例子。
structure(list(id = c(3010013, 3010013, 3010013, 3010013, 3010013,
3010013, 3010013, 3010013, 3010013, 3010013, 3010013, 3010013,
3010013, 3010013, 3010013, 3010013, 3010013, 3010013, 3010013,
3010013), DateCollected = structure(c(1131408000, 1131408000,
1131408000, 1131408000, 1131494400, 1131580800, 1131580800, 1131580800,
1131580800, 1131667200, 1131753600, 1131840000, 1131926400, 1131926400,
1131926400, 1131926400, 1131926400, 1131926400, 1132012800, 1132099200
), class = c("POSIXct", "POSIXt"), tzone = "UTC"), Test = c("Cyclosporine",
"Cyclosporine", "Cyclosporine", "Cyclosporine", "Cyclosporine",
"Cyclosporine", "Cyclosporine", "Cyclosporine", "Cyclosporine",
"Cyclosporine", "Cyclosporine", "Cyclosporine", "Cyclosporine",
"Cyclosporine", "Cyclosporine", "Cyclosporine", "Cyclosporine",
"Cyclosporine", "Cyclosporine", "Cyclosporine"), Result = c(222,
233, 287, 368, 200, 167, 236, 286, 295, 313, 292, 252, 308, 358,
982, 1905, 1965, 3881, 327, 400), Units = c("ug/L", "ug/L", "ug/L",
"ug/L", "ug/L", "ug/L", "ug/L", "ug/L", "ug/L", "ug/L", "ug/L",
"ug/L", "ug/L", "ug/L", "ug/L", "ug/L", "ug/L", "ug/L", "ug/L",
"ug/L")), row.names = c(NA, -20L), class = c("tbl_df", "tbl",
"data.frame"))
回答 3
Stack Overflow用户
回答已采纳
发布于 2020-05-05 17:50:37
在base R中,我们可以使用duplicated
df[!duplicated(df[c('id', 'DateCollected', 'Test')]),]或者在filter和duplicated中使用dplyr
library(dplyr)
df %>%
filter(!duplicated(select(., id, DateCollected, Test)))
# A tibble: 9 x 5
# id DateCollected Test Result Units
# <dbl> <dttm> <chr> <dbl> <chr>
#1 3010013 2005-11-08 00:00:00 Cyclosporine 222 ug/L
#2 3010013 2005-11-09 00:00:00 Cyclosporine 200 ug/L
#3 3010013 2005-11-10 00:00:00 Cyclosporine 167 ug/L
#4 3010013 2005-11-11 00:00:00 Cyclosporine 313 ug/L
#5 3010013 2005-11-12 00:00:00 Cyclosporine 292 ug/L
#6 3010013 2005-11-13 00:00:00 Cyclosporine 252 ug/L
#7 3010013 2005-11-14 00:00:00 Cyclosporine 308 ug/L
#8 3010013 2005-11-15 00:00:00 Cyclosporine 327 ug/L
#9 3010013 2005-11-16 00:00:00 Cyclosporine 400 ug/L Stack Overflow用户
发布于 2020-05-05 17:35:19
使用dplyr软件包
> df %>% group_by(DateCollected) %>%
summarize(id=first(id), first(Test), first(Result), first(Units)) %>%
ungroup()
## A tibble: 9 x 5
# DateCollected id `first(Test)` `first(Result)` `first(Units)`
# <dttm> <dbl> <chr> <dbl> <chr>
#1 2005-11-08 00:00:00 3010013 Cyclosporine 222 ug/L
#2 2005-11-09 00:00:00 3010013 Cyclosporine 200 ug/L
#3 2005-11-10 00:00:00 3010013 Cyclosporine 167 ug/L
#4 2005-11-11 00:00:00 3010013 Cyclosporine 313 ug/L
#5 2005-11-12 00:00:00 3010013 Cyclosporine 292 ug/L
#6 2005-11-13 00:00:00 3010013 Cyclosporine 252 ug/L
#7 2005-11-14 00:00:00 3010013 Cyclosporine 308 ug/L
#8 2005-11-15 00:00:00 3010013 Cyclosporine 327 ug/L
#9 2005-11-16 00:00:00 3010013 Cyclosporine 400 ug/L Stack Overflow用户
发布于 2020-05-05 17:45:54
您可以只希望从单个行中group_by列。在这种情况下,您可能只需要重复的第一行id、DateCollected和Test组合。
library(dplyr)
df %>%
group_by(id, DateCollected, Test) %>%
slice(1)输出
# A tibble: 9 x 5
# Groups: id, DateCollected, Test [9]
id DateCollected Test Result Units
<dbl> <dttm> <chr> <dbl> <chr>
1 3010013 2005-11-08 00:00:00 Cyclosporine 222 ug/L
2 3010013 2005-11-09 00:00:00 Cyclosporine 200 ug/L
3 3010013 2005-11-10 00:00:00 Cyclosporine 167 ug/L
4 3010013 2005-11-11 00:00:00 Cyclosporine 313 ug/L
5 3010013 2005-11-12 00:00:00 Cyclosporine 292 ug/L
6 3010013 2005-11-13 00:00:00 Cyclosporine 252 ug/L
7 3010013 2005-11-14 00:00:00 Cyclosporine 308 ug/L
8 3010013 2005-11-15 00:00:00 Cyclosporine 327 ug/L
9 3010013 2005-11-16 00:00:00 Cyclosporine 400 ug/L 页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/61618888
复制相关文章
相似问题

