R:合并要导出到excel的行
R:合并要导出到excel的行
提问于 2019-12-30 19:55:51
如果列中的值相同(在唯一标识符组中),则必须合并excel的行。我附上了当前openxlsx输出和所需输出的照片。
我知道,在SAS中,您可以使用PROC报告,它会自动完成这一任务,所以我确信有一种方法可以做到。我尝试了柔性化,但我也需要一些条件格式,但它无法做到。
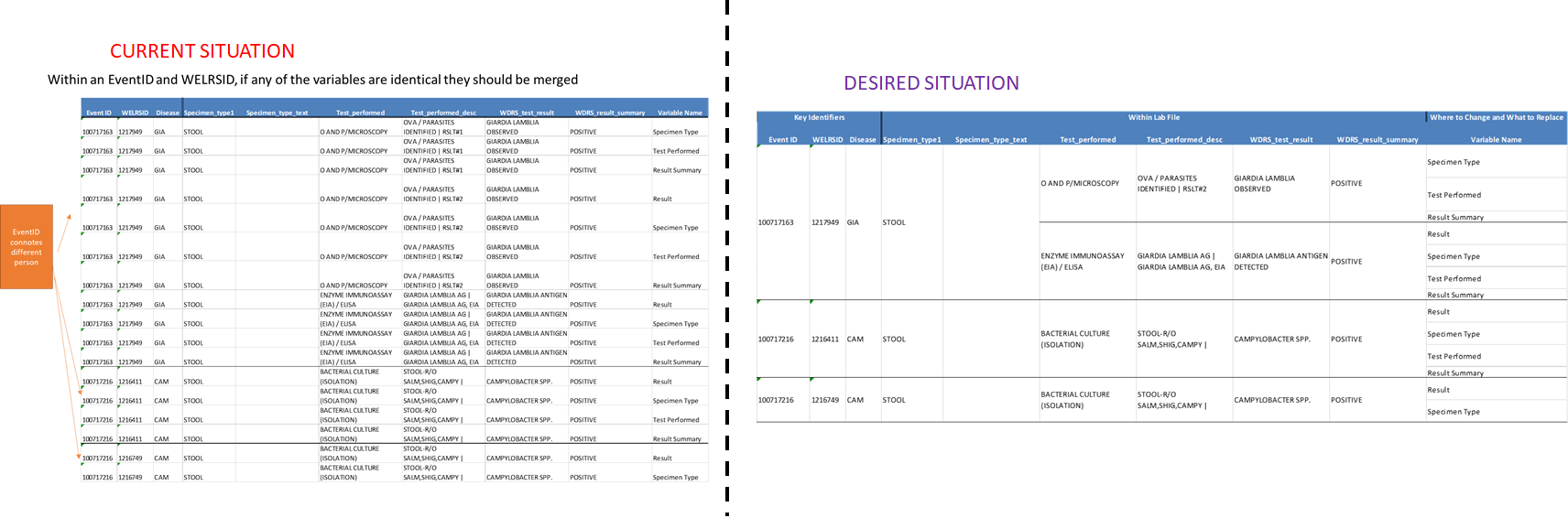
编辑:
数据如下:
structure(list(`Event ID` = c("100717163", "100717163", "100717163",
"100717163", "100717163", "100717163", "100717163", "100717163",
"100717163", "100717163", "100717163", "100717163", "100717163",
"100717163", "100717163", "100717163", "100717216", "100717216",
"100717216", "100717216", "100717216", "100717216", "100717216",
"100717216"), WELRSID = c("1215288", "1215288", "1215288", "1215288",
"1217949", "1217949", "1217949", "1217949", "1217949", "1217949",
"1217949", "1217949", "1217949", "1217949", "1217949", "1217949",
"1216411", "1216411", "1216411", "1216411", "1216749", "1216749",
"1216749", "1216749"), Disease = c("GIA", "GIA", "GIA", "GIA",
"GIA", "GIA", "GIA", "GIA", "GIA", "GIA", "GIA", "GIA", "GIA",
"GIA", "GIA", "GIA", "CAM", "CAM", "CAM", "CAM", "CAM", "CAM",
"CAM", "CAM"), Specimen_type1 = c("STOOL", "STOOL", "STOOL",
"STOOL", "STOOL", "STOOL", "STOOL", "STOOL", "STOOL", "STOOL",
"STOOL", "STOOL", "STOOL", "STOOL", "STOOL", "STOOL", "STOOL",
"STOOL", "STOOL", "STOOL", "STOOL", "STOOL", "STOOL", "STOOL"
), Specimen_type_text = c(NA_character_, NA_character_, NA_character_,
NA_character_, NA_character_, NA_character_, NA_character_, NA_character_,
NA_character_, NA_character_, NA_character_, NA_character_, NA_character_,
NA_character_, NA_character_, NA_character_, NA_character_, NA_character_,
NA_character_, NA_character_, NA_character_, NA_character_, NA_character_,
NA_character_), Test_performed = c("ENZYME IMMUNOASSAY (EIA) / ELISA",
"ENZYME IMMUNOASSAY (EIA) / ELISA", "ENZYME IMMUNOASSAY (EIA) / ELISA",
"ENZYME IMMUNOASSAY (EIA) / ELISA", "O AND P/MICROSCOPY", "O AND P/MICROSCOPY",
"O AND P/MICROSCOPY", "O AND P/MICROSCOPY", "O AND P/MICROSCOPY",
"O AND P/MICROSCOPY", "O AND P/MICROSCOPY", "O AND P/MICROSCOPY",
"ENZYME IMMUNOASSAY (EIA) / ELISA", "ENZYME IMMUNOASSAY (EIA) / ELISA",
"ENZYME IMMUNOASSAY (EIA) / ELISA", "ENZYME IMMUNOASSAY (EIA) / ELISA",
"BACTERIAL CULTURE (ISOLATION)", "BACTERIAL CULTURE (ISOLATION)",
"BACTERIAL CULTURE (ISOLATION)", "BACTERIAL CULTURE (ISOLATION)",
"BACTERIAL CULTURE (ISOLATION)", "BACTERIAL CULTURE (ISOLATION)",
"BACTERIAL CULTURE (ISOLATION)", "BACTERIAL CULTURE (ISOLATION)"
), Test_performed_desc = c("GIARDIA LAMBLIA AG | GIARDIA LAMBLIA AG, EIA",
"GIARDIA LAMBLIA AG | GIARDIA LAMBLIA AG, EIA", "GIARDIA LAMBLIA AG | GIARDIA LAMBLIA AG, EIA",
"GIARDIA LAMBLIA AG | GIARDIA LAMBLIA AG, EIA", "OVA / PARASITES IDENTIFIED | RSLT#1",
"OVA / PARASITES IDENTIFIED | RSLT#1", "OVA / PARASITES IDENTIFIED | RSLT#1",
"OVA / PARASITES IDENTIFIED | RSLT#1", "OVA / PARASITES IDENTIFIED | RSLT#2",
"OVA / PARASITES IDENTIFIED | RSLT#2", "OVA / PARASITES IDENTIFIED | RSLT#2",
"OVA / PARASITES IDENTIFIED | RSLT#2", "GIARDIA LAMBLIA AG | GIARDIA LAMBLIA AG, EIA",
"GIARDIA LAMBLIA AG | GIARDIA LAMBLIA AG, EIA", "GIARDIA LAMBLIA AG | GIARDIA LAMBLIA AG, EIA",
"GIARDIA LAMBLIA AG | GIARDIA LAMBLIA AG, EIA", "STOOL-R/O SALM,SHIG,CAMPY |",
"STOOL-R/O SALM,SHIG,CAMPY |", "STOOL-R/O SALM,SHIG,CAMPY |",
"STOOL-R/O SALM,SHIG,CAMPY |", "STOOL-R/O SALM,SHIG,CAMPY |",
"STOOL-R/O SALM,SHIG,CAMPY |", "STOOL-R/O SALM,SHIG,CAMPY |",
"STOOL-R/O SALM,SHIG,CAMPY |"), WDRS_test_result = c("GIARDIA LAMBLIA ANTIGEN DETECTED",
"GIARDIA LAMBLIA ANTIGEN DETECTED", "GIARDIA LAMBLIA ANTIGEN DETECTED",
"GIARDIA LAMBLIA ANTIGEN DETECTED", "GIARDIA LAMBLIA OBSERVED",
"GIARDIA LAMBLIA OBSERVED", "GIARDIA LAMBLIA OBSERVED", "GIARDIA LAMBLIA OBSERVED",
"GIARDIA LAMBLIA OBSERVED", "GIARDIA LAMBLIA OBSERVED", "GIARDIA LAMBLIA OBSERVED",
"GIARDIA LAMBLIA OBSERVED", "GIARDIA LAMBLIA ANTIGEN DETECTED",
"GIARDIA LAMBLIA ANTIGEN DETECTED", "GIARDIA LAMBLIA ANTIGEN DETECTED",
"GIARDIA LAMBLIA ANTIGEN DETECTED", "CAMPYLOBACTER SPP.", "CAMPYLOBACTER SPP.",
"CAMPYLOBACTER SPP.", "CAMPYLOBACTER SPP.", "CAMPYLOBACTER SPP.",
"CAMPYLOBACTER SPP.", "CAMPYLOBACTER SPP.", "CAMPYLOBACTER SPP."
), WDRS_result_summary = c("POSITIVE", "POSITIVE", "POSITIVE",
"POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE",
"POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE",
"POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE", "POSITIVE",
"POSITIVE", "POSITIVE", "POSITIVE"), WDRSresult_notcoded = c(NA_character_,
NA_character_, NA_character_, NA_character_, NA_character_, NA_character_,
NA_character_, NA_character_, NA_character_, NA_character_, NA_character_,
NA_character_, NA_character_, NA_character_, NA_character_, NA_character_,
NA_character_, NA_character_, NA_character_, NA_character_, NA_character_,
NA_character_, NA_character_, NA_character_), Test_result = c("POSITIVE | POSITIVE",
"POSITIVE | POSITIVE", "POSITIVE | POSITIVE", "POSITIVE | POSITIVE",
"GIARDIA LAMBLIA CYST | GIARDIA LAMBLIA CYSTS.", "GIARDIA LAMBLIA CYST | GIARDIA LAMBLIA CYSTS.",
"GIARDIA LAMBLIA CYST | GIARDIA LAMBLIA CYSTS.", "GIARDIA LAMBLIA CYST | GIARDIA LAMBLIA CYSTS.",
"GIARDIA LAMBLIA TROPHOZOITE | GIARDIA LAMBLIA TROPHOZOITES",
"GIARDIA LAMBLIA TROPHOZOITE | GIARDIA LAMBLIA TROPHOZOITES",
"GIARDIA LAMBLIA TROPHOZOITE | GIARDIA LAMBLIA TROPHOZOITES",
"GIARDIA LAMBLIA TROPHOZOITE | GIARDIA LAMBLIA TROPHOZOITES",
"POSITIVE | POSITIVE", "POSITIVE | POSITIVE", "POSITIVE | POSITIVE",
"POSITIVE | POSITIVE", "CAMPYLOBACTER SPECIES |", "CAMPYLOBACTER SPECIES |",
"CAMPYLOBACTER SPECIES |", "CAMPYLOBACTER SPECIES |", "CAMPYLOBACTER SPECIES |",
"CAMPYLOBACTER SPECIES |", "CAMPYLOBACTER SPECIES |", "CAMPYLOBACTER SPECIES |"
), `Variable Name` = structure(c(1L, 3L, 4L, 2L, 1L, 3L, 4L,
2L, 1L, 3L, 4L, 2L, 1L, 3L, 4L, 2L, 1L, 3L, 4L, 2L, 1L, 3L, 4L,
2L), .Label = c("Result", "Result Summary", "Specimen Type",
"Test Performed"), class = "factor"), `Change to this (only if Red)` = c("GIARDIA LAMBLIA ANTIGEN DETECTED",
"STOOL", "ENZYME IMMUNOASSAY (EIA) / ELISA", "POSITIVE", "GIARDIA LAMBLIA OBSERVED",
"STOOL", "O AND P/MICROSCOPY", "POSITIVE", "GIARDIA LAMBLIA OBSERVED",
"STOOL", "O AND P/MICROSCOPY", "POSITIVE", "GIARDIA LAMBLIA ANTIGEN DETECTED",
"STOOL", "ENZYME IMMUNOASSAY (EIA) / ELISA", "POSITIVE", "CAMPYLOBACTER SPP.",
"STOOL", "BACTERIAL CULTURE (ISOLATION)", "POSITIVE", "CAMPYLOBACTER SPP.",
"STOOL", "BACTERIAL CULTURE (ISOLATION)", "POSITIVE"), Error = c("No Error",
"No Error", "No Error", "No Error", "No Error", "No Error", "No Error",
"No Error", "No Error", "No Error", "No Error", "No Error", "No Error",
"No Error", "No Error", "No Error", "No Error", "No Error", "No Error",
"No Error", "No Error", "No Error", "No Error", "No Error"),
Error2 = c(0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0)), row.names = c(NA, -24L), class = c("tbl_df",
"tbl", "data.frame"))码
addWorksheet(wb,“数据”)
hs1 <- createStyle(fgFill = "#4F81BD", halign = "CENTER", textDecoration = "Bold",
border = c("Bottom"), fontColour = "white", borderStyle = "double")
hs2 <- createStyle(fgFill = "#4F81BD", halign = "CENTER", textDecoration = "Bold",
border = c("Bottom", "Right"), fontColour = "white", borderStyle = "double")
title <- createStyle(fgFill = "#4F81BD", halign = "CENTER", textDecoration = "Bold", border = "Left", fontColour = "white", borderStyle = "double")
duplicate <- createStyle(border = "Bottom")
text <- createStyle(wrapText = TRUE)
highlighting <- createStyle(fontColour = "red")
writeData(wb, "data", excel2, startRow = 2, headerStyle = hs1)
writeData(wb, "data", x = "Key Identifiers", startRow = 1, startCol = 1)
writeData(wb, "data", x = "Within Lab File", startRow = 1, startCol = 4)
writeData(wb, "data", x = "Where to Change and What to Replace", startRow = 1, startCol = 12)
mergeCells(wb, "data", cols = c(1:3), rows = 1)
mergeCells(wb, "data", cols = c(12:13), rows = 1)
mergeCells(wb, "data", cols = c(4:11), rows = 1)
addStyle(wb, "data", rows = 1, cols = 1, gridExpand = TRUE, style = title)
addStyle(wb, "data", rows = 1, cols = 4, gridExpand = TRUE, style = title)
addStyle(wb, "data", rows = 1, cols = 12, gridExpand = TRUE, style = title)
addStyle(wb, "data", rows = 2, cols = 3, gridExpand = TRUE, style = hs2)
addStyle(wb, "data", rows = 2, cols = 11, gridExpand = TRUE, style = hs2)
addStyle(wb, "data", rows = 2, cols = 13, gridExpand = TRUE, style = hs2)
addStyle(wb, "data", text, rows = c(2:nrow(excel)), cols = c(1:15), stack = TRUE, gridExpand =TRUE)
setColWidths(wb, "data", cols = c(1:15), widths = c(10, 10, 8, 15, 24, 24, 24, 24, 24, 24, 24, 16, "auto", 15, 15))
setColWidths(wb, "data", cols = c(14:15), hidden = TRUE)
conditionalFormatting(wb, "data", cols = 13, rows = c(3:nrow(excel)), rule = "O3>=1", style = highlighting)
conditionalFormatting(wb, "data", cols = 1:13, rows = c(3:nrow(excel)), rule = "$B3 != $B4", style = duplicate)
conditionalFormatting(wb, "data", cols = 2, rows = c(3:nrow(excel)), rule = "$B3 != $B4", color = "blue", showValue = FALSE,
)
saveWorkbook(wb, "Data Dashboard.xlsx", overwrite = TRUE)回答 1
Stack Overflow用户
发布于 2019-12-31 16:57:51
不是完全修复,而是能够产生合并单元格的错觉。
empty <- createStyle(fontColour = "white")
conditionalFormatting(wb, "data", cols = 2, rows = c(4:nrow(excel)), rule = "$B4 = $B3", style = empty)
conditionalFormatting(wb, "data", cols = 3, rows = c(4:nrow(excel)), rule = "$C4 = $C3", style = empty)
conditionalFormatting(wb, "data", cols = 4, rows = c(4:nrow(excel)), rule = "AND($D4=$D3,$B4 = $B3)", style = empty)
conditionalFormatting(wb, "data", cols = 5, rows = c(4:nrow(excel)), rule = "AND($E4=$E3,$B4 = $B3)", style = empty)页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/59536200
复制相关文章
相似问题

