单细胞分析中的叠加条形图
单细胞分析中的叠加条形图
提问于 2020-01-11 23:16:41
我正试图用下面的代码用ggplot2绘制一个堆叠的桶形图
barplot <- ggplot() + geom_bar(aes(y = percentage, x = TBD, fill = TBD), data = charts.data, stat="identity")我想为我的单细胞分析创建一个条形图,它有两个条件,一个是经过处理的,一个是未经处理的。我想要显示的是,在每个条件下,不同的单元格类型所占的百分比,看看处理后是否会对不同的单元类型产生影响。
如何在每种情况下确定每种单元格类型的百分比,然后再绘制桶形图?
dput(head(comparison))的输出
structure(c(6051L, 1892L, 1133L, 893L, 148L, 868L, 5331L, 3757L,
1802L, 1061L, 2786L, 704L), .Dim = c(6L, 2L), .Dimnames = structure(list(c("Fibroblast", "T cell", "Macrophage", "Stellate", "Acinar", "Endothelial"), c("treated", "untreated")), .Names = c("",
"")), class = "table")dput(head(cell_cycle_data))的输出
structure(list(orig.ident = c("treated", "treated", "treated",
"treated", "treated", "treated"), nCount_RNA = c(1892, 307, 1348,
3699, 4205, 4468), nFeature_RNA = c(960L, 243L, 765L, 1612L,
1341L, 1644L), percent.mt = c(0.211416490486258, 1.62866449511401,
4.45103857566766, 4.4065963773993, 0.0713436385255648, 3.87197851387645
), RNA_snn_res.0.5 = structure(c(11L, 11L, 5L, 6L, 11L, 13L), .Label = c("0", "1", "2", "3", "4", "5", "6", "7", "8", "9", "10", "11", "12",
"13", "14", "15", "16", "17", "18", "19"), class = "factor"), seurat_clusters = structure(c(11L, 11L, 5L, 6L, 11L, 13L), .Label = c("0", "1", "2", "3", "4", "5", "6", "7", "8", "9", "10", "11", "12", "13", "14", "15", "16", "17", "18", "19"), class = "factor"), S.Score = c(0.476893835992198, -0.0200784617568548, -0.0335915198305002, -0.0247184276246385, 0.010785196602457, 0.0190008903712199), G2M.Score = c(0.204441469200986, 0.173804859670862, -0.0313235510969097, -0.0376796363661889, -0.0559526905696905, -0.0122031631356698), Phase = structure(c(3L, 2L, 1L, 1L, 3L, 3L), .Label = c("G1", "G2M", "S"), class = "factor"), old.ident = structure(c(7L,7L, 1L, 4L, 7L, 9L), .Label = c("Fibroblast", "T cell", "Macrophage", "Stellate", "Acinar", "Endothelial", "Tumor", "B cell", "Mast cell", "Ductal", "Islets of Langerhans"), class = "factor")), row.names = c("treated_AAACGCTAGCGGGTTA-1", "treated_AAAGGTAAGTACAGAT-1", "treated_AAAGTGAGTTTGATCG-1", "treated_AAATGGACAAAGTGTA-1",
"treated_AACAAAGGTCGACTTA-1", "treated_AACAGGGTCCTAGCCT-1"), class = "data.frame")dput(tail(comparison))的输出
structure(list(orig.ident = c("untreated", "untreated", "untreated",
"untreated", "untreated", "untreated"), nCount_RNA = c(901, 823,
1184, 1835, 1147, 1407), nFeature_RNA = c(482L, 479L, 649L, 1043L,
604L, 709L), percent.mt = c(1.77580466148724, 2.91616038882138,
4.22297297297297, 3.86920980926431, 2.0052310374891, 4.05117270788913
), RNA_snn_res.0.5 = structure(c(7L, 7L, 7L, 14L, 7L, 7L), .Label = c("0",
"1", "2", "3", "4", "5", "6", "7", "8", "9", "10", "11", "12",
"13", "14", "15", "16", "17", "18", "19"), class = "factor"),
seurat_clusters = structure(c(7L, 7L, 7L, 14L, 7L, 7L), .Label = c("0",
"1", "2", "3", "4", "5", "6", "7", "8", "9", "10", "11",
"12", "13", "14", "15", "16", "17", "18", "19"), class = "factor"),
S.Score = c(-0.0320858200243315, 0.0304725660342869, 0.0215996091745327,
0.0384166213301423, 0.144956251122548, -0.0242770509986111
), G2M.Score = c(0.0904224391544142, 0.050148242050667, -0.0178041670730754,
-0.0112596867977946, -0.0519554524339088, -0.0136533184257381
), Phase = structure(c(2L, 2L, 3L, 3L, 3L, 1L), .Label = c("G1",
"G2M", "S"), class = "factor"), old.ident = structure(c(5L,
5L, 5L, 5L, 5L, 5L), .Label = c("Fibroblast", "T cell", "Macrophage",
"Stellate", "Acinar", "Endothelial", "Tumor", "B cell", "Mast cell",
"Ductal", "Islets of Langerhans"), class = "factor")), row.names = c("untreated_TTTGGTTGTCTAATCG-18",
"untreated_TTTGGTTTCCCGAGGT-18", "untreated_TTTGTTGAGAACTGAT-18",
"untreated_TTTGTTGAGCTCGGCT-18", "untreated_TTTGTTGAGTGCCTCG-18",
"untreated_TTTGTTGCACGGTGCT-18"), class = "data.frame")回答 1
Stack Overflow用户
发布于 2020-01-12 05:53:14
如果不知道数据的结构,就很难猜出示例的好代码是什么。
但是,如果我们假设每个条件都有,那么您就有一个单个单元格的列表,每个单元格都有对应于它们的单元格类型的特定标签,如下面的示例:
set.seed(123)
Untreated <- data.frame(Cell_Type = sample(LETTERS[1:4],10, replace = TRUE))
Treated <- data.frame(Cell_Type =sample(LETTERS[1:4],25, replace = TRUE))
Cell_Type
1 C
2 C
3 C
4 B
5 C
6 B
... ...您可以使用dplyr进行第一次bind_rows
library(dplyr)
Untreated <- Untreated %>% mutate(Condition = "Untreated")
Treated <- Treated %>% mutate(Condition = "Treated")
DF <- bind_rows(Untreated, Treated)
Cell_Type Condition
1 C Untreated
2 C Untreated
3 C Untreated
4 B Untreated
5 C Untreated
6 B Untreated然后,您可以将每个单元格类型的数量计数到每个条件中,并将其表示为百分比:
DF <- DF %>% group_by(Condition, Cell_Type) %>%
summarise(Nb = n()) %>%
mutate(C = sum(Nb)) %>%
mutate(percent = Nb/C*100)
# A tibble: 7 x 5
# Groups: Condition [2]
Condition Cell_Type Nb C percent
<chr> <chr> <int> <int> <dbl>
1 Treated A 7 25 28.
2 Treated B 7 25 28.
3 Treated C 6 25 24
4 Treated D 5 25 20
5 Untreated A 1 10 10
6 Untreated B 4 10 40
7 Untreated C 5 10 50 然后,您可以将结果绘制为每个条件的叠加条形图,并根据Cell_Type填充每种颜色:
library(ggplot2)
ggplot(DF, aes(x = Condition, y = percent, fill = Cell_Type))+
geom_bar(stat = "identity")+
geom_text(aes(label = paste(percent,"%")), position = position_stack(vjust = 0.5))
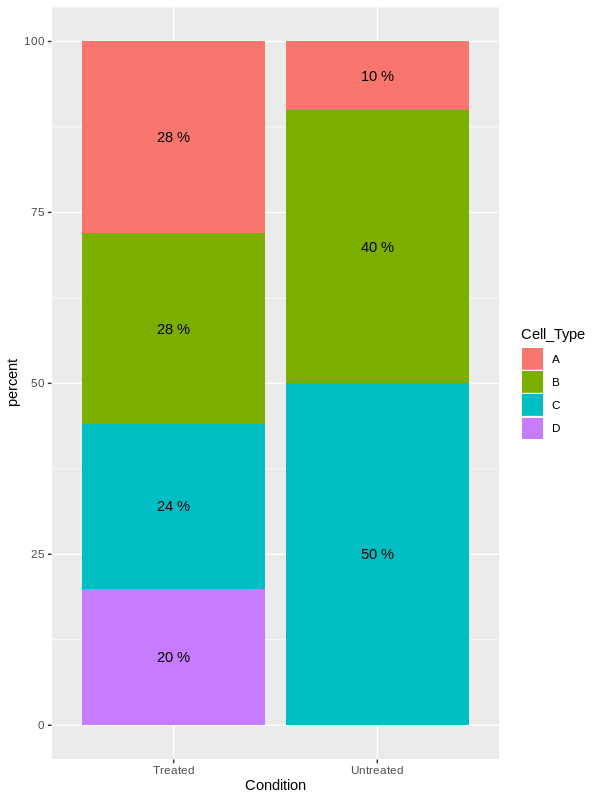
编辑:使用OP提供的数据绘图
使用您在问题中提供的数据,您可以:
df <- structure(c(6051L, 1892L, 1133L, 893L, 148L, 868L, 5331L, 3757L,
1802L, 1061L, 2786L, 704L), .Dim = c(6L, 2L), .Dimnames = structure(list(c("Fibroblast", "T cell", "Macrophage", "Stellate", "Acinar", "Endothelial"), c("treated", "untreated")), .Names = c("",
"")), class = "table")
df <- data.frame(df)它提供了以下数据:
Var1 Var2 Freq
1 Fibroblast treated 6051
2 T cell treated 1892
3 Macrophage treated 1133
4 Stellate treated 893
5 Acinar treated 148
6 Endothelial treated 868
7 Fibroblast untreated 5331
8 T cell untreated 3757
9 Macrophage untreated 1802
10 Stellate untreated 1061
11 Acinar untreated 2786
12 Endothelial untreated 704然后,您可以重命名列,为每个条件计算每种单元格类型的百分比:
library(dplyr)
DF <- df %>% rename(Cell_Type = Var1, Condition = Var2) %>%
group_by(Condition) %>%
mutate(Percent = Freq / sum(Freq)*100)
# A tibble: 12 x 4
# Groups: Condition [2]
Cell_Type Condition Freq Percent
<fct> <fct> <int> <dbl>
1 Fibroblast treated 6051 55.1
2 T cell treated 1892 17.2
3 Macrophage treated 1133 10.3
4 Stellate treated 893 8.13
5 Acinar treated 148 1.35
6 Endothelial treated 868 7.90
7 Fibroblast untreated 5331 34.5
8 T cell untreated 3757 24.3
9 Macrophage untreated 1802 11.7
10 Stellate untreated 1061 6.87
11 Acinar untreated 2786 18.0
12 Endothelial untreated 704 4.56然后,对于作图部分:
library(ggplot2)
ggplot(DF, aes(x = Condition, y = Percent, fill = Cell_Type))+
geom_bar(stat = "identity")+
geom_text(aes(label = paste(round(Percent,2),"%")), position = position_stack(vjust = 0.5))
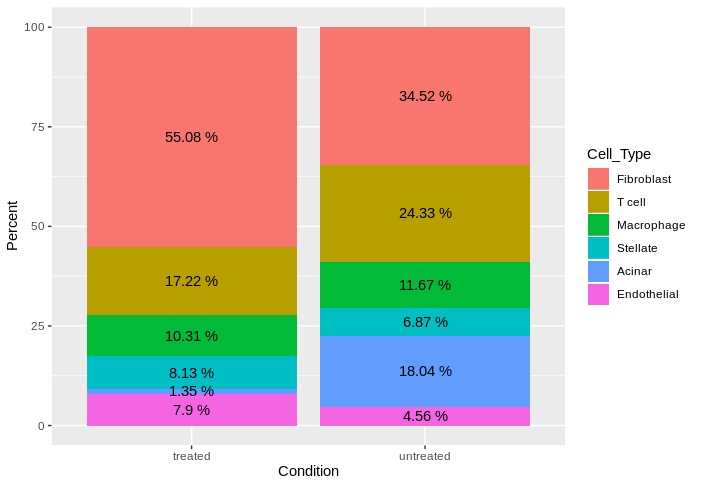
它能回答你的问题吗?
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/59699523
复制相关文章
相似问题

