适用于小信号模型的pyhf收敛失效拟合
适用于小信号模型的pyhf收敛失效拟合
提问于 2020-02-06 07:04:40
(这是我们( pyhf团队)最近遇到的一个问题,我们认为这个问题很好,值得分享。所以我们在这里发布了一个修改过的版本。
我试着用pyhf v0.4.0做一个简单的假设检验。我正在使用的模型有一个小的信号,所以我需要扫描信号强度几乎所有的出路到mu=100。然而,我一直有一个收敛问题。为什么适配不能收敛?
下面是我的环境、我正在使用的代码以及我的错误。
环境
$ "$(which python3)" --version
Python 3.7.5
$ python3 -m venv "${HOME}/.venvs/example"
$ . "${HOME}/.venvs/example/bin/activate"(example) $ python -m pip install --upgrade pip setuptools wheel
(example) $ cat requirements.txt
pyhf~=0.4.0
black
(example) $ python -m pip install -r requirements.txt
(example) $ pip list
Package Version
------------------ --------
appdirs 1.4.3
attrs 19.3.0
black 19.10b0
Click 7.0
importlib-metadata 1.5.0
jsonpatch 1.25
jsonpointer 2.0
jsonschema 3.2.0
numpy 1.18.1
pathspec 0.7.0
pip 20.0.2
pkg-resources 0.0.0
pyhf 0.4.0
pyrsistent 0.15.7
PyYAML 5.3
regex 2020.1.8
scipy 1.4.1
setuptools 45.1.0
six 1.14.0
toml 0.10.0
tqdm 4.42.1
typed-ast 1.4.1
wheel 0.34.2
zipp 2.1.0代码
# example.py
import pyhf
from pyhf import Model, infer
def main():
signal=[0.00000000e+00,2.16147594e-04,4.26391320e-04,8.53157029e-04,
7.95947245e-04,1.85458682e-03,3.15515589e-03,4.22895664e-03,
4.65887617e-03,7.35380863e-03,8.71947686e-03,7.94697901e-03,
1.02721341e-02,9.24346489e-03,9.38926633e-03,9.68742497e-03,
8.11072856e-03,7.71003446e-03,6.80873211e-03,5.43234586e-03,
4.98376829e-03,4.72218222e-03,3.40645378e-03,3.44950579e-03,
2.61473009e-03,2.18345641e-03,2.00960464e-03,1.33786215e-03,
1.18440675e-03,8.36366201e-04,5.99855228e-04,4.27406780e-04,
2.71607026e-04,1.81370902e-04,1.03710513e-04,4.42737056e-05,
2.25835175e-05,1.04470885e-05,4.08162922e-06,3.20004812e-06,
3.37990384e-07,6.72843977e-07,0.00000000e+00,9.08675772e-08,
0.00000000e+00]
bkgrd=[1.47142981e+03,9.07095061e+02,9.11188195e+02,7.06123452e+02,
6.08054685e+02,5.23577562e+02,4.41672633e+02,4.00423307e+02,
3.59576067e+02,3.26368076e+02,2.88077216e+02,2.48887339e+02,
2.20355981e+02,1.91623853e+02,1.57733823e+02,1.32733279e+02,
1.12789438e+02,9.53141118e+01,8.15735557e+01,6.89604141e+01,
5.64245978e+01,4.49094779e+01,3.95547919e+01,3.13005748e+01,
2.55212288e+01,1.93057913e+01,1.48268648e+01,1.13639821e+01,
8.64408136e+00,5.81608649e+00,3.98839138e+00,2.61636610e+00,
1.55906281e+00,1.08550560e+00,5.57450828e-01,2.25258250e-01,
2.05230728e-01,1.28735312e-01,6.13798028e-02,2.00805073e-02,
5.91436617e-02,0.00000000e+00,0.00000000e+00,0.00000000e+00,
0.00000000e+00]
spec = {
"channels": [
{
"name": "singlechannel",
"samples": [
{
"name": "signal",
"data": signal,
"modifiers": [
{"name": "mu", "type": "normfactor", "data": None}
],
},
{"name": "background", "data": bkgrd, "modifiers": [],},
],
}
]
}
model = pyhf.Model(spec)
hypo_tests = pyhf.infer.hypotest(
1.0,
model.expected_data([0]),
model,
0.5,
[(0, 80)],
return_expected_set=True,
return_test_statistics=True,
qtilde=True,
)
print(hypo_tests)
if __name__ == "__main__":
main()错误
(example) $ python example.py
/home/jovyan/.venvs/example/lib/python3.7/site-packages/pyhf/tensor/numpy_backend.py:253: RuntimeWarning: divide by zero encountered in log
return n * np.log(lam) - lam - gammaln(n + 1.0)
/home/jovyan/.venvs/example/lib/python3.7/site-packages/pyhf/tensor/numpy_backend.py:253: RuntimeWarning: invalid value encountered in multiply
return n * np.log(lam) - lam - gammaln(n + 1.0)
ERROR:pyhf.optimize.opt_scipy: fun: nan
jac: array([nan])
message: 'Iteration limit exceeded'
nfev: 1300003
nit: 100001
njev: 100001
status: 9
success: False
x: array([0.499995])
Traceback (most recent call last):
File "example.py", line 65, in <module>
main()
File "example.py", line 59, in main
qtilde=True,
File "/home/jovyan/.venvs/example/lib/python3.7/site-packages/pyhf/infer/__init__.py", line 82, in hypotest
asimov_data = generate_asimov_data(asimov_mu, data, pdf, init_pars, par_bounds)
File "/home/jovyan/.venvs/example/lib/python3.7/site-packages/pyhf/infer/utils.py", line 8, in generate_asimov_data
bestfit_nuisance_asimov = fixed_poi_fit(asimov_mu, data, pdf, init_pars, par_bounds)
File "/home/jovyan/.venvs/example/lib/python3.7/site-packages/pyhf/infer/mle.py", line 62, in fixed_poi_fit
**kwargs,
File "/home/jovyan/.venvs/example/lib/python3.7/site-packages/pyhf/optimize/opt_scipy.py", line 47, in minimize
assert result.success
AssertionError回答 1
Stack Overflow用户
回答已采纳
发布于 2020-02-06 07:38:25
从模型的角度来看,背景估计不应该是零,所以在模型中添加一个1e-7的epsilon,然后添加1%的背景不确定性。虽然这里的问题是信号强度的合理间隔在μ ∈ [0,10]之间。如果你的模型是这样的,你对这个范围内的信号强度不敏感,那么你应该测试一个新的信号模型,它是由某个尺度因子缩放的原始信号。
环境
为了便于可视化,让我们对环境进行一些扩展
(example) $ cat requirements.txt
pyhf~=0.4.0
black
matplotlib~=3.1
altair~=4.0代码
# answer.py
import pyhf
from pyhf import Model, infer
import numpy as np
import matplotlib.pyplot as plt
import pyhf.contrib.viz.brazil
def invert_interval(test_mus, hypo_tests, test_size=0.05):
cls_obs = np.array([test[0] for test in hypo_tests]).flatten()
cls_exp = [
np.array([test[1][i] for test in hypo_tests]).flatten() for i in range(5)
]
crossing_test_stats = {"exp": [], "obs": None}
for cls_exp_sigma in cls_exp:
crossing_test_stats["exp"].append(
np.interp(
test_size, list(reversed(cls_exp_sigma)), list(reversed(test_mus))
)
)
crossing_test_stats["obs"] = np.interp(
test_size, list(reversed(cls_obs)), list(reversed(test_mus))
)
return crossing_test_stats
def main():
unscaled_signal=[0.00000000e+00,2.16147594e-04,4.26391320e-04,8.53157029e-04,
7.95947245e-04,1.85458682e-03,3.15515589e-03,4.22895664e-03,
4.65887617e-03,7.35380863e-03,8.71947686e-03,7.94697901e-03,
1.02721341e-02,9.24346489e-03,9.38926633e-03,9.68742497e-03,
8.11072856e-03,7.71003446e-03,6.80873211e-03,5.43234586e-03,
4.98376829e-03,4.72218222e-03,3.40645378e-03,3.44950579e-03,
2.61473009e-03,2.18345641e-03,2.00960464e-03,1.33786215e-03,
1.18440675e-03,8.36366201e-04,5.99855228e-04,4.27406780e-04,
2.71607026e-04,1.81370902e-04,1.03710513e-04,4.42737056e-05,
2.25835175e-05,1.04470885e-05,4.08162922e-06,3.20004812e-06,
3.37990384e-07,6.72843977e-07,0.00000000e+00,9.08675772e-08,
0.00000000e+00]
bkgrd=[1.47142981e+03,9.07095061e+02,9.11188195e+02,7.06123452e+02,
6.08054685e+02,5.23577562e+02,4.41672633e+02,4.00423307e+02,
3.59576067e+02,3.26368076e+02,2.88077216e+02,2.48887339e+02,
2.20355981e+02,1.91623853e+02,1.57733823e+02,1.32733279e+02,
1.12789438e+02,9.53141118e+01,8.15735557e+01,6.89604141e+01,
5.64245978e+01,4.49094779e+01,3.95547919e+01,3.13005748e+01,
2.55212288e+01,1.93057913e+01,1.48268648e+01,1.13639821e+01,
8.64408136e+00,5.81608649e+00,3.98839138e+00,2.61636610e+00,
1.55906281e+00,1.08550560e+00,5.57450828e-01,2.25258250e-01,
2.05230728e-01,1.28735312e-01,6.13798028e-02,2.00805073e-02,
5.91436617e-02,0.00000000e+00,0.00000000e+00,0.00000000e+00,
0.00000000e+00]
scale_factor = 500
signal = np.asarray(unscaled_signal) * scale_factor
epsilon = 1e-7
background = np.asarray(bkgrd) + epsilon
spec = {
"channels": [
{
"name": "singlechannel",
"samples": [
{
"name": "signal",
"data": signal.tolist(),
"modifiers": [
{"name": "mu", "type": "normfactor", "data": None}
],
},
{
"name": "background",
"data": background.tolist(),
"modifiers": [
{
"name": "uncert",
"type": "shapesys",
"data": (0.01 * background).tolist(),
},
],
},
],
}
]
}
model = pyhf.Model(spec)
init_pars = model.config.suggested_init()
par_bounds = model.config.suggested_bounds()
data = model.expected_data(init_pars)
cls_obs, cls_exp = pyhf.infer.hypotest(
1.0,
data,
model,
init_pars,
par_bounds,
return_expected_set=True,
return_test_statistics=True,
qtilde=True,
)
# Show that the scale factor chosen gives reasonable values
print(f"Observed CLs for µ=1: {cls_obs[0]:.2f}")
print("-----")
for idx, n_sigma in enumerate(np.arange(-2, 3)):
print(
"Expected {}CLs for µ=1: {:.3f}".format(
" " if n_sigma == 0 else "({} σ) ".format(n_sigma),
cls_exp[idx][0],
)
)
# Perform hypothesis test scan
_start = 0.1
_stop = 4
_step = 0.1
poi_tests = np.arange(_start, _stop + _step, _step)
print("\nPerforming hypothesis tests\n")
hypo_tests = [
pyhf.infer.hypotest(
mu_test,
data,
model,
init_pars,
par_bounds,
return_expected_set=True,
return_test_statistics=True,
qtilde=True,
)
for mu_test in poi_tests
]
# This is all you need. Below is just to demonstrate.
# Upper limits on signal strength
results = invert_interval(poi_tests, hypo_tests)
print(f"Observed Limit on µ: {results['obs']:.2f}")
print("-----")
for idx, n_sigma in enumerate(np.arange(-2, 3)):
print(
"Expected {}Limit on µ: {:.3f}".format(
" " if n_sigma == 0 else "({} σ) ".format(n_sigma),
results["exp"][idx],
)
)
# Visualize the "Brazil band"
fig, ax = plt.subplots()
fig.set_size_inches(7, 5)
ax.set_title("Hypothesis Tests")
ax.set_ylabel("CLs")
ax.set_xlabel(f"µ (for Signal x {scale_factor})")
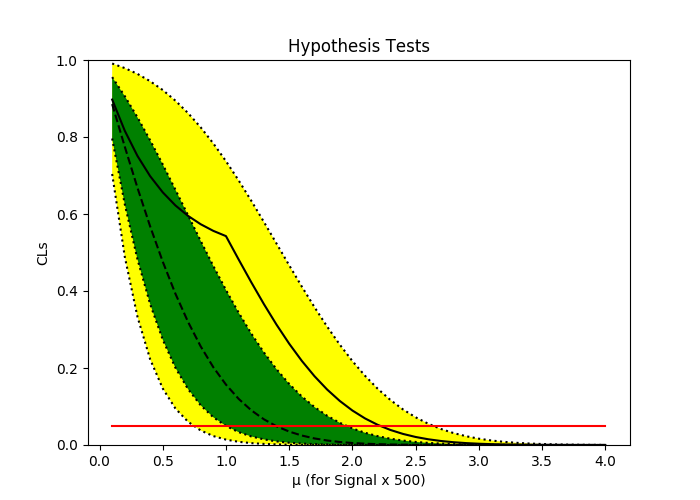
pyhf.contrib.viz.brazil.plot_results(ax, poi_tests, hypo_tests)
fig.savefig("brazil_band.pdf")
if __name__ == "__main__":
main()输出
信号需要缩放的值可以通过尝试几个缩放因子值来确定,直到mu=1的信号强度的CLs值开始看起来合理(比1e-3更大的值)。在这个特殊的例子中,500的比例因子似乎没问题。
非标度信号强度的上限就是观察到的极限除以尺度因子,在这种情况下没有明显的灵敏度。
(example) $ python answer.py
Observed CLs for µ=1: 0.54
-----
Expected (-2 σ) CLs for µ=1: 0.014
Expected (-1 σ) CLs for µ=1: 0.049
Expected CLs for µ=1: 0.157
Expected (1 σ) CLs for µ=1: 0.403
Expected (2 σ) CLs for µ=1: 0.737
Performing hypothesis tests
Observed Limit on µ: 2.22
-----
Expected (-2 σ) Limit on µ: 0.746
Expected (-1 σ) Limit on µ: 0.998
Expected Limit on µ: 1.392
Expected (1 σ) Limit on µ: 1.953
Expected (2 σ) Limit on µ: 2.638
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/60089405
复制相关文章
相似问题

