"Unnest“重叠时间间隔
"Unnest“重叠时间间隔
提问于 2020-12-22 16:54:29
我正试图为一组以领先/滞后方式工作的过滤器创建一些情节。
关于领先/滞后的简短描述:
当一个新的过滤器上线时,它被放置在滞后位置,这意味着水通过主过滤器(也称铅过滤器)后通过它。当铅过滤器堵塞时,电流滞后滤波器被移动到引线位置。总之,过滤器从滞后位置开始,然后被撞到引线位置。
从视觉上看,你可以这样想象:
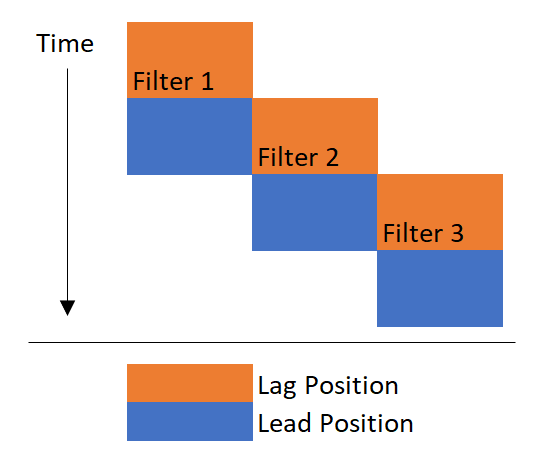
我需要做的是“不嵌套”(因为没有一个更好的词)有重叠的时间。换句话说,我希望每个过滤器都有一个连续运行的时间戳,而不管它所处的领先/滞后位置。
这些数据的结构如下:
data <- structure(list(record_timestamp = structure(c(1608192000, 1608192060, 1608192120, 1608192180, 1608192240, 1608192300, 1608192360, 1608192420, 1608192480, 1608192540, 1608192600, 1608192660, 1608192720, 1608192780, 1608192840, 1608192900, 1608192960, 1608193020, 1608193080, 1608193140, 1608193200, 1608193260, 1608193320, 1608193380, 1608193440, 1608193500, 1608193560, 1608193620, 1608193680, 1608193740, 1608193800), class = c("POSIXct", "POSIXt"), tzone = "UTC"), flow = c(20, 20, 20, 20, 20, 20, 20, 20, 20, 20, 20, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10), lag_start = structure(c(1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260), class = c("POSIXct", "POSIXt"), tzone = "UTC"), lead_start = structure(c(NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260), class = c("POSIXct", "POSIXt"), tzone = "UTC"), changeout_interval = new("Interval", .Data = c(0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 660, 0, 0, 0, 0, 0, 0, 0, 0, 0, 600, 0, 0, 0, 0, 0, 0, 0, 0, 0, NA), start = structure(c(1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192000, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260, 1608193260 ), tzone = "UTC", class = c("POSIXct", "POSIXt")), tzone = "UTC")), class = c("spec_tbl_df", "tbl_df", "tbl", "data.frame"), row.names = c(NA, -31L), spec = structure(list( cols = list(record_timestamp = structure(list(), class = c("collector_character", "collector")), flow = structure(list(), class = c("collector_double", "collector")), polish_start = structure(list(), class = c("collector_character", "collector")), lead_start = structure(list(), class = c("collector_character", "collector"))), default = structure(list(), class = c("collector_guess", "collector")), skip = 1), class = "col_spec"))我对最终结果的设想数据看起来是这样的:
end_data <- structure(list(record_timestamp = structure(c(1608192000, 1608192060,1608192120, 1608192180, 1608192240, 1608192300, 1608192360, 1608192420,1608192480, 1608192540, 1608192600, 1608192660, 1608192720, 1608192780,1608192840, 1608192900, 1608192960, 1608193020, 1608193080, 1608193140,1608193200, 1608192660, 1608192720, 1608192780, 1608192840, 1608192900,1608192960, 1608193020, 1608193080, 1608193140, 1608193200, 1608193260,1608193320, 1608193380, 1608193440, 1608193500, 1608193560,1608193620,1608193680, 1608193740, 1608193800), class = c("POSIXct", "POSIXt"), tzone = "UTC"), flow = c(20, 20, 20, 20, 20, 20, 20, 20, 20, 20, 20, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA), lag_start = structure(c(1608192000, 1608192000, 1608192000,1608192000, 1608192000, 1608192000, 1608192000, 1608192000,1608192000,1608192000, 1608192000, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660, 1608192660,1608192660, 1608192660, 1608192660, 1608192660, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA), class = c("POSIXct", "POSIXt"), tzone = "UTC"), lead_start = structure(c(NA, NA, NA, NA, NA, NA, NA, NA,NA, NA, NA, 1608192660, 1608192660, 1608192660, 1608192660,1608192660, 1608192660, 1608192660, 1608192660, 1608192660,1608192660, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, 1608193260,1608193260, 1608193260, 1608193260, 1608193260, 1608193260,1608193260, 1608193260, 1608193260, 1608193260), class = c("POSIXct","POSIXt"), tzone = "UTC"), filter_id = c(1, 1, 1, 1, 1, 1,1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2)), class = c("spec_tbl_df", "tbl_df", "tbl", "data.frame"), row.names = c(NA, -41L), spec = structure(list(cols = list(record_timestamp = structure(list(), class = c("collector_character","collector")), flow = structure(list(), class = c("collector_double","collector")), polish_start = structure(list(), class = c("collector_character","collector")), lead_start = structure(list(), class = c("collector_character", "collector")), filter_id = structure(list(), class = c("collector_double","collector"))), default = structure(list(), class = c("collector_guess","collector")), skip = 1), class = "col_spec"))这将使时间戳加倍,但它将允许更容易地绘制,因为我可以在group_by列上使用filter_id。
到目前为止,我有一组时间间隔,为每个过滤器,从开始到结束,领导通过滞后。这是密码:
intervals <- data %>%
distinct(lag_start, .keep_all = TRUE) %>%
mutate(changeout_interval = interval(lag_start, lead(lag_start, 2))) %>%
select(record_timestamp, changeout_interval)从那里,我如何过滤所有的时间戳,属于每一个时间间隔?就像有条件的pivot_longer。
最终目标是能够用几行ggplot2来绘制过滤器的完整生命周期,包括领导和滞后。下面是我对情节的设想:
grouped_data <- data %>%
group_by(lag_start) %>%
mutate(elapsed_time = difftime(record_timestamp,
record_timestamp[1],
units = "mins"),
total_flow = cumsum(flow))
ggplot(grouped_data, aes(x = elapsed_time, y = total_flow)) +
geom_line(aes(color = as.factor(lag_start)))但是,这个图不包括每个过滤器的流,当它变成引线位置时。
回答 1
Stack Overflow用户
回答已采纳
发布于 2020-12-22 19:12:28
使用dense_rank按lag_start对筛选器进行分组,然后每个过滤器创建一个记录。这使得信息以宽格式保存,因为interval和end_data具有不同的数据结构。
library(dplyr)
library(lubridate)
data %>%
select(-changeout_interval) %>% # example only as interval appeared to calculate this
mutate(filter_id = dense_rank(lag_start)) %>%
group_by(filter_id) %>%
slice(1) %>%
ungroup() %>%
mutate(lead_start = lead(lead_start), lead_end = lead(lead_start), changeout_interval = interval(lag_start, lead_end))
# A tibble: 3 x 7
record_timestamp flow lag_start lead_start filter_id lead_end
<dttm> <dbl> <dttm> <dttm> <int> <dttm>
1 2020-12-17 08:00:00 20 2020-12-17 08:00:00 2020-12-17 08:11:00 1 2020-12-17 08:21:00
2 2020-12-17 08:11:00 15 2020-12-17 08:11:00 2020-12-17 08:21:00 2 NA
3 2020-12-17 08:21:00 10 2020-12-17 08:21:00 NA 3 NA 更新,以回应对问题的澄清补充。使用相同的dense_rank方法,然后通过pivot_longer切换到长格式,从而使cumsum需求更易于绘制。
library(dplyr)
library(tidyr)
library(ggplot2)
plot_data <- data %>%
select(-changeout_interval) %>% # example only as interval appeared to calculate this
mutate(filter_lag = dense_rank(lag_start),
filter_lead = filter_lag - 1) %>%
select(-lag_start, -lead_start) %>%
pivot_longer(cols = starts_with("filter_"),
names_to = "position",
names_prefix = "filter_",
values_to = "filter") %>%
filter(filter > 0) %>% # drops the starting filter as data shows no lead filter?
group_by(filter) %>%
mutate(elapsed_time = difftime(record_timestamp, record_timestamp[1], units = "mins"),
rolling_flow = cumsum(flow))绘制elapsed_time和rolling_flow
ggplot(plot_data, aes(x = as.numeric(elapsed_time),
y = rolling_flow,
color = factor(filter))) +
geom_line()
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/65412698
复制相关文章
相似问题

