比较顶点查找表和边缘列表的有效方法,并将顶点标签分配给任何匹配的边缘
比较顶点查找表和边缘列表的有效方法,并将顶点标签分配给任何匹配的边缘
提问于 2021-03-01 14:36:03
我有两个数据帧。首先,包含顶点名称列表的查找表:
lookup <- data.frame(Name=c("Bob","Jane"))然后我有一个边缘列表,如下所示:
edges <- data.frame(vertex1 = c("Bob","Bill","Bob","Jane","Bill","Jane","Bob","Jane","Bob","Bill","Bob"
,"Jane","Bill","Jane","Bob","Jane","Jane","Jill","Jane","Susan","Susan"),
edgeID = c(1,1,1,1,1,1,2,2,1,1,1,1,1,1,2,2,3,3,3,3,3),
vertex2 = c("Bill","Bob","Jane","Bob","Jane","Jill","Jane","Bob","Bill","Bob"
,"Jane","Bob","Jane","Bill","Jane","Bob","Jill","Jane","Susan","Jane","Jill"))对于“查找”表中的每个唯一顶点,我想迭代“边”表,并标记每个edgeID,其中查找$Name就在顶点中。
我可以用下面的脚本来完成这个任务:
library(igraph)
g <- graph_from_data_frame(edges[c(1, 3, 2)], directed = FALSE)
do.call(
rbind,
c(
make.row.names = FALSE,
lapply(
as.character(lookup$Name),
function(nm) {
z <- c(nm, V(g)$name[distances(g, nm) == 1])
cbind(group = nm, unique(subset(edges, vertex1 %in% z & vertex2 %in% z)))
}
)
)
) group vertex1 edgeID vertex2
1 Bob Bob 1 Bill
2 Bob Bill 1 Bob
3 Bob Bob 1 Jane
4 Bob Jane 1 Bob
5 Bob Bill 1 Jane
6 Bob Bob 2 Jane
7 Bob Jane 2 Bob
8 Bob Jane 1 Bill
9 Jane Bob 1 Bill
10 Jane Bill 1 Bob
11 Jane Bob 1 Jane
12 Jane Jane 1 Bob
13 Jane Bill 1 Jane
14 Jane Jane 1 Jill
15 Jane Bob 2 Jane
16 Jane Jane 2 Bob
17 Jane Jane 1 Bill
18 Jane Jane 3 Jill
19 Jane Jill 3 Jane
20 Jane Jane 3 Susan
21 Jane Susan 3 Jane
22 Jane Susan 3 Jill问题是,对于大型边缘列表来说,这似乎效率低下。在我的实际数据中,“查找”有3263个观测值,而“边”有167,775,170个观测值。我已经尝试在一个16核和100 of或RAM的亚马逊EC2实例上运行上面的脚本,但是没有结束(使用"future_lapply“而不是"lapply”来允许并行处理)。有什么办法可以让这件事更有效率/更快吗?
这并不是我唯一需要对边缘进行分组的时候,我希望能找到一种在时间和亚马逊账单方面不那么昂贵的方法。
回答 1
Stack Overflow用户
回答已采纳
发布于 2021-03-01 16:06:55
我认为您可以首先缩小原始的data.frame edges,然后可以避免每次迭代在lapply中使用unique。
下面的代码可能会加速一些,但不确定它是如何在实际数据中获得的。
edges.unique <- unique(edges[c(1, 3, 2)])
g <- graph_from_data_frame(edges.unique, directed = FALSE)
do.call(
rbind,
c(
make.row.names = FALSE,
lapply(
lookup$Name,
function(nm) {
z <- colnames(d <- distances(g, nm))[which(d < 2)]
cbind(group = nm, subset(edges.unique, vertex1 %in% z & vertex2 %in% z))
}
)
)
)更新
edges.unique <- unique(
transform(
edges[c("vertex1", "vertex2", "edgeID")],
vertex1 = ifelse(vertex1 < vertex2, vertex1, vertex2),
vertex2 = ifelse(vertex1 < vertex2, vertex2, vertex1)
)
)
g <- graph_from_data_frame(edges.unique, directed = FALSE)
res <- do.call(
rbind,
c(
make.row.names = FALSE,
lapply(
lookup$Name,
function(nm) {
z <- colnames(d <- distances(g, nm))[which(d < 2)]
cbind(group = nm, subset(edges.unique, vertex1 %in% z & vertex2 %in% z))
}
)
)
)给出
> res
group vertex1 vertex2 edgeID
1 Bob Bill Bob 1
2 Bob Bob Jane 1
3 Bob Bill Jane 1
4 Bob Bob Jane 2
5 Jane Bill Bob 1
6 Jane Bob Jane 1
7 Jane Bill Jane 1
8 Jane Jane Jill 1
9 Jane Bob Jane 2
10 Jane Jane Jill 3
11 Jane Jane Susan 3
12 Jane Jill Susan 3当您键入plot(g)时,您将看到下面的简化
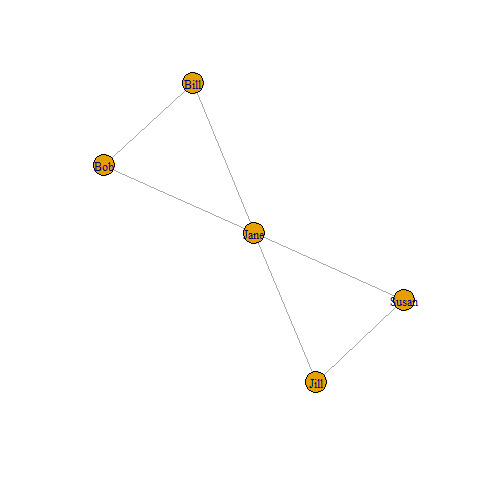
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/66424061
复制相关文章
相似问题

