使用ggnewscale::new_scale()和R中的ggplot2将图例拆分成两个或多个列
使用ggnewscale::new_scale()和R中的ggplot2将图例拆分成两个或多个列
提问于 2021-08-11 15:11:29
在following Stefan's very helpful answer to this post (他使用ggnewscale::new_scale()的地方)之后,我现在被困在以下问题上:
如何将自定义ggnewscale 中的自定义图例排列为多个垂直列?与其类似,通常使用诸如ggplot2中的guides(scale_shape_manual=guide_legend(ncol=2))这样的命令来完成。
最小可重现性示例:
# fictional data in the scheme of https://stackoverflow.com/q/66804487/16642045
mutated <- list()
for(i in 1:10) {
mutated[[i]] <- data.frame(Biological.Replicate = rep(1,4),
Reagent.Conc = c(10000, 2500, 625, 156.3),
Reagent = rep(1,8),
Cell.type = rep(LETTERS[i],4),
Mean.Viable.Cells.1 = rep(runif(n = 10, min = 0, max = 1),4))
}
mutated <- do.call(rbind.data.frame, mutated)用户"stefan“回答后的修改代码如下所示:
# from https://stackoverflow.com/a/66808609/16642045
library(ggplot2)
library(ggnewscale)
library(dplyr)
library(magrittr)
mutated <- mutated %>%
mutate(Cell.type = as.factor(Cell.type),
Reagent = factor(Reagent,
levels = c("0", "1", "2")))
mean_mutated <- mutated %>%
group_by(Reagent, Reagent.Conc, Cell.type) %>%
split(.$Cell.type)
layer_geom_scale <- function(Cell.type) {
list(geom_point(mean_mutated[[Cell.type]], mapping = aes(shape = Reagent)),
geom_line(mean_mutated[[Cell.type]], mapping = aes(group = Reagent, linetype = Reagent)),
scale_linetype_manual(name = Cell.type, values = c("solid", "dashed", "dotted"), drop=FALSE),
scale_shape_manual(name = Cell.type, values = c(15, 16, 4), labels = c("0", "1", "2"), drop=FALSE)
)
}
my_plot <-
ggplot(mapping = aes(
x = as.factor(Reagent.Conc),
y = Mean.Viable.Cells.1)) +
layer_geom_scale(unique(mutated$Cell.type)[1])
for(current_Cell.type_index in 2:length(unique(mutated$Cell.type))) {
my_plot <-
my_plot +
ggnewscale::new_scale("shape") +
ggnewscale::new_scale("linetype") +
layer_geom_scale(unique(mutated$Cell.type)[current_Cell.type_index])
}
my_plot这导致:
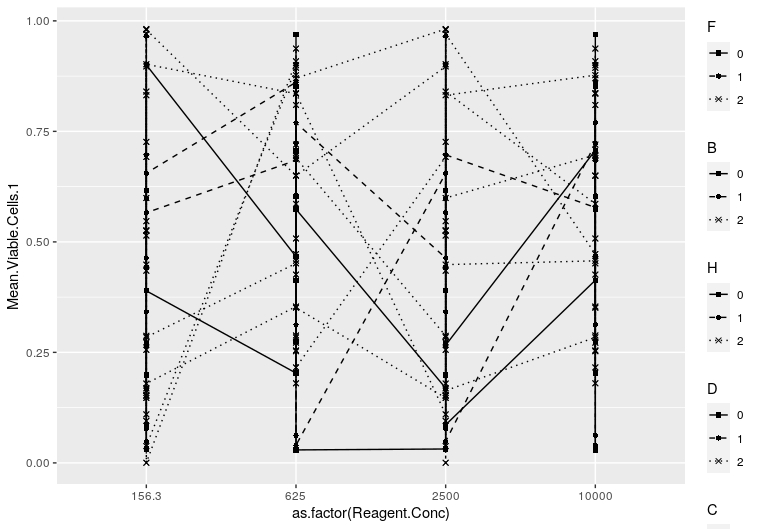
现在,--我希望在两列中并行显示传奇,和我尝试过这样做(但没有成功):
my_plot +
guides(scale_shape_manual=guide_legend(ncol=2))编辑:一张我希望传奇故事被安排的图片,
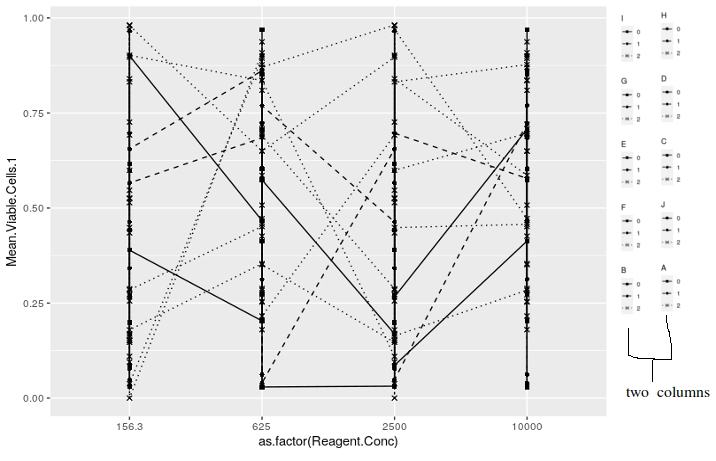
有人能帮我吗?
谢谢!
回答 1
Stack Overflow用户
发布于 2021-08-12 08:47:33
注意:这个答案解决了问题澄清之前的问题,编辑#4及更高版本。
水平传说
添加theme(legend.box = "horizontal")将使图例元素并排出现。
多列与ggnewscale
在全球范围内使用guides将导致在 ggnewscale更新之后对scales 进行修改。在这方面,只更新变量RKO:
layer_geom_scale <- function(cell_type, color) {
list(geom_point(mean_mutated[[cell_type]], mapping = aes(shape = Reagent), color = color),
geom_line(mean_mutated[[cell_type]], mapping = aes(group = Reagent, linetype = Reagent), color = color),
scale_linetype_manual(name = cell_type, values = c("solid", "dashed", "dotted"), drop=FALSE),
scale_shape_manual(name = cell_type, values = c(15, 16, 4), labels = c("0", "1", "2"), drop=FALSE)
)
}
my_plot <-
ggplot(mapping = aes(
x = as.factor(Reagent.Conc),
y = Mean.Viable.Cells.1)) +
layer_geom_scale("HCT", "#999999") +
ggnewscale::new_scale("linetype") +
ggnewscale::new_scale("shape") +
layer_geom_scale("RKO", "#E69F00") +
theme(legend.box = "horizontal") +
guides(shape = guide_legend(ncol = 2),
linetype = guide_legend(ncol = 2))
my_plot
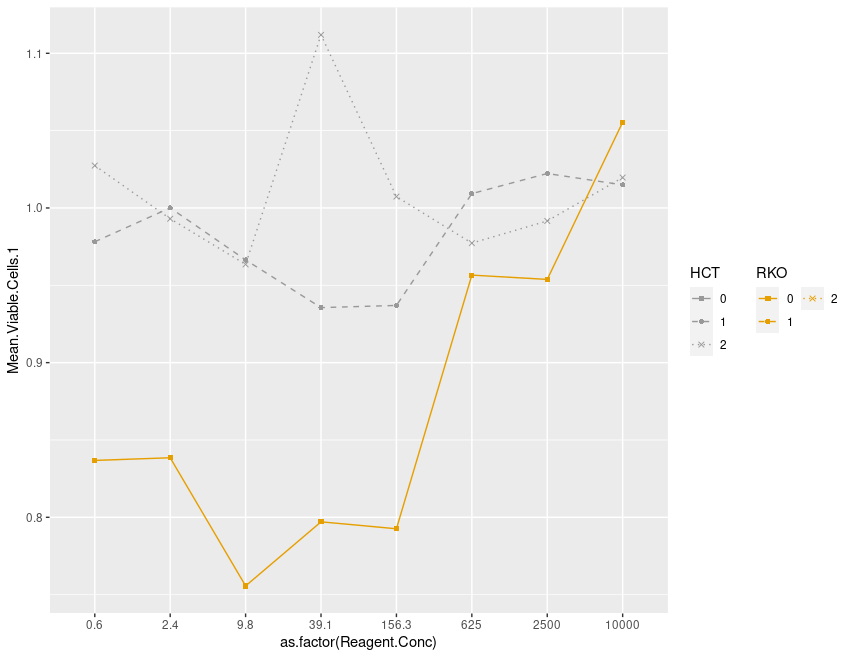
要修改所有变量的相同比例,应该在刻度定义中添加guide:
layer_geom_scale <- function(cell_type, color) {
list(geom_point(mean_mutated[[cell_type]], mapping = aes(shape = Reagent), color = color),
geom_line(mean_mutated[[cell_type]], mapping = aes(group = Reagent, linetype = Reagent), color = color),
scale_linetype_manual(name = cell_type, values = c("solid", "dashed", "dotted"), drop=FALSE,
guide = guide_legend(ncol = 2)),
scale_shape_manual(name = cell_type, values = c(15, 16, 4), labels = c("0", "1", "2"), drop=FALSE,
guide = guide_legend(ncol = 2))
)
}
my_plot <-
ggplot(mapping = aes(
x = as.factor(Reagent.Conc),
y = Mean.Viable.Cells.1)) +
layer_geom_scale("HCT", "#999999") +
ggnewscale::new_scale("linetype") +
ggnewscale::new_scale("shape") +
layer_geom_scale("RKO", "#E69F00") +
theme(legend.box = "horizontal")
my_plot
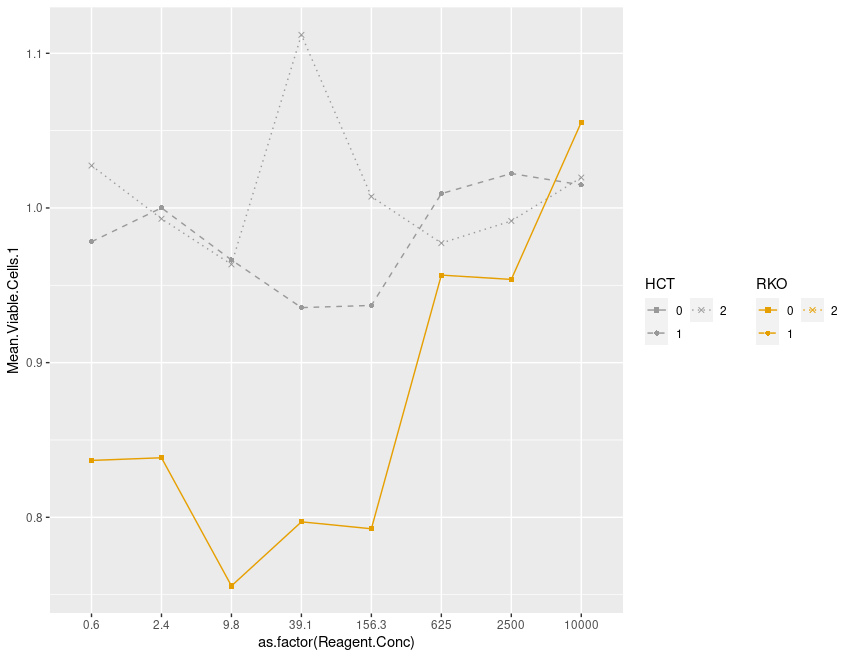
原始数据
# from https://stackoverflow.com/q/66804487/16642045
mutated <- structure(list(
Biological.Replicate = c(1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L),
Reagent.Conc = c(10000, 2500, 625, 156.3,
39.1, 9.8, 2.4, 0.6,
10000, 2500, 625, 156.3,
39.1, 9.8, 2.4, 0.6,
10000, 2500, 625, 156.3,
39.1, 9.8, 2.4, 0.6),
Reagent = c(1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L,
2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L,
0L, 0L, 0L, 0L,
0L, 0L, 0L, 0L),
Cell.type = c("HCT", "HCT", "HCT", "HCT",
"HCT", "HCT", "HCT", "HCT",
"HCT", "HCT", "HCT", "HCT",
"HCT", "HCT", "HCT", "HCT",
"RKO", "RKO", "RKO", "RKO",
"RKO", "RKO", "RKO", "RKO"),
Mean.Viable.Cells.1 = c(1.014923966, 1.022279854, 1.00926559, 0.936979842,
0.935565248, 0.966403395, 1.00007073, 0.978144524,
1.019673384, 0.991595836, 0.977270557, 1.007353643,
1.111928183, 0.963518289, 0.993028364, 1.027409034,
1.055452733, 0.953801253, 0.956577449, 0.792568337,
0.797052961, 0.755623576, 0.838482346, 0.836773918)),
row.names = 9:32,
class = "data.frame")
# from https://stackoverflow.com/a/66808609/16642045
library(ggplot2)
library(ggnewscale)
library(dplyr)
library(magrittr)
mutated <- mutated %>%
mutate(Cell.type = factor(Cell.type,
levels = c("HCT", "RKO")),
Reagent = factor(Reagent,
levels = c("0", "1", "2")))
mean_mutated <- mutated %>%
group_by(Reagent, Reagent.Conc, Cell.type) %>%
split(.$Cell.type)页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/68744642
复制相关文章
相似问题

