R中单变量异常值的快速检测/删除方法
R中单变量异常值的快速检测/删除方法
提问于 2022-12-02 15:57:11
是否有比下面的例子更快的方法来检测R中的异常值?
需求:异常值应该由结果向量中的NA来表示。
vals = c(6.4, 1.786, 5.934, 6.689, 6.098, 6.177, 6.768, 6.31, 6.164,
1.543, 6.242, 6.107, 6.708, 6.184, 6.102, 6.495, 6.423, 6.489,
5.264, 5.09, 5.915, 6.114, 5.395, 5.991, 6.732, 6.143, 6.657,
5.563, 5.173, 5.439, 4.305, 6.867, 5.007, 6.37, 6.193, 5.504,
6.333, 6.25, 0.206, 5.911, 5.496, 0.093, 6.554, 6.25, 6.526,
6.202, 6.305, 5.977, 6.476, 5.903, 5.758, 5.117, 6.985, 6.485,
0.763, 5.368, 5.146, 3.079, 5.823, 5.627, 6.077, 6.346, 5.301,
5.555, 6.02, 6.914, 5.896, 5.458, 6.473, 7.348, 7.649, 6.464,
6.545, 6.673, 6.618, 6.659)
detect_outliers = function(x, na.rm = TRUE, ...) {
qnt = stats::quantile(x, probs=c(.25, .75), na.rm = na.rm, ...)
H = 1.5 * stats::IQR(x, na.rm = na.rm)
y = x
y[x < (qnt[1] - H)] = NA
y[x > (qnt[2] + H)] = NA
y
}
detect_outliers2 = function(x, ...) {
out = suppressMessages(univOutl::boxB(x, ...))
x[out$outliers] = NA
x
}
detect_outliers3 = function(x) {
out = graphics::boxplot(x, plot=FALSE)$out
x[fastmatch::`%fin%`(x, out)] = NA
x
}
detect_outliers4 = function(x) {
out = grDevices::boxplot.stats(x)$out
x[fastmatch::`%fin%`(x, out)] = NA
x
}
detect_outliers5 = function(x) {
out = rstatix::identify_outliers(data.frame(x))
x[fastmatch::`%fin%`(x, out$x)] = NA
x
}
detect_outliers6 = function(x) {
dev = abs(x-median(x)) # absolute deviation from median
MAD = median(abs(dev)) # median absolute deviation
sd = MAD/0.67449
x[dev > 2*sd] = NA
x
}
rbenchmark::benchmark("detect_outliers" = detect_outliers(vals),
"detect_outliers2" = detect_outliers2(vals),
"detect_outliers3" = detect_outliers3(vals),
"detect_outliers4" = detect_outliers4(vals),
"detect_outliers5" = detect_outliers5(vals),
"detect_outliers6" = detect_outliers6(vals),
replications = 1000,
columns = c("test", "replications", "elapsed",
"relative", "user.self", "sys.self"))基准测试结果
test replications elapsed relative user.self sys.self
1 detect_outliers 1000 0.198 3.600 0.198 0.001
2 detect_outliers2 1000 0.350 6.364 0.331 0.019
3 detect_outliers3 1000 0.105 1.909 0.105 0.000
4 detect_outliers4 1000 0.070 1.273 0.070 0.000
5 detect_outliers5 1000 5.245 95.364 5.224 0.004
6 detect_outliers6 1000 0.055 1.000 0.055 0.001野值移除
df = data.frame(method = factor(c(rep("detect_outliers", length(vals)),
rep("detect_outliers2", length(vals)),
rep("detect_outliers3", length(vals)),
rep("detect_outliers4", length(vals)),
rep("detect_outliers5", length(vals)),
rep("detect_outliers6", length(vals))),
levels = rev(c("detect_outliers",
"detect_outliers2",
"detect_outliers3",
"detect_outliers4",
"detect_outliers5",
"detect_outliers6"))),
orig = rep(vals, 6),
outlier_removed = c(detect_outliers(vals),
detect_outliers2(vals),
detect_outliers3(vals),
detect_outliers4(vals),
detect_outliers5(vals),
detect_outliers6(vals)))
df$is_outlier = factor(ifelse(is.na(df$outlier_removed), "yes", "no"),
levels = c("yes", "no"))
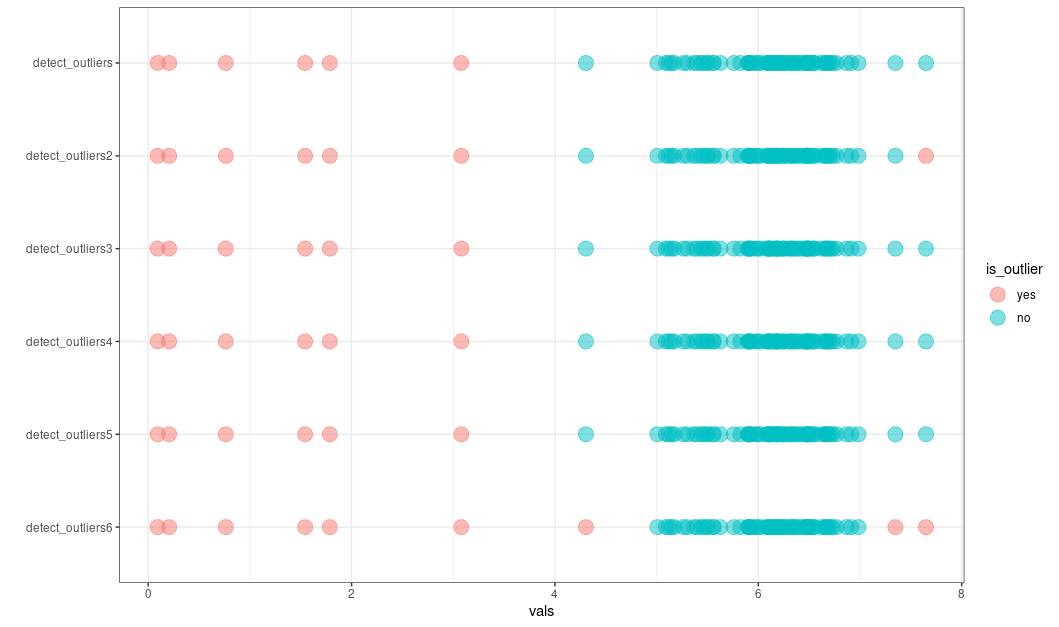
ggplot2::ggplot(df, ggplot2::aes(x = method,
y = orig,
color = is_outlier)) +
ggplot2::geom_point(alpha = 0.5, size = 5) +
ggplot2::theme_bw() +
ggplot2::labs(x = "", y = "vals") +
ggplot2::coord_flip()
回答 1
Stack Overflow用户
发布于 2022-12-02 16:44:55
您可以使用identify_outliers() rstatix包。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/74658406
复制相关文章
相似问题

